Abstract
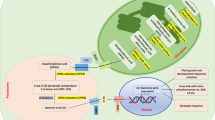
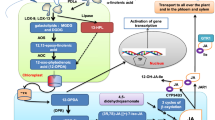
Coumarins, also known as 1,2-benzopyrones, comprise a large class of secondary metabolites that are ubiquitously found throughout the plant kingdom. In many plant species, coumarins are particularly important for iron acquisition and plant defence. Here, we show that COUMARIN SYNTHASE (COSY) is a key enzyme in the biosynthesis of coumarins. Arabidopsis thaliana cosy mutants have strongly reduced levels of coumarin and accumulate o-hydroxyphenylpropanoids instead. Accordingly, cosy mutants have reduced iron content and show growth defects when grown under conditions in which there is a limited availability of iron. Recombinant COSY is able to produce umbelliferone, esculetin and scopoletin from their respective o-hydroxycinnamoyl-CoA thioesters by two reaction steps—a trans–cis isomerization followed by a lactonization. This conversion happens partially spontaneously and is catalysed by light, which explains why the need for an enzyme for this conversion has been overlooked. The combined results show that COSY has an essential function in the biosynthesis of coumarins in organs that are shielded from light, such as roots. These findings provide routes to improving coumarin production in crops or by microbial fermentation.



Similar content being viewed by others
Data availability
References
Sun, H. et al. Scopoletin is a phytoalexin against Alternaria alternata in wild tobacco dependent on jasmonate signalling. J. Exp. Bot. 65, 4305–4315 (2014).
Gnonlonfin, G. J. B., Sanni, A. & Brimer, L. Review scopoletin—a coumarin phytoalexin with medicinal properties. Crit. Rev. Plant Sci. 31, 47–56 (2012).
Chong, J. et al. Downregulation of a pathogen-responsive tobacco UDP-Glc:phenylpropanoid glucosyltransferase reduces scopoletin glucoside accumulation, enhances oxidative stress, and weakens virus resistance. Plant Cell 14, 1093–1107 (2002).
Stringlis, I. A. et al. MYB72-dependent coumarin exudation shapes root microbiome assembly to promote plant health. Proc. Natl Acad. Sci. USA 115, E5213–E5222 (2018).
Rodríguez-Celma, J. & Schmidt, W. Reduction-based iron uptake revisited: on the role of secreted iron-binding compounds. Plant Signal Behav. 8, e26116 (2013).
Schmid, N. B. et al. Feruloyl-CoA 6′-hydroxylase1-dependent coumarins mediate iron acquisition from alkaline substrates in Arabidopsis. Plant Physiol. 164, 160–172 (2014).
Schmidt, H. et al. Metabolome analysis of Arabidopsis thaliana roots identifies a key metabolic pathway for iron acquisition. PLoS ONE 9, e102444 (2014).
Siwinska, J. et al. Scopoletin 8-hydroxylase: a novel enzyme involved in coumarin biosynthesis and iron-deficiency responses in Arabidopsis. J. Exp. Bot. 69, 1735–1748 (2018).
Tsai, H.-H. et al. Scopoletin 8-hydroxylase-mediated fraxetin production is crucial for iron mobilization. Plant Physiol. 177, 194–207 (2018).
Rajniak, J. et al. Biosynthesis of redox-active metabolites in response to iron deficiency in plants. Nat. Chem. Biol. 14, 442–450 (2018).
Matsumoto, S., Mizutani, M., Sakata, K. & Shimizu, B.-I. Molecular cloning and functional analysis of the ortho-hydroxylases of p-coumaroyl coenzyme A/feruloyl coenzyme A involved in formation of umbelliferone and scopoletin in sweet potato, Ipomoea batatas (L.) Lam. Phytochemistry 74, 49–57 (2012).
Vialart, G. et al. A 2-oxoglutarate-dependent dioxygenase from Ruta graveolens L. exhibits p-coumaroyl CoA 2′-hydroxylase activity (C2′H): a missing step in the synthesis of umbelliferone in plants. Plant J. 70, 460–470 (2012).
Kai, K. et al. Scopoletin is biosynthesized via ortho-hydroxylation of feruloyl CoA by a 2-oxoglutarate-dependent dioxygenase in Arabidopsis thaliana. Plant J. 55, 989–999 (2008).
Karamat, F. et al. CYP98A22, a phenolic ester 3′-hydroxylase specialized in the synthesis of chlorogenic acid, as a new tool for enhancing the furanocoumarin concentration in Ruta graveolens. BMC Plant Biol. 12, 152 (2012).
Tuominen, L. K., Johnson, V. E. & Tsai, C.-J. Differential phylogenetic expansions in BAHD acyltransferases across five angiosperm taxa and evidence of divergent expression among Populus paralogues. BMC Genom. 12, 236 (2011).
Vanholme, R. et al. Caffeoyl shikimate esterase (CSE) is an enzyme in the lignin biosynthetic pathway in Arabidopsis. Science 341, 1103–1106 (2013).
Vanholme, R. et al. A systems biology view of responses to lignin biosynthesis perturbations in Arabidopsis. Plant Cell 24, 3506–3529 (2012).
Yu, X.-H., Gou, J.-Y. & Liu, C.-J. BAHD superfamily of acyl-CoA dependent acyltransferases in Populus and Arabidopsis: bioinformatics and gene expression. Plant Mol. Biol. 70, 421–442 (2009).
Schmid, M. et al. A gene expression map of Arabidopsis thaliana development. Nat. Genet. 37, 501–506 (2005).
Ahn, Y. O. et al. Scopolin-hydrolyzing β-glucosidases in roots of Arabidopsis. Plant Cell Physiol. 51, 132–143 (2010).
Dorey, S. et al. Spatial and temporal induction of cell death, defense genes, and accumulation of salicylic acid in tobacco leaves reacting hypersensitively to a fungal glycoprotein elicitor. Mol. Plant Microbe Interact. 10, 646–655 (1997).
Beuerle, T. & Pichersky, E. Enzymatic synthesis and purification of aromatic coenzyme A esters. Anal. Biochem. 302, 305–312 (2002).
Levsh, O. et al. Dynamic conformational states dictate selectivity toward the native substrate in a substrate-permissive acyltransferase. Biochemistry 55, 6314–6326 (2016).
Walker, A. M. et al. Elucidation of the structure and reaction mechanism of sorghum hydroxycinnamoyltransferase and its structural relationship to other coenzyme A-dependent transferases and synthases. Plant Physiol. 162, 640–651 (2013).
Ma, X., Koepke, J., Panjikar, S., Fritzsch, G. & Stöckigt, J. Crystal structure of vinorine synthase, the first representative of the BAHD superfamily. J. Biol. Chem. 280, 13576–13583 (2005).
Rodríguez-Celma, J. et al. Mutually exclusive alterations in secondary metabolism are critical for the uptake of insoluble iron compounds by Arabidopsis and Medicago truncatula. Plant Physiol. 162, 1473–1485 (2013).
Fourcroy, P. et al. Involvement of the ABCG37 transporter in secretion of scopoletin and derivatives by Arabidopsis roots in response to iron deficiency. New Phytol. 201, 155–167 (2014).
Rizhsky, L., Liang, H. & Mittler, R. The water-water cycle is essential for chloroplast protection in the absence of stress. J. Biol. Chem. 278, 38921–38925 (2003).
Yang, S.-M., Shim, G. Y., Kim, B.-G. & Ahn, J.-H. Biological synthesis of coumarins in Escherichia coli. Microb. Cell Fact. 14, 65 (2015).
Lin, Y., Sun, X., Yuan, Q. & Yan, Y. Combinatorial biosynthesis of plant-specific coumarins in bacteria. Metab. Eng. 18, 69–77 (2013).
Kuromori, T. et al. A collection of 11 800 single-copy Ds transposon insertion lines in Arabidopsis. Plant J. 37, 897–905 (2004).
Ito, T. et al. A new resource of locally transposed Dissociation elements for screening gene-knockout lines in silico on the Arabidopsis genome. Plant Physiol. 129, 1695–1699 (2002).
Sundaresan, V. et al. Patterns of gene action in plant development revealed by enhancer trap and gene trap transposable elements. Genes Dev. 9, 1797–1810 (1995).
Shimada, T. L., Shimada, T. & Hara-Nishimura, I. A rapid and non-destructive screenable marker, FAST, for identifying transformed seeds of Arabidopsis thaliana. Plant J. 61, 519–528 (2010).
Ramakers, C., Ruijter, J. M., Deprez, R. H. L. & Moorman, A. F. M. Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neurosci. Lett. 339, 62–66 (2003).
Czechowski, T., Stitt, M., Altmann, T., Udvardi, M. K. & Scheible, W.-R. Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol. 139, 5–17 (2005).
Kale, N. S. et al. MetaboLights: an open-access database repository for metabolomics data. Curr. Protoc. Bioinform. 53, 14.13.11–14.13.18 (2016).
R Core Team. R: A language and environment for statistical computing (R Foundation for Statistical Computing, 2017).
Murashige, T. & Skoog, F. A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol. Plant. 15, 473–497 (1962).
Rao, G. K., Gowda, N. B. & Ramakrishna, R. A. Raney nickel–catalyzed hydrogenation of unsaturated carboxylic acids with sodium borohydride in water. Synth. Commun. 42, 893–904 (2012).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinform. 10, 241 (2009).
Remmert, M., Biegert, A., Hauser, A. & Söding, J. HHblits: lightning-fast iterative protein sequence searching by HMM-HMM alignment. Nat. Methods 9, 173–175 (2012).
Guex, N., Peitsch, M. C. & Schwede, T. Automated comparative protein structure modeling with SWISS-MODEL and Swiss-PdbViewer: a historical perspective. Electrophoresis 30, S162–S173 (2009).
Benkert, P., Biasini, M. & Schwede, T. Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics 27, 343–350 (2011).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Hiscox, J. D. & Israelstam, G. F. A method for the extraction of chlorophyll from leaf tissue without maceration. Can. J. Bot. 57, 1332–1334 (1979).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Huson, D. H. et al. Dendroscope: an interactive viewer for large phylogenetic trees. BMC Bioinform. 8, 460 (2007).
Acknowledgements
We thank A. Bleys for help with preparing the manuscript. We acknowledge funding through the framework of the IWT-SBO (project ARBOREF), the Hercules program of Ghent University for the Synapt Q-Tof (grant no. AUGE/014), the Bijzonder Onderzoeksfonds-Zware Apparatuur of Ghent University for the FT-ICR-MS instrument (174PZA05) and the Multidisciplinary Research Partnership Biotechnology for a Sustainable Economy (01MRB510W) of Ghent University. R.V. was supported by the Research Foundation—Flanders with a postdoctoral fellowship; L.S., L.H. and B.D.M. by the Agency for Innovation by Science and Technology, Flanders (IWT-Vlaanderen); and X.L. and J.L. by the China Scholarship Council. H.K. and J.R. were funded by the DOE Great Lakes Bioenergy Research Center (DOE BER Office of Science DE-SC0018409) and research in the Schmidt laboratory was supported by MoST.
Author information
Authors and Affiliations
Contributions
R.V., L.S., W.S. and W.B. designed the experiments. R.V., L.S., K.C.S., H.K., X.L., B.D.M., L.H., J.L., G.G., H.-H.T. and K.M. performed experiments. R.V., L.S., G.G., K.M., J.H., B.V., H.-H.T. and J.R. collected and analysed data; R.V. and W.B. wrote the manuscript with the help of all of the authors.
Corresponding author
Ethics declarations
Competing interests
A patent (Belgium Patent publication no: WO 2018/046526 Al) for the use of COSY for the production of coumarins in plants and microorganisms has been published.
Additional information
Peer review information: Nature Plants thanks Chang Jun Liu and **g Ke Weng and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–23, legend of Supplementary Table 1, Supplementary Tables 2 and 3, Supplementary Table 4 legend, Supplementary Tables 5 and 6.
Supplementary Tables 1 and 4
Supplementary Table 1: a full list of peaks detected in the root extracts of cosy-1 and cosy-2. Supplementary Table 4: a full list of peaks detected in the inflorescence stem extracts of cosy-1 and cosy-2.
Rights and permissions
About this article
Cite this article
Vanholme, R., Sundin, L., Seetso, K.C. et al. COSY catalyses trans–cis isomerization and lactonization in the biosynthesis of coumarins. Nat. Plants 5, 1066–1075 (2019). https://doi.org/10.1038/s41477-019-0510-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-019-0510-0
- Springer Nature Limited