Abstract
The global emergence of many severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) variants jeopardizes the protective antiviral immunity induced after infection or vaccination. To address the public health threat caused by the increasing SARS-CoV-2 genomic diversity, the National Institute of Allergy and Infectious Diseases within the National Institutes of Health established the SARS-CoV-2 Assessment of Viral Evolution (SAVE) programme. This effort was designed to provide a real-time risk assessment of SARS-CoV-2 variants that could potentially affect the transmission, virulence, and resistance to infection- and vaccine-induced immunity. The SAVE programme is a critical data-generating component of the US Government SARS-CoV-2 Interagency Group to assess implications of SARS-CoV-2 variants on diagnostics, vaccines and therapeutics, and for communicating public health risk. Here we describe the coordinated approach used to identify and curate data about emerging variants, their impact on immunity and effects on vaccine protection using animal models. We report the development of reagents, methodologies, models and notable findings facilitated by this collaborative approach and identify future challenges. This programme is a template for the response to rapidly evolving pathogens with pandemic potential by monitoring viral evolution in the human population to identify variants that could reduce the effectiveness of countermeasures.
Similar content being viewed by others
Main
SARS-CoV-2, the aetiological agent of coronavirus disease 2019 (COVID-19), has caused a devastating pandemic resulting in more than 6 million deaths worldwide (https://covid19.who.int). With continuous transmission cycles occurring around the world, SARS-CoV-2 variants have arisen with mutations throughout its genome, including in the spike protein gene, the principal antigenic target of all SARS-CoV-2 vaccines currently in use1,2. The rapid emergence of variants—the latest being Omicron in November 2021—has raised concerns about how new mutations affect virus replication, infectivity, transmission and infection, and vaccine-induced immunity. This rapid genetic evolution of SARS-CoV-2 created an immediate need to monitor and characterize variants for potential resistance to medical countermeasures.
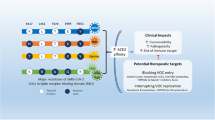
The US Department of Health and Human Services established the SARS-CoV-2 Interagency Group (SIG) to maximize coordination between the Centers for Disease Control and Prevention, the National Institutes of Health (NIH), the Food and Drug Administration, the Biomedical Advanced Research and Development Authority and Department of Defense for the US public health response to the COVID-19 pandemic3. The National Institute of Allergy and Infectious Diseases (NIAID) formed the SAVE consortium in January 2021 as a critical data-generating component for the SIG and to facilitate rapid data sharing with global partners and the scientific community (Fig. 1). The SAVE programme provides a comprehensive real-time risk assessment of emerging mutations in SARS-CoV-2 strains that could affect transmissibility, virulence, and infection- or vaccine-induced immunity. SAVE was constructed as a rational, structured and iterative risk-assessment pipeline with a goal of providing critical data to support SIG actions and ensure the effectiveness of countermeasures against emerging variants.
The SAVE programme is divided into three working groups to provide real-time risk assessments of SARS-CoV-2 variants on infection and vaccine-induced immunity. The early-detection and analysis group curates and prioritizes emerging SARS-CoV-2 variants. The in vitro group evaluates the effect of SARS-CoV-2 variants on humoral and cell-mediated immune responses. The in vivo group uses animal models to test vaccine efficacy, transmission, and define immune mechanisms and correlates of protection. These data are fed into the SIG, which coordinates between different US government agencies to assess the impact of variants on critical SARS-CoV-2 countermeasures, including vaccines, therapeutics and diagnostics. This iterative approach allows for information flow between the SAVE programme and the SIG to continue prioritizing and testing SARS-CoV-2 variants.
The SAVE programme is composed of an international team of scientists with expertise in virology, immunology, vaccinology, structural biology, bioinformatics, viral genetics and evolution. Each team member is responsible for key contributions ranging from curation of viral mutations, bioinformatics analysis, development of new reagents, assay development and testing, in vitro characterization, and in vivo model development and countermeasure testing. The SAVE programme is divided into three working groups: (1) the early-detection and analysis group; (2) the in vitro group; and (3) the in vivo group. The early-detection group uses public databases and analysis tools to curate and prioritize emerging SARS-CoV-2 variants. The in vitro group evaluates the impact of SARS-CoV-2 variants on humoral and cell-mediated immune responses using in vitro assays. The in vivo group uses small and large animal models to test vaccine efficacy, transmission, and define immune mechanisms and correlates of protection. A common theme across these subgroups is the integration of orthogonal experimental and computational approaches to validate findings and strengthen the evidence for recommendations. Collaborative efforts between the early-detection geneticists and evolutionary biologists, and the in vitro group virologists/immunologists enable the rapid determination of relationships between viral evolution and neutralization sensitivity. In turn, these results enable the in vivo team to assess and evaluate vaccine protection in animal studies. The SAVE programme has regularly scheduled (usually weekly) meetings that include individual subgroup meetings and an all-hands meeting, which serves as an opportunity to share key information across groups and align priorities for the most urgent experimental questions. NIAID programme staff and intramural and extramural scientists share leadership responsibilities. Collaboration within and across these groups has accelerated research and discovery due to the immediate and open sharing of ideas, reagents, protocols and data4,5,6,7,8,9,10,11,12. The SAVE group routinely invites scientists from international sites to present a real-time assessment of SARS-CoV-2 variants and infections within their region. The SAVE group coordinates with the Biodefense and Emerging Infections (BEI) Research Resources Repository, the World Reference Center for Emerging Viruses and Arboviruses (WRCEVA) and the World Health Organization (WHO) to distribute SARS-CoV-2 isolates, proteins and plasmids. The SAVE group also has an open-face sharing policy in which findings are quickly disseminated through preprint servers while manuscripts undergo formal peer review. The head-to-head comparison, review and discussion of unpublished data has yielded real-time peer review that would otherwise take months to achieve.
The early-detection and analysis group
SARS-CoV-2 genome sequencing data have been shared in public databases. As of December 2021, GISAID—the most widely used database for SARS-CoV-2—has more than 6.5 million sequences deposited with more than 150,000 sequences added weekly. This depth and rate of growth of genetic information for an emerging virus is unprecedented, providing a unique resource to track virus evolution. From late 2020, the emergence of variants of concern (VOCs) with an increased risk to global public health prompted scientists to establish variant detection and tracking pipelines (such as Outbreak.info13, CDC SARS-CoV-2 Variant Classifications and Definitions14 and the BV-BRC SARS-CoV-2 Real-time Tracking and Early Warning System for Variants and Lineages of Concern (https://www.bv-brc.org)). The early-detection and analysis group was assembled to establish a systematic approach to identify and predict SARS-CoV-2 variants that might increase virus replication, transmission and/or escape immunity. The team’s main goal is to select and prioritize variants for development of key experimental reagents (for example, spike proteins for binding assays and pseudoviruses (PSVs) for neutralization assays) and viruses for challenge studies, as well as to inform the in vitro and in vivo groups about predicted variant properties to guide their experiments. The initial and primary focus has been on variants with mutations in the spike protein that might lead to antibody escape, with subsequent analyses considering T cell escape, infectivity and transmission. Other important characteristics—such as replication fitness and virulence—and genomic regions outside of the spike gene are also evaluated. The process is collaborative and iterative, with seven teams using independent models and methodologies to prioritize mutations and lineages as well as rank importance for downstream testing. Although the focus is on human infections, the early-detection group also monitors variants circulating in animal populations, such as mink and deer, as they represent a potential reservoir source.
Methodology
Genomic surveillance consists of weekly downloads of SARS-CoV-2 genomes from GISAID/GENBANK, quality filtering, alignment, and the identification of variant or co-variant substitutions. The main focus has been on potential antibody escape to identify mutations in key epitopes in the receptor-binding domain (RBD) and the N-terminal domain (NTD) supersite, but regions proximal to the furin-cleavage site or experiencing convergent/parallel evolution are also considered. The dynamics of these spike substitutions, as a function of time and geographical spread, are evaluated considering sequence prevalence and viral population growth rate, including comparative analyses to other variants co-circulating in a given geographical location (Fig. 2a). One example of recurrent substitutions with phenotypic relevance are those near to the furin-cleavage site, which result in enhanced spike cleavage and infectivity15,16. These mutations have been identified in different variants and in newly expanding lineages. Some teams take into account vaccine coverage when prioritizing an emerging lineage for analysis.
a, The trajectory of SARS-CoV-2 variant sequence prevalence over a one-year period, 1 January 2021 to 31 December 31 2021, tracking frequencies of weekly counts based on PANGO lineage designations. The data in the graphs are based on the 4.8 million SARS-CoV-2 sequences sampled in 2021 and made available through the GISAID Initiative. Updated graphs can be found online (https://cov.lanl.gov; the tracking tool is called Embers). Global summary and status of five continents. Europe and North America remain the most highly sampled regions of the world, biasing the global sampling. b, Tangle plots for comparative prioritization of circulating variants across subgroups. The list of variants to prioritize was built collectively by the whole group and prioritized by individual teams to arrive at a consensus list. Each column graph refers to the prioritization order made by each subteam for circulating variants in December 2021 (top, highest priority; bottom, lowest priority): A, Cambridge University; B, LANL; C, ISMMS; D, JCVI/BV-BRC; E, UCR SOM; F, Broad Institute; G, WRAIR. The final consensus ranking of the 43 variants was produced by ordering the lineages by their mean rank across the different teams, who also have the option to defer from ranking a lineage or to assign multiple lineages a tied ranking and, after discussion with the group, determine priority categories. The dashed arrow indicates the order of priority. The colours refer to each PANGO lineage tracked, but blocks of the same colour can also refer to different variants within a PANGO lineage. For example, in addition to the coloured Delta AY.* sublineages indicated, Delta has 26 subvariants (purple) with different combinations of mutations that are being prioritized for analysis.
The rankings are split into two broadly distinct methodologies, each with slight variations: one is based on convergent evolution as the main signal for selection and functional impact of mutations (that is, the Cambridge and Walter Reed Army Institute of Research (WRAIR) teams); whereas the other is anchored on prevalence and growth patterns of mutations and defined lineages (that is, the Los Alamos National Lab (LANL), Icahn School of Medicine at Mount Sinai (ISMMS), J. Craig Venter Institute/Bacterial Viral Bioinformatic Resource Center (JCVI/BV-BRC), UC Riverside and Broad Institute teams) (Fig. 2b).
The functional impact of mutations
Cambridge prioritizes substitutions that are likely to cause immune escape by looking at both experimentally determined escape from polyclonal sera and the effect of mutations on spike protein structure. Substitutions are given higher priority if they appear to be emerging and if they are in a different Barnes class17 from previously observed substitutions, and lower priority if they have already been tested experimentally. The WRAIR team tracks the prevalence of substitutions at a set of sites selected based on the strength of the interaction with known SARS-CoV-2 antibodies (using complex structures in the Protein Data Bank; https://www.rcsb.org) as well as structural information or knowledge from deep mutational scanning or mutagenesis studies. Weight scores for ranking are also given for various characteristics, such as the fold increase in detection over time and geographical spread or population growth in the context of high vaccination coverage.
Prevalence and growth patterns
The ISMMS team has a similar approach, whereby variants are ranked on the basis of an aggregate score for sequence prevalence increase and genetic changes of concern in sites of importance associated with functional changes (such as ACE2 binding, antibody escape) but also assigns weight to mutations in the active sites of viral enzymes. Moreover, data from surveillance cohorts in the New York City metropolitan area are used to assess lineages associated with local outbreaks and breakthrough infections after vaccination. LANL identifies emergent mutational patterns within the spike, RBD and NTD supersite to determine global and regional sampling frequencies. Variant dynamics and global spread are tracked at multiple geographical levels using a suite of tools5 (https://cov.lanl.gov/). The JCVI/BV-BRC team uses an algorithm combining sequence prevalence dynamics with functional impact predictions to rank emerging variants. Each mutation is given a sequence-prevalence score, reflecting geographically localized prevalence changes, and a functional impact score, on the basis of the location of the mutation within important spike protein regions and whether studies have demonstrated significant changes in either antibody- or ACE2-receptor binding18,19,20,21. UC Riverside uses relative growth in the prevalence of specific substitutions and deletions/insertions to identify the fastest growing variants and mutation combinations (https://coronavirus3d.org). For the final variant and subvariant ranking, additional criteria are included, such as their potential impact on protein structure (by modelling) and the re-emergence of individual mutations in previously undescribed combinations in new variants. Finally, the team from the Broad Institute, similar to the UC Riverside team, examines the accelerated growth of a variant relative to its peers, across multiple geographical regions, but fits a binomial logistic regression to each lineage’s proportion over time. Moreover, they fit hierarchical multinomial logistic regression models across geographical regions22.
Challenges for the early-detection and analysis group
The early-detection and analysis group has faced six main challenges in identifying emerging variants for functional testing: (1) the newest data are the most subject to bias and the least representative because of small numbers. The longer that one waits, the more accurate the data, but the greater the delay in identifying newly emergent variants for evaluation. (2) Disentanglement of epidemiological from evolutionary effects. A variant might show increased sequence prevalence within a geographical region due to founder effects, or increased incidence could be conferred by epidemiological factors rather than an evolutionary fitness advantage. An example of a founder effect is Delta AY.25, which is very common in North America but not increasing in frequency over time (Fig. 2a), versus AY.4.2, which was first sampled well after Delta was increasing in the UK and was constantly increasing in frequency in 27 countries where it was found and, furthermore, it never significantly decreased relative to other Delta variants once it emerged, suggesting positive selection. (3) Selective pressures on the virus are in flux, and mutations may be transient due to a balance with requirements for retention of fitness. Pressures are exerted by the host at the level of transmission, epidemiological interventions and immune evasion. (4) Under-representation of variant spread and evolution in countries with limited sampling and sequencing capacity. Although some parts of the world have an abundance of sequencing data(such as the UK and USA), others are under-represented (such as the African continent and China). There is an urgent need to increase sampling and sequencing capacity in resource-poor countries. (5) Variability in data quality. The submission of consensus assemblies without underlying raw read-level data means that quality cannot be independently evaluated. Erroneous genome sequences due to technical artifacts, low coverage or bioinformatic strategies that default to ancestral bases in regions without sequence coverage can affect the accuracy of variant amino acid calls23. (6) The database curation quality-control steps can filter on the basis of criteria that do not apply uniformly across lineages. The B.1.621/Mu lineage had an unexpected stop codon in ORF3a that caused B.1.621 sequences to be flagged during automated uploads to the GISAID database, which initially led Mu to be undercounted. This can lead to a false understanding of the dynamics of a given variant lineage globally. Despite these challenges, our prioritization methods continue to evolve as more information becomes available. These efforts have allowed for the rapid generation of reagents for multiple variants before they have spread extensively in the USA and have been critical for guiding the in vitro and in vivo groups. A list of regularly updated prioritized variants is available online (https://docs.google.com/spreadsheets/d/167uJP9LfJN07410sWaMSKU1Se-4XX687j8IgVX4MV_w/edit?usp=sharing).
The in vitro group
The in vitro group performs antibody binding, neutralization, Fc effector and T cell stimulation assays to understand how SARS-CoV-2 variants affect vaccine- and infection-induced immunity. The in vitro group serves as a critical intermediary between the early detection and analysis and in vivo groups by providing valuable data to confirm variant lineage prioritization, and ranking viruses for prioritized in vivo challenge studies. The in vitro group was initially tasked with develo** key reagents (for example, spike and RBD antigens, and plasmids for generating PSVs) and procuring biospecimens (such as authentic viruses and sera/plasma from infected and vaccinated individuals). At the beginning of 2021, reagents for generating data—including variant virus isolates, recombinant infectious clones, recombinant variant spike proteins for antibody binding assays, variant-specific expression plasmids for PSV particle entry inhibition assays and variant-specific sera—were not widely available (Fig. 3a). A key lesson from this process is that the streamlining of administrative procedures for reagent sharing facilitates data generation that directly informs urgent policy- and decision-making. A substantial and ongoing challenge requiring numerous administrative steps is to obtain authentic virus isolates from domestic and international sources. To expedite this process, we developed a pipeline between SAVE investigators to isolate, propagate and sequence emerging viruses. This effort led to cataloguing and isolating hundreds of SARS-CoV-2 variants representing over 40 lineages. For more difficult to obtain SARS-CoV-2, additional efforts have been made to generate infectious clones37. The grid lines represent twofold dilution of antiserum. The y and x axes represent antigenic distance. Circles, antigens; squares, sera. d, T cell responses to SARS-CoV-2 variants. Sequencing data are curated for coding mutations (pink boxes). Curated mutations are tested on convalescent T cell responses using functional assays (activation-induced marker (AIM) assays; green boxes). Immune Epitope Database (IEDB) and the immunocode multiplex identification of T cell receptor antigen specificity dataset (MIRA) are analysed to generate curated peptide sets of immunodominant epitopes (blue boxes). Data are integrated to produce a ranked score list of variant epitope changes weighted by their likelihood to disrupt epitope binding and the relative size of the affected population (grey boxes). MPs, megapools. Partially created using BioRender.
At the start of the pandemic, the correlates of immune protection were unknown for COVID-19. Multiple teams within the in vitro group conducted assessments of vaccine-induced serum neutralization using parallel but independent methods across laboratories. Studies with clinical samples show neutralizing antibody titres are a strong predictor of protection against severe disease27. As such, a major undertaking of the in vitro group has been to use neutralization assays to assess the effect of spike mutations on the inhibitory activity of clinically approved monoclonal antibodies and serum/plasma from vaccinated or infected individuals. One of the strengths of the in vitro group is the use of orthogonal SARS-CoV-2 neutralization assays based on authentic live viruses, PSVs and chimeric viruses. An initial task of this group was to compare neutralization assay platforms across 12 independent laboratories using a defined serum panel from individuals vaccinated with the Pfizer and Moderna vaccines. Using either the ancestral wild-type virus (Wuhan-1) or more recent variants (for example, the Beta variant shown in Fig. 3b), team members performed neutralization assays that varied on the basis of live virus assay readouts (foci, plaques, cytopathic effect, luciferase and fluorescence), target cells, and expression of ACE2 and/or TMPRSS2 on target cells28,29,30,31,32,33,34,35. This type of performance testing has highlighted differences between assay platforms, cell targets and readouts that can impact neutralization potency. Nonetheless, in most cases, there was considerable congruence across platforms. Another area of emphasis is using variant-infected serum/plasma samples to visualize the antigenic evolution of spike through a process called antigenic cartography36,37 (Fig. 3c). This two-dimensional map provides a landscape of how spike mutations drive loss in neutralizing activity.
For many viruses, the affinity and magnitude of antibody binding to viral glycoproteins associates with virus-neutralizing activity, and a strong correlation has been shown for SARS-CoV-238,39,40,41. Investigating the correlation between the neutralizing and binding activity of vaccine-induced antibodies showed that spike mutations alter this slope, and virus neutralization is often more affected than antibody binding42,43. This has been confirmed through different platforms measuring changes in binding to either native spike proteins or the RBD, including ELISA44 and multiplexed spike antigen detection platforms7. One potential explanation for this is that many more binding than neutralizing epitopes exist on the spike protein. Some antibodies that have neutralizing activity against the wild-type virus may lose activity to variants, yet overall binding is still maintained—a phenomenon observed for other viruses (such as influenza virus45).
Binding antibodies can still have a considerable protective effect, irrespective of neutralizing activity due to Fc effector functions, as seen with influenza virus or Ebola virus46,47,48. The humoral immune response restricts microbes through the coordinated effort of the Fab (antigen-binding) and Fc (constant) domains49. After infection or vaccination, polyclonal antibodies are induced that target pathogens at multiple sites through their Fab domains. Fab domains that directly or indirectly hinder virus entry are neutralizing; however, the remaining ‘non-neutralizing’ antibodies can bind to and opsonize the pathogen to form immune complexes, or bind to spike proteins on the surface of infected cells. Once complexed, the Fc domains act as molecular beacons that draw in immune cells through Fc-gamma receptors (FcγRs), providing instructions on how the immune system should destroy the antibody-opsonized material. Fc-effector functions of antibodies are linked to natural resolution of COVID-1950,51,52,53, correlate with vaccine-mediated protection from infection in animal models54,55,56 and are associated with protection after the transfer of passive convalescent serum or monoclonal antibodies57,58,59,60. Although emerging variants of SARS-CoV-2 can escape neutralizing antibodies, their substitutions alter a limited fraction of the overall humoral immune response to the SARS-CoV-2 spike56,61. Thus, Fc-effector functions have more resilience in the face of variation across spike, for both mRNA and the adenoviral 26 (Ad26) vaccines, offering mechanisms through which antibodies may continue to confer protection despite esca** neutralization.
Growing evidence from animal models and human studies indicates that CD4+ and CD8+ T cells have protective roles in preventing severe disease and death from SARS-CoV-2 infection6,62,63,64. T cells are an attractive target for intervention as they are less susceptible to viral escape than antibodies6,65. This is largely for two reasons: (1) in convalescent individuals, T cells can target peptides derived from the entire proteome, not just surface-exposed epitopes; and (2) HLA-restriction and diversity creates interpersonal variation in the repertoire of targets, limiting the immunological pressure on any one epitope. Given the presumed role of T cells in limiting severe disease and their potential for sustaining protection against variant mutation, the SAVE in vitro group included assessment of T cell responses. The goal was to determine empirical drift from vaccination and infection-induced immunity, and to develop tools to predict the impact of variant-associated mutations on immunodominant T cell responses.
The T cell investigations follow two parallel approaches to assess the impact of variant mutations on T cell reactivity and a broad range of different variants (Fig. 3d). The first involves measuring the overall reactivity against the entire spike protein (in the case of vaccination) or the entire proteome (in the case of infection) and expressing the results as the fold difference relative to the ancestral sequences. A parallel approach characterizes the mutational impact on specific single epitopes, and monitors whether individuals with decreased T cell reactivity have responses that selectively recognize certain epitopes in the context of particular HLA types. Regarding the first approach, at the general population level, the results to date have detected a limited impact of mutations within spike after natural infection or mRNA vaccination6 against the most concerning variants at the time the study was performed (B.1.1.7, B.1.351, P.1 and B.1.427/429). These findings were corroborated66 and expanded to adenoviral-vector-based vaccination67. However, in a minority of individuals, two- to threefold decreases in the CD8+ T cell responses against the B.1.351/Beta and B.1.427/429/Epsilon variants were noted6. These findings suggest that a more in-depth characterization at the single-epitope level is required to understand the mechanisms behind the reduced CD8+ T cell response in specific individuals. Moreover, it is critical to monitor and predict the effect of emerging circulating variants on T cell reactivity, particularly regarding the most concerning (to date) B.1.617.2/Delta variant (including the AY.* sublineages) and B.1.1.529/Omicron variant. The experimental data will be used to confirm and improve the bioinformatics analysis and infer the impact of current and upcoming variants on SARS-CoV-2 specific T cell responses.
Advanced computational tools for assessing SARS-CoV-2 genome mutations on HLA binding have enabled prediction of the effect of mutations within a VOC on T cell reactivity. Owing to the broad diversity of HLA genotypes, T cell escape at the population level is not likely, as demonstrated for multiple VOCs6. However, previous work on HIV and influenza virus has identified associations between specific HLA class I alleles, disease severity68,69 and vaccine efficacy70. We anticipate that, as SARS-CoV-2 continues to spread globally, T cell immunity will eventually drive viral evolution. In these situations, specific HLA alleles may become associated with a reduced ability to mount responses against dominant T cell epitopes, which may affect clinical outcomes. The T cell subgroup has developed a computational pipeline to assess the effects of specific mutations on HLA binding by also ranking all individual mutations on a T cell escape score, based on experimentally verified and predicted T cell responses (Fig. 3d). This ranking will provide early identification of specific mutations associated with T cell escape, particularly CD8+ T cells, and testable hypotheses for T cell experiments. In our preliminary analyses of VOCs, the B.1.617.2 variant was identified as the first in which mutations were associated with reduced HLA binding at the population level. These data suggest that T cell cross-reactivity to B.1.617.2 may be reduced in some individuals. Owing to the extensive number of SARS-CoV-2 viral genomes, and large-scale clinical cohorts that are being studied, the T cell SAVE group plans to assemble a database linking HLA genotypes with clinical outcome and viral genomes, which may provide a unique opportunity to study HLA associations with clinical disease and viral evolution at a resolution that has not previously been attempted.