Abstract
Fructosamine-3-kinases (FN3Ks) are a conserved family of repair enzymes that phosphorylate reactive sugars attached to lysine residues in peptides and proteins. Although FN3Ks are present across the Tree of Life and share detectable sequence similarity to eukaryotic protein kinases, the biological processes regulated by these kinases are largely unknown. To address this knowledge gap, we leveraged the FN3K CRISPR Knock-Out (KO) HepG2 cell line alongside an integrative multi-omics study combining transcriptomics, metabolomics, and interactomics to place these enzymes in a pathway context. The integrative analyses revealed the enrichment of pathways related to oxidative stress response, lipid biosynthesis (cholesterol and fatty acids), and carbon and co-factor metabolism. Moreover, enrichment of nicotinamide adenine dinucleotide (NAD) binding proteins and localization of human FN3K (HsFN3K) to mitochondria suggests potential links between FN3K and NAD-mediated energy metabolism and redox balance. We report specific binding of HsFN3K to NAD compounds in a metal and concentration-dependent manner and provide insight into their binding mode using modeling and experimental site-directed mutagenesis. Our studies provide a framework for targeting these understudied kinases in diabetic complications and metabolic disorders where redox balance and NAD-dependent metabolic processes are altered.
Similar content being viewed by others
Introduction
Reducing sugars including ribose and glucose can spontaneously react with free amines on peptides, proteins, and lipids through a process called glycation1. This process can occur either endogenously or exogenously via the foods we consume2. Because increased glycation can negatively impact biomolecular functions through the formation of stable adducts, known as ketosamines3, organisms have evolved enzymatic repair systems as a defense mechanism. Fructosamine-3 kinases (FN3Ks) are one such family of repair enzymes that use Adenosine triphosphate (ATP) to de-glycate lysine residues in proteins4,5,6. They do so by catalyzing the transfer of the gamma phosphate from ATP to the 3’ hydroxyl group in the ketosamine substrate, which rapidly breaks down into inorganic phosphate and 3-deoxyglucosone (3-DG), regenerating the original lysine6. There is emerging evidence that altered FN3K expression and increased ketosamine levels contribute to metabolic disorders7,8 through the production of Advanced Glycation End (AGEs) products9 and reactive oxygen species (ROS)10.
Deglycation by FN3Ks is an ancient mechanism for protein repair as FN3K homologs are found throughout the tree of life11,12. While lower eukaryotes and prokaryotes possess a single copy of the FN3K gene, most tetrapod genomes contain two: FN3K and FN3K Related Protein (FN3KRP). Interestingly, two independent gene duplication events have led to the tetrapods having two copies: one in reptiles/birds and the other in placental mammals13. It is believed that in lower organisms, FN3K orthologs repair proteins glycated by ribose-5-phosphate, a potent glycating metabolite formed from the conserved pentose phosphate pathway11,12. Despite the remarkable conservation of FN3Ks across diverse organisms and their emerging role in protein repair, the biological and cellular processes linked to this important class of proteins remain unknown.
FN3K activity is highly substrate-specific with human FN3K (HsFN3K) phosphorylating ketosamines resulting from glycation of both L and D orientation sugars, whereas FN3KRP orthologs are limited to only D-orientation sugars11,12,14. At the transcriptional level, in human tissues, HsFN3K is highly expressed in the brain, kidney, liver, heart muscle and adrenal gland (Fig. 1a) whereas HsFN3KRP expression levels are somewhat uniform throughout the different tissues (Fig. 1a). The paralogs are also reported to be localized to distinct subcellular compartments. Based on immunohistochemistry (IHC) studies on HsFN3K in HepG2 cells, HsFN3K is reported to be localized in mitochondria whereas HsFN3KRP is localized in nucleoplasm in different cell lines15.
a Tissue-specific expression levels of human FN3K (HsFN3K) and human FN3KRP (HsFN3KRP), based on data from the Protein Atlas Database. b The levels of HsFN3K RNA expression across various types of cancer, arranged by median expression values. The figure was produced using cBioPortal57. Unit for mRNA expression levels: RSEM (Batch normalized from Illumina HiSeq_RNASeqV2) (log2(value + 1)). c Western blot analysis displaying the total quantities of HsFN3K and HsFN3KRP in both wild-type and FN3K knockout (KO) HepG2 cells. The blots were from separate gels.
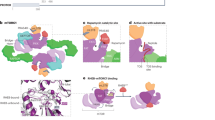
Previously, we reported the crystal structure of FN3K ortholog from Arabidopsis thaliana (AtFN3K), revealing an evolutionarily conserved redox regulation mechanism involving conserved cysteines in the ATP binding P-loop16. Specifically, we demonstrated that AtFN3K adopts a disulfide-linked dimer that can be reversed under reducing conditions and that the equivalent cysteine in HsFN3K (C24HsFN3K) performs an analogous role conferring redox sensitivity and disulfide-mediated oligomerization. Furthermore, a recent study has proposed a potential link between HsFN3K deglycation function and the development of hepatocellular carcinoma (liver cancer) through the involvement of the nuclear transcription factor Nuclear factor erythroid 2-related factor 2 (NRF2)17, which controls the expression of multiple antioxidant enzymes18. Acting as an adaptor protein, Kelch-like ECH-associated protein (KEAP1) interacts and directly inhibits Nuclear factor erythroid 2-related factor 2 (NRF2) by facilitating its ubiquitination through Cullin3 (CUL3) and marking it for proteasomal degradation19. KEAP1 has redox-sensitive cysteines that undergo modifications during oxidative stress, impeding the interaction between KEAP1 and NRF2. The progressive buildup of NRF2 triggers the induction of antioxidant enzymes, further underscoring the role of HsFN3K in maintaining cellular redox balance and altered expression in liver cancer (Fig. 1b). It is worth noting that higher HsFN3K expression levels are also observed in ocular/eye and adrenal cancers (Fig. 1b). However, despite the altered expression of HsFN3K in multiple cancer types and their established roles in metabolic diseases such as diabetes and associated complications20,21, our understanding of FN3K’s functions within human cells and across different species, particularly in the context of redox regulation, remains incomplete.
Here we leverage the CRISPR Knock Out (KO) of HsFN3K in HepG2 liver cancer cell line (Fig. 1c, Supplementary Fig. 1) and employ a multi-omics approach integrating transcriptomics, metabolomics, and interactomics to illuminate the cellular pathways associated with HsFN3K. Notably, knocking out HsFN3K did not change protein levels for the paralog HsFN3KRP (Fig. 1c). Completely knocking out HsFN3K in HepG2 cells resulted in upregulation of lipid biosynthesis (fatty acid and cholesterol) and oxidative stress response pathways. Joint enrichment analysis of the transcriptomics and the metabolomics datasets revealed enrichment of pathways related to glutathione and co-factor metabolisms, including Coenzyme A (CoA) and nicotinate/nicotinamide metabolism. Interestingly, several of the HsFN3K interacting partners were involved in fatty acid and pyruvate metabolism pathways, which we further validated using immunoprecipitation in HepG2 cells. We found that HsFN3K interacts with Fatty acid synthase (FASN) and Lactate dehydrogenase A (LDHA) in cytoplasm. Moreover, by performing enrichment analysis on the integrative network combining transcriptomics and interactomics, we identified enrichment of Nicotinamide dinucleotide (NAD) binding superfamily and thiolase domains. Finally, we validated HsFN3K binding to NAD compounds using Differential Scanning Fluorimetry (DSF). We further probed the binding specificity through mutagenesis and identified key residues in the adenosine as well as substrate binding pockets, revealing that HsFN3K is inhibited by NADH.
Results
To investigate the molecular pathways regulated by HsFN3K, we utilized an integrative multi-omics approach. By leveraging the FN3K Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) Knock-Out (KO) HepG2 cell lines, we identified genes, metabolites, and pathways altered by HsFN3K knock out. Enrichment analyses on individual datasets as well as combined datasets (metabolomics and transcriptomics) (Fig. 2a, b) highlighted key pathways (Supplementary Fig. 2a) placing these enzymes in a biological context.
Transcriptomic profiling and pathway analysis of HsFN3K KO cells reveal alterations in metallothionein binding, lipid metabolism, and oxidative stress response pathways
For the transcriptomics, we conducted RNA sequencing on total RNA from the KO and the wild type (WT) cells. The obtained reads were analyzed using the HISAT2-StringTie-Ballgown22 suite of tools where the reads are first mapped against the human reference using HISAT223, followed by transcript assembly and quantification using StringTie24. Finally, based on these quantified transcripts in both the KO and WT cell lines, Ballgown25 was used to perform a differential expression analysis. Overall, we identified 408 differentially expressed genes (DEGs) (greater than 2 log-fold change and p-value < 0.05) in the CRISPR KO cells relative to WT cells) (Fig. 3a). Out of the 408 genes, 305 genes were upregulated. The DEGs also include 23 genes that are RNA genes or pseudogenes (Supplementary Dataset File 1).
The most upregulated genes include metallothioneins (MT) paralogs such as MT1E (11-fold) and MT1G (15-fold) and members of the Cytochrome P450 family such as CYP24A1 (10-fold) and CYP17A1 (8.5-fold) (Fig. 3a). Moreover, genes in cholesterol synthesis and homeostasis: Proprotein convertase subtilisin/kexin type 9 (PCSK9) (10.5-fold), methylsterol monooxygenase 1 (MSMO1) (10-fold), mevalonate diphosphate decarboxylase (MVD) (10-fold), mevalonate kinase (MVK) (8-fold), and 3-hydroxy-3-methylglutaryl-CoA synthase 1 (HMGCS1) (9.5-fold) were also highly upregulated (Fig. 3a).
The most downregulated genes include the kelch domain containing 7B (KLHDC7B) (9.2-fold) and its long noncoding RNA counterpart CTA-384D8.31 (10.7-fold) (Fig. 3a). Other RNA genes such as growth arrest specific 5 (GAS5) (4.2-fold) and small nucleolar RNA host gene 22 (SNHG22) (4.1-fold) were also downregulated (Fig. 3a). Moreover, there was down regulation of genes involved in cell growth secreted protein acidic and cysteine rich (SPARC) (4.3-fold) and in cell adhesion nephronectin (NPNT) (8.9-fold) in the KO (Fig. 3a). Interestingly, S-adenosylmethionine (SAM) synthesizing gene methionine adenosyltransferase 2 A (MAT2A) (4.3-fold) was among the top 10 most down-regulated genes (Fig. 3a).
After identifying differentially expressed genes, we performed enrichment analysis at the pathway level, prioritizing them according to their fold enrichment. This approach enabled us to effectively quantify the overrepresentation of specific pathways by comparing the total number of genes identified in each pathway against the background frequency of genes annotated for that pathway26. Notably, the pathway involving “Metallothioneins bind metals” emerged as the most significantly enriched pathway, as determined by fold enrichment (Fig. 3b). Metallothioneins are low molecular weight (6–7 kDa) cysteine-rich proteins that play crucial roles in metal homeostasis and the oxidative stress response pathway, providing protection against DNA damage and cytotoxicity27.
Following this, we observed significant enrichment in pathways associated with lipid biosynthesis and signaling, particularly in cholesterol and fatty acid metabolism (Fig. 3b). Other notable pathways were related to “Oxidative stress response” and related pathways such as “Binding and Uptake of Ligands by Scavenger Receptors”, “FOXO-mediated transcription of oxidative stress, metabolic, and neuronal genes”, “Glutathione metabolism” as well as “Ferroptosis”. Finally, “Vitamins and co-factor metabolism” as well as related pathways such as “Folate metabolism” and “Porphyrin and chlorophyll metabolism” were also enriched (Fig. 3b).
Joint pathway analysis using transcriptomics and metabolomics datasets reveal enrichment of glutathione, carbon, and co-factor metabolisms including nicotinate/nicotinamide metabolism
We previously identified differentially abundant metabolites by comparing the metabolic profiles of FN3K KO and WT HepG2 cells16. Metabolite set enrichment analysis (MSEA) of the differentially abundant metabolites revealed an enrichment in several pathways related to amino acid metabolism, in addition to pathways such as “Glutathione metabolism”, “Pyruvate metabolism”, and “Pantothenate and CoA Biosynthesis” (Supplementary Fig. 2b). Integrating this with our transcriptomics data, we conducted a joint pathway analysis (Fig. 2a) using MetaboAnalyst23 was first used to map the reads to the human reference genome separately for the KO and WT samples. This was performed using the default parameters. Then, the alignment was passed to StringTie24 for assembly and quantification of the transcripts. The assembled transcripts were then passed to Ballgown25 to perform statistical tests to identify differentially expressed transcripts between the WT and KO samples. We used Ballgown’s stattest function to perform a statistical comparison of the FPKM values for all assembled transcripts at the gene level and report the p-value and the fold change for genes that were significantly differentially expressed. Only genes with p-value < 0.05 and a log fold change > 0.2 were considered as significantly differentially expressed and used in subsequent analyses.
Pathway enrichment analysis
We collected the UniProt IDs for i) all the genes that were found to be differentially expressed in the RNA-Seq analysis and ii) the genes that were identified as interacting partners using IP. This list was uploaded individually or merged together into Cytoscape50 and the STRING Cytoscape plugin51 was used to build a STRING-based network of all these genes. We then used the STRING plugin to perform an enrichment analysis on this network of genes to find enriched pathways. The enrichment analyses on transcriptomics and interactomics datasets was performed using stringApp which uses KEGG52, Reactome53, and Wiki pathways54. We used a redundancy cutoff of 0.2 within STRING to remove pathways that had a high overlap of genes. We further filtered this list to remove small pathways that had 5 or less genes with an FDR value below 0.05 (Supplementary Dataset File 1). For metabolomics, we used Metaboanalyst which uses KEGG and Human Metabolome database.
We used fold enrichment, a quantitative measure of the over-representation of a pathway, to rank order the obtained pathways. Fold enrichment is obtained by comparing the background frequency of total genes annotated for that pathway on humans to the actual genes that mapped to that same pathway from the list that was provided. Specifically, fold enrichment for each pathway was calculated using the following proportion:
where,
-
n is the total number of genes from our list that match the pathway.
-
M is the total number of genes in our list.
-
Nb is the total number of background genes (total genes in that pathway) in humans.
-
Nt is the total number of genes in humans (19566, for human genome used by the STRING database version 11.5).
Differential Scanning Fluorimetry (DSF)
Approximately 7 µM of Human FN3K WT or mutants (C24A, F39V, D234A, W219) were mixed with 1:500 SYPRO ORANGE (Sigma) in buffer (20 mM HEPES pH 7.4, 150 mM NaCl, 5% glycerol). The nucleotides or nicotinamides were added in the presence or absence of 6 mM MgCl2 to make the final concentration of 5 mM. The solution was then heated gradually from 20 °C to 95 °C at the rate of 0.3 °C, and the fluorescence was monitored using the StepOne Plus Real-time PCR instrument. Data was analyzed using Prism.
Homology modeling, docking and ConSurf Analysis
The crystal structure of the plant homolog from A. thaliana (PDB ID: 6OID) was used as a template for the homology model of HsFN3K. ADP was modeled in the pocket by aligning the crystal structure with the homology model. Magnesium ion was docked on the homology model using the Metal-Ion Binding (MIB)38 webserver. NADH was docked onto the homology model using scaffold docking with ADP as a template on the Acedock37 webserver. Chain A of the homology model was used as a query for the ConSurf55 server.
Subcellular Fractionation, immunoprecipitation (IP) and Western Blot
HepG2 cells were cultured on a 10 cm dish until they reached 80–90% confluency. Subcellular fractionation was conducted following a protocol from “Subcellular Fractionation: A Laboratory Manual”56. Briefly, cells from a 10 cm dish were lysed in 550 μL ice-cold RSB buffer (10 mM Tris-HCl, pH 7.5, 10 mM NaCl, 1.5 mM MgCl2) for 10 minutes and subjected to dounce homogenization. The lysed cells were then transferred to a microcentrifuge tube, and 400 µL of 2.5x homogenization buffer (12.5 mM Tris-HCl, pH 7.5, 525 mM mannitol, 175 mM sucrose, 2.5 mM EDTA) was added. The mixture was centrifuged at 1300 g to pellet nuclei (the nuclear fraction), which was then washed with 1x homogenization buffer (5 mM Tris-HCl, pH 7.5, 210 mM mannitol, 70 mM sucrose, 1 mM EDTA) to remove contaminants. The supernatant was transferred to a new tube and subjected to a repeat 1300 g centrifugation to ensure all nuclei were removed. The final supernatant was then transferred to a new tube and centrifuged at 17000 g for 15 minutes to pellet the mitochondria (the mitochondrial fraction was washed with 1x homogenization buffer to eliminate contaminants). The remaining supernatant was designated as the cytosolic fraction and centrifuged at 17000 g to clear mitochondrial debris. Nuclear and mitochondrial fractions were resuspended in 1 mL RIPA buffer (50 mM Tris-HCl, pH 7.4, 150 mM NaCl, 1% NP-40, 0.1% SDS, 0.5% deoxycholate) and sonicated, while the cytosolic fraction was dialyzed in RIPA buffer using Amicon centrifugal filters. Protein concentrations in each fraction were determined using a BioTek Synergy plate reader, all the subcellular fractions were normalized to 1 mg/ml total proteins. Equal amounts of protein (400ug) were subjected to immunoprecipitation with anti-FN3K antibody (1:200 dilution) (Invitrogen, Catalog #: PA5–28603) and 20 µL agarose protein A/G plus beads (Pierce). After 16 hours of incubation at 4 °C, the beads were washed three times in RIPA buffer, and the bound proteins were eluted in 1x Laemmli buffer and resolved on 10% SDS-PAGE gels. Western blotting was carried out by transferring proteins onto a PVDF membrane using a semi-dry transfer system. Interacting proteins were detected using antibodies against fatty acid synthase (1:2000 dilution) (Invitrogen, Catalog #: MA-531490) and LDHA (1:1000 dilution) (Cell Signaling Technology, Catalog #: C4B5). Antibodies against Vinculin (1:1000 dilution) (Cell Signaling Technology, Catalog #: E1E9V), Histone H3 (1:5000 dilution) (Cell Signaling Technology, Catalog #: D1H2), and SDHA (1:1000 dilution) (Cell Signaling Technology, Catalog #: 5839 S) served as subcellular markers. Detection of bound antibodies was performed (chemiluminescence) using the LI-COR Odyssey M imaging system.
Immunofluorescence
HepG2 WT and KO cells were fixed with 4% p-formaldehyde for 30 min and washed three times with 1XPBS followed by blocking with 3% normal goat serum for 1 h. Cells were incubated with 1ug/ml FN3K (1:50 dilution) (Invitrogen, PA5–66239) and Tomm20 (1:100 dilution) (Abnova, H00009804-M01) antibodies at 4 °C for 24 h. Cells were washed three times with 1XPBS and incubated with secondary antibodies, Alexa Fluor rabbit 488 (rabbit-FN3K, green) and Alexa Fluor mouse 555 (mouse-Tomm20, red) for 24 h. Secondary antibodies were removed by washing three times with 1XPBS and nuclear staining was performed with 1ug/ml DAPI (blue) for 5 min. Cells were given final wash with 1XPBS followed by visualizing under Zeiss Observer.D1 microscope for imaging.
Interaction of LDHA with Purified 6X-Histidine Tag HsFN3K
25 µL of 50% Co2+ Talon beads were prepared by washing three times with Phosphate-Buffered Saline (PBS). To these prepared beads, 400 µL of the HepG2 cytosolic fraction (from 1 mg/ml total proteins) was added, followed by the addition of 45 µL of 1 mg/mL His-tagged (HsFN3K) (wild-type and mutants as indicated). After 16 hours of incubation at 4 °C, the beads were washed three times in RIPA buffer. Bound proteins were eluted in 1x Laemmli buffer and resolved on 10% SDS-PAGE gels. Western blotting was performed as described above using anti-LDHA (1:1000 dilution) (Cell Signaling Technology, Catalog #: C4B5) and anti-His Tag (1:6000 dilution) (Cell Signaling Technology, Catalog #: 27E8) antibodies.
NADH inhibition assay
A mixture containing 10.0 µg (5 µL) of HsFN3K WT and its mutants, along with 50 mM Ribulose-N-α-Ac-lysine (5 µL) to reach a final concentration of 5 mM, was prepared. This mixture was then combined with 20 µL of a specially prepared solution comprising 150 µL of 5X Kinase Buffer (containing 160 mM HEPES at pH 7.4, 80 mM MgCl2, 1.2 M NaCl, and 40% v/v Glycerol), 15 µL of 250 mM Phosphoenolpyruvic acid (PEP), 45 µL of a PK/LDH mixture [ranging from 600 to 1000 units/mL for pyruvate kinase (PK) and 900 to 1400 units/mL for lactic dehydrogenase (LDH)], and 90 µL of H2O. To test inhibition, 10 µL of NADH stock at varying concentrations was added. The reaction was initiated with the addition of 10 µL of 2.5 mM ATP, achieving a final concentration of 0.5 mM ATP. The total volume for each reaction well was brought to 50 µL. Subsequently, the 96-well plate was placed into a Biotek Synergy H4 plate reader, where the absorbance at 340 nm was continuously monitored at 35 °C for two and a half hours. For storage, proteins were kept in Buffer D (comprising 25 mM HEPES, pH 7.4, 300 mM NaCl, and 10% v/v Glycerol), which was also employed as the mock buffer when necessary.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All relevant data supporting the findings of this study are available within the article (and its supplementary information files). RNA expression data for the WT and the FN3K KO samples have been deposited in the Gene Expression Omnibus database under the accession code GSE242555.
References
Gkogkolou, P. & Böhm, M. Advanced glycation end products. Key players in skin aging? Derm.-Endocrinol. 4, 259–270 (2012).
Uribarri, J. et al. Dietary advanced glycation end products and their role in health and disease. Adv. Nutr. 6, 461–473 (2015).
Baynes, J. W. et al. The Amadori product on protein: structure and reactions. Prog. Clin. Biol. Res. 304, 43–67 (1989).
Bommer, G. T., Van Schaftingen, E. & Veiga-da-Cunha, M. Metabolite repair enzymes control metabolic damage in glycolysis. Trends Biochem Sci. 45, 228–243 (2020).
Collard, F., Delpierre, G., Stroobant, V., Matthijs, G. & Van Schaftingen, E. A Mammalian Protein Homologous to Fructosamine-3-Kinase Acting on Psicosamines and Ribulosamines but not on Fructosamines. Diabetes 52, 2888–2895 (2003).
Szwergold, B. S., Howell, S. & Beisswenger, P. J. Human fructosamine-3-kinase: purification, sequencing, substrate specificity, and evidence of activity in vivo. Diabetes 50, 2139–2147 (2001).
Alderawi, A. et al. FN3K expression in COPD: a potential comorbidity factor for cardiovascular disease. BMJ Open Respir. Res. 7, https://doi.org/10.1136/bmjresp-2020-000714 (2020).
Dunmore, S. J. et al. Evidence that differences in fructosamine-3-kinase activity may be associated with the glycation gap in human diabetes. Diabetes 67, 131–136 (2018).
Chen, J. H., Lin, X., Bu, C. & Zhang, X. Role of advanced glycation end products in mobility and considerations in possible dietary and nutritional intervention strategies. Nutr. Metab. (Lond.) 15, 72 (2018).
Cepas, V., Collino, M., Mayo, J. C. & Sainz, R. M. Redox Signaling and Advanced Glycation Endproducts (AGEs) in Diet-Related Diseases. Antioxidants (Basel) 9, https://doi.org/10.3390/antiox9020142 (2020).
Fortpied, J., Gemayel, R., Stroobant, V. & van Schaftingen, E. Plant ribulosamine/erythrulosamine 3-kinase, a putative protein-repair enzyme. Biochem. J. 388, 795–802 (2005).
Gemayel, R. et al. Many fructosamine 3kinase homologues in bacteria are ribulosamine/erythrulosamine 3-kinase potentially involved in protein deglycation. FEBS J. 274, 4360–4374 (2007).
Cunningham, F. et al. Ensembl 2022. Nucleic Acids Res. 50, D988–d995 (2022).
Collard, F. et al. Fructosamine 3-kinase-related protein and deglycation in human erythrocytes. Biochem. J. 382, 137–143 (2004).
Thul, P. J. et al. A subcellular map of the human proteome. Science 356, eaal3321 (2017).
Shrestha, S. et al. A redox-active switch in fructosamine-3-kinases expands the regulatory repertoire of the protein kinase superfamily. Sci Signal 13, https://doi.org/10.1126/scisignal.aax6313 (2020).
Sanghvi, V. R. et al. The Oncogenic Action of NRF2 Depends on De-glycation by Fructosamine-3-Kinase. Cell 178, 807–819.e821 (2019).
Rushmore, T. H., Morton, M. R. & Pickett, C. B. The antioxidant responsive element. Activation by oxidative stress and identification of the DNA consensus sequence required for functional activity. J. Biol. Chem. 266, 11632–11639 (1991).
McMahon, M., Itoh, K., Yamamoto, M. & Hayes, J. D. Keap1-dependent proteasomal degradation of transcription factor Nrf2 contributes to the negative regulation of antioxidant response element-driven gene expression. J. Biol. Chem. 278, 21592–21600 (2003).
Avemaria, F. et al. Possible role of fructosamine 3-kinase genoty** fot the management of diabetic patients. Clin. Chem. Lab. Med. 53, 1315–1320 (2015).
Avemaria, F. et al. Genetic variability of the fructosamine-3-kinase gene in diabetic patients. Clin. Chem. Lab. Med. 49, 803–808 (2011).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genoty** with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295, https://doi.org/10.1038/nbt.3122.
Frazee, A. C. et al. Ballgown bridges the gap between transcriptome assembly and expression analysis. Nat. Biotechnol. 33, 243–246, https://doi.org/10.1038/nbt.3172.
Dalmer, T. R. A. & Clugston, R. D. Gene ontology enrichment analysis of congenital diaphragmatic hernia-associated genes. Pediatric Res. 85, 13–19, https://doi.org/10.1038/s41390-018-0192-8.
Si, M. & Lang, J. The roles of metallothioneins in carcinogenesis. J. Hematol. Onco.y 11, 107, https://doi.org/10.1186/s13045-018-0645-x.
Chong, J., Wishart, D. S. & **a, J. Using MetaboAnalyst 4.0 for Comprehensive and Integrative Metabolomics Data Analysis. Curr. Protocols Bioinform. 68, https://doi.org/10.1002/cpbi.86 (2019).
Utriainen, M. & Morris, J. H. clusterMaker2: a major update to clusterMaker, a multi-algorithm clustering app for Cytoscape. BMC Bioinforma. 24, 134 (2023).
Ellgaard, L. & Ruddock, L. W. The human protein disulphide isomerase family: substrate interactions and functional properties. EMBO Rep. 6, 28–32 (2005).
Jensen-Urstad, A. P. & Semenkovich, C. F. Fatty acid synthase and liver triglyceride metabolism: housekeeper or messenger? Biochimica et. biophysica acta 1821, 747–753 (2012).
Quan, J., Bode, A. M. & Luo, X. ACSL family: The regulatory mechanisms and therapeutic implications in cancer. Eur. J. Pharmacol. 909, 174397 (2021).
Valvona, C. J., Fillmore, H. L., Nunn, P. B. & Pilkington, G. J. The Regulation and Function of Lactate Dehydrogenase A: Therapeutic Potential in Brain Tumor. Brain Pathol. 26, 3–17 (2016).
Tang, Y. et al. Overexpression of PCK1 gene antagonizes hepatocellular carcinoma through the activation of gluconeogenesis and suppression of glycolysis pathways. Cell Physiol. Biochem. 47, 344–355 (2018).
**ao, W., Wang, R. S., Handy, D. E. & Loscalzo, J. NAD(H) and NADP(H) Redox Couples and Cellular Energy Metabolism. Antioxid. Redox Signal 28, 251–272 (2018).
Harijan, R. K., Dalwani, S., Kiema, T. R., Venkatesan, R. & Wierenga, R. K. Thiolase: a versatile biocatalyst employing coenzyme A-thioester chemistry for making and breaking C-C bonds. Annu Rev. Biochem. 92, 351–384 (2023).
Ruiz-Carmona, S. et al. rDock: a fast, versatile and open source program for docking ligands to proteins and nucleic acids. PLoS Comput. Biol. 10, e1003571 (2014).
Lu, C. H. et al. MIB2: metal ion-binding site prediction and modeling server. Bioinformatics 38, 4428–4429 (2022).
Šileikytė, J., Sundalam, S., David, L. L. & Cohen, M. S. Chemical Proteomics Approach for Profiling the NAD Interactome. J. Am. Chem. Soc. 143, 6787–6791 (2021).
Uhlén, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Rafaeloff-Phail, R. et al. Biochemical regulation of mammalian AMP-activated protein kinase activity by NAD and NADH. J. Biol. Chem. 279, 52934–52939 (2004).
Del Toro, N. et al. The IntAct database: efficient access to fine-grained molecular interaction data. Nucleic Acids Res. 50, D648–d653 (2022).
Oughtred, R. et al. The BioGRID database: A comprehensive biomedical resource of curated protein, genetic, and chemical interactions. Protein Sci. 30, 187–200 (2021).
Berthiaume, J. M., Kurdys, J. G., Muntean, D. M. & Rosca, M. G. Mitochondrial NAD(+)/NADH Redox State and Diabetic Cardiomyopathy. Antioxid. Redox Signal 30, 375–398 (2019).
Calabrese, G., Morgan, B. & Riemer, J. Mitochondrial glutathione: regulation and functions. Antioxid. Redox Signal 27, 1162–1177 (2017).
Chung, H. S., Wang, S. B., Venkatraman, V., Murray, C. I. & Van Eyk, J. E. Cysteine oxidative posttranslational modifications: emerging regulation in the cardiovascular system. Circ. Res. 112, 382–392 (2013).
**ao, H. et al. A quantitative tissue-specific landscape of protein redox regulation during aging. Cell 180, 968–983.e924 (2020).
Alderawi, A. et al. FN3K expression in COPD: a potential comorbidity factor for cardiovascular disease. https://doi.org/10.1136/bmjresp-2020-000714.
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. A.-O. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown.
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Doncheva, N. T., Morris, J. H., Gorodkin, J. & Jensen, L. J. Cytoscape StringApp: Network Analysis and Visualization of Proteomics Data. J. Proteome Res. 18, 623–632 (2019).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Gillespie, M. et al. The reactome pathway knowledgebase 2022. Nucleic Acids Res. 50, D687–d692 (2022).
Martens, M. et al. WikiPathways: connecting communities. Nucleic Acids Res. 49, D613–d621 (2021).
Ashkenazy, H. et al. ConSurf 2016: an improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Res. 44, W344–350, (2016).
Pryor, P. R. Subcellular fractionation : a laboratory manual. (Cold Spring Harbor Laboratory Press, 2015).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Kutmon, M. et al. PathVisio 3: an extendable pathway analysis toolbox. PLoS Comput. Biol. 11, e1004085 (2015).
Acknowledgements
Funding for NK from NIGMS (R35 GM139656) is acknowledged. We thank members of the NK lab and Dr. Dominic Byrne from the University of Liverpool for valuable feedback. NIH funding (S10RR028859) to Professor IJ Amster for purchasing Orbitrap for mass spec analysis is acknowledged.
Author information
Authors and Affiliations
Contributions
N.K. conceptualized the research and secured funding. N.K. and S.S. supervised the overall project. S.S. and S.K. conducted the experiments and data collection. S.S. and R.T performed data and statistical analysis. N.K., S.S., S.K., and R.T. drafted and revised manuscripts. All authors discussed the results, reviewed the manuscript, and approved the final version to be published.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shrestha, S., Taujale, R., Katiyar, S. et al. Multi-omics reveals new links between Fructosamine-3-Kinase (FN3K) and core metabolic pathways. npj Syst Biol Appl 10, 64 (2024). https://doi.org/10.1038/s41540-024-00390-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41540-024-00390-0
- Springer Nature Limited