Abstract
Activation of the KRAS oncogene is a source of replication stress, but how this stress is generated and how it is tolerated by cancer cells remain poorly understood. Here we show that induction of KRASG12V expression in untransformed cells triggers H3K27me3 and HP1-associated chromatin compaction in an RNA transcription dependent manner, resulting in replication fork slowing and cell death. Furthermore, elevated ATR expression is necessary and sufficient for tolerance of KRASG12V-induced replication stress to expand replication stress-tolerant cells (RSTCs). PrimPol is phosphorylated at Ser255, a potential Chk1 substrate site, under KRASG12V-induced replication stress and promotes repriming to maintain fork progression and cell survival in an ATR/Chk1-dependent manner. However, ssDNA gaps are generated at heterochromatin by PrimPol-dependent repriming, leading to genomic instability. These results reveal a role of ATR-PrimPol in enabling precancerous cells to survive KRAS-induced replication stress and expand clonally with accumulation of genomic instability.
Similar content being viewed by others
Introduction
Genomic instability is a hallmark of cancer and is a driving force for the acquisition of other characteristics of cancer1. In hereditary cancers, mutations in DNA repair genes cause genomic instability that promotes cancer development. On the other hand, recent high-throughput sequencing studies have revealed that sporadic cancers have fewer defects in DNA repair genes2. Instead, more than half of the top 20 genes mutated in human cancers are associated with features of sustained proliferation, raising the possibility that genes driving cell proliferation, such as oncogenes, are embracing complex environmental changes that disrupt cellular processes and contribute to genomic instability3. Such cellular disruptions involve physiological dysregulation of DNA replication and are widely referred to as replication stress (RS). A large body of evidence indicates that during the early stages of tumorigenesis, oncogenes cause RS, leading to genomic instability4. While replication cannot be completed in most of these distressed cells, resulting in programmed cell death or senescence5, a limited number continue to proliferate and expand clonally in noncancer tissues6. These findings raise the question of how cells adapt to RS while driving genomic instability during early carcinogenic processes.
KRAS is one of the most frequently mutated oncogenes in human cancers and is considered to be an important early driver of many tumors7. KRAS mutations commonly lead to substitution of a single amino acid (G12) in the KRAS protein with another amino acid, which causes constitutive activation of the downstream RAS signaling cascade and induces oncogenic events8,9. For instance, such mutations cause loss of normal growth barriers and disruption of tissue homeostasis resulting in increased exposure of mutant KRAS cells to stress-inducing conditions such as RS10,11. Recent evidence suggests that strong overexpression of oncogenic Ras in normal fibroblasts increases RNA synthesis accompanied by R-loop accumulation, which interferes with replication fork progression and causes RS, whereas inhibition of origin firing failed to abrogate RAS-induced RS in the same system12. However, the obstacles to abrogating fork progression have not been fully addressed, and whether a similar mechanism is active in normal human epithelial cells expressing oncogenic KRAS mutants—a frequent situation in human carcinogenesis—remains elusive.
Ataxia Telangiectasia and Rad3-related kinase (ATR), a master regulator of RS, and its critical substrate, Checkpoint Kinase 1 (Chk1), regulate cell cycle checkpoints and facilitate DNA repair via their substrates13,14,15,16,17,18,19. Therefore, ATR and Chk1 have long been considered tumor suppressors20,21,22. However, recent studies have shown that ATR and Chk1 are required for cell survival in response to oncogenic RAS expression during tumorigenesis, suggesting that the ATR-Chk1 kinase pathway protects cells from deleterious and chronic RS induced by oncogenes, thus promoting RS tolerance (RST)23,24,25,26. As a mechanism of RST in human cells, repriming of DNA synthesis at stalled replication forks promotes discontinuous replication at leading strands with single-stranded DNA (ssDNA) gaps27,28. Human primase and DNA-directed polymerase (PrimPol) is a major factor that mediates the repriming process during the RS response29. PrimPol needs to be tightly regulated during DNA replication to prevent aberrant repriming, fork speeding, and chromosomal breakage, which increase the risk of genomic instability30,87 with the human genome (GRCh38) (https://gdc.cancer.gov/about-data/gdc-data-processing/gdc-reference-files) and transcriptome data (GENCODE version 3188) as reference datasets. Transcripts per million (TPM) values were calculated using the StringTie (2.0.4) program89. This study was approved by the Institutional Review Board of the National Cancer Center.
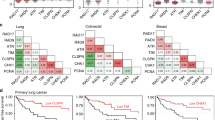
The samples were grouped depending on ATR expression, PrimPol expression, KRAS mutation status, and the previous/current smoking status of the patient. The low and high expression groups for each gene were defined as follows. The cohort was divided into two groups (High and Low) such that the average expression levels in the Low group (Av_Low) and High group (Av_High) met the following condition: the ratio of the average expression levels [(Av_High)/(Av_Low)] is minimized (typically ~2) or not <1.5. The OS of each group was analyzed by the Kaplan‒Meier method, with the log-rank (Mantel‒Cox) test.
Western blot analysis
Cells were lysed in sampling buffer (120 mM Tris-HCl (pH 6.8), 12% glycerol, 4% SDS, 200 mM DTT, 0.004% bromophenol blue) and denatured at 95 °C for 5 min. The protein concentration in the cell lysate was estimated by the Bradford assay (XL-Bradford, Apro Science, Cat# KY-1031). Equal amounts of protein were separated by SDS‒PAGE (10% SDS-polyacrylamide gel) at 120 V for 2 h and were then transferred to polyvinylidene difluoride (PVDF) membranes (Immobilon-P, Millipore, Cat# IPVH00010) at 150 mA for ~16 h at 4 °C. The membranes were blocked with 5% skim milk in TBST (25 mM Tris-HCl (pH 7.8), 140 mM NaCl, 0.1% Tween 20) for 20 min at room temperature (RT) and probed with the following primary antibodies diluted in 5% skim milk in TBST (supplemented with 0.02% of NaN3): anti-pan-RAS (1:1000, Santa Cruz, Cat# sc-166691), anti-αTubulin (1:5000, MBL, Cat# PM054), anti-phospho-ATR (Thr1989) (1:3000, Cell Signaling Technology, Cat# 30632), anti-ATR (1:3000, GeneTex, Cat# GTX128146), anti-phospho-Chk1 (Ser317) (1:500, Cell Signaling Technology, Cat# 12302), anti-phospho-Chk1 (Ser345) (1:500, Cell Signaling Technology, Cat# 2348), anti-Chk1 (1:100, Santa Cruz, Cat# sc-8408), anti-phospho-RB (Ser807/811) (1:1000, Cell Signaling Technology, Cat# 8516), anti-E-cadherin (1:1000, Cell Signaling Technology, Cat# 3195), anti-Vimentin (1:1000, Cell Signaling Technology, Cat# 5741), anti-PrimPol (1:3000, Proteintech, Cat# 29824-1-AP), anti-myc tag (1:1000, MBL, Cat# 192-3), anti-HistoneH1.4 (1:1000, Cell Signaling Technology, Cat# 41328), anti-GAPDH (1:1000, Wako, Cat# 010-25521), anti-phospho-Chk1 (Ser296) (1:500, Cell Signaling Technology, Cat# 90178), anti-RNaseH1 (1:1000, Abcam, Cat# ab56560), anti-HP1α (1:1000, Millipore, Cat# 05-689), anti-HP1β (1:1000, Cell Signaling Technology, Cat# 8676). After 3 washes with TBST, the membranes were incubated with Horseradish Peroxidase (HRP)-conjugated AffiniPure Goat Anti-Rabbit or Mouse IgG (H + L) (1:5000, Jackson ImmunoResearch, Cat# 111-035-003 or Cat# 115-035-003) in 5% skim milk for 1 h at RT and were then washed with TBST 5 times. The blots were developed with Western Lightning Plus ECL Reagent (PerkinElmer, Cat# NEL105001EA) according to the manufacturer’s instructions and imaged with an LAS 3000 luminescent image analyzer (Fujifilm).
Long-term time-to-progression (TTP) assay
A total of 2 × 105 cells were plated in 6-cm plates with or without 0.1 μM 4OHT. After 3 or 4 days of incubation, cells were collected and counted. This 2D culture process was repeated for ~30 days depending on the deleterious effects of 4OHT-induced KRASG12V. The daily growth rate (μ) was calculated according to the formula: μ = (ln N – ln N’)/t, where N is the number of cells at the current week, N’ is the number of cells plated at the previous week, and t is the culture days. Based on the daily growth rate (μ) calculated at every split time, the total current cell number (Nx) was estimated according to the formula ln Nx = ln N0 + μ * T, where N0 is the number of cells plated at the initial split point and T is the total number of incubation days.
Anchorage-independent growth assay
A total of 2 × 104 cells were suspended in 1 ml of 0.4% soft agar solution (SeaPlaque agarose, Lonza, SeaPlaqueTM, Cat# 50101) and then plated on 1.5 ml of 0.7% bottom agar. Top medium containing 0.1 μM of 4OHT was added every 3 ~ 4 days. After ~30 days of culture at 37 °C, cells were stained with 0.01% crystal violet dissolved in 20% MtOH for 1 h at RT and were then imaged using a LAS-3000 imager (Fujifilm).
RNA extraction and real-time PCR
A total of 1 × 105 cells were plated in 6-well plates. Following treatments, total RNA was purified using ISOGEN II (Nippon Gene, Cat# 311-07361) according to the manufacturer’s instructions. Extracted RNA was resuspended in TE buffer (pH 8.0, NipponGene, Cat# 314-90021).
To detect ATR, PrimPol and ActinB mRNA, reverse transcription of the mRNA library was carried out using a High-Capacity RNA-to-cDNA Kit (Applied Biosystems, Cat# 4387406) according to the manufacturer’s instructions. SYBR Green real-time PCR was performed using a KAPA SYBR FAST ABI Prism qPCR Kit (Kapa Biosystems, Cat# KK4604). The following primers were used at 500 nM: ATR-F, 5'-GGTATGCTCTCACTTCCATG-3'; ATR-R, 5'-GTCAGAAGAAACACACATCG-3'; PRIMPOL-F, 5'-TGTGGCTTTGGAGGTTACTGA-3'; PRIMPOL-R, 5'-TTCTACTGAAGTGCCGATACTGT-3'; Hs_ACTB_1_SG QuantiTect Primer Assay (QIAGEN, ID# QT00095431).
To detect miR185, reverse transcription was carried out using a TaqMan™ MicroRNA Reverse Transcription Kit (Applied Biosystems, Cat# 4366596) and TaqMan™ MicroRNA Assays (Applied Biosystems, RNU48: Assay ID# 001006, miR185: Assay ID# 002271) according to the manufacturer’s instructions. TaqMan real-time PCR was performed using TaqMan™ Fast Advanced Master Mix (Applied Biosystems, Cat# 4444556) according to the manufacturer’s instructions.
PCR was performed using a StepOnePlus thermocycler (Applied Biosystem). The comparative Ct (ΔΔCt) method was selected to validate the quantification of mRNA or miRNA expression.
Proliferation assay
A total of 2000 cells were seeded in 96-well plates and treated with 0.1 μM 4OHT. The numbers of proliferating cell were determined 7 days after 4OHT treatment using PrestoBlueTM. According to the manufacturer’s instructions, PrestoBlueTM Cell Viability Reagent (Invitrogen, Cat# A13262) was added directly to the cells and incubated for 1 h at 37 °C. The optical density (excitation: 560 nm, emission: 590 nm) was measured with a Biotek Epoch microplate reader and analyzed with Gen5 2.05 software (Biotek).
For HP1α KD effect validation, 1.5 ×105 cells were transfected with 1 nM siRNA for 2 days and then replated on 96-well plates with 0.1 μM 4OHT. After 5 days of incubation, cell proliferation was evaluated by Cell Titer-Glo 2.0 Assay (Promega, Cat# G9243) according to the manufacturer’s instructions. The luminescence was measured with a Biotek Epoch microplate reader and analyzed with Gen5 2.05 software (Biotek).
DNA fiber assay and S1 fiber assay
The standard DNA fiber assay was previously described53. A total of 1 ×105 cells were plated in 6-well plates. Following treatments, cells were sequentially labeled with 100 μM IdU (5-iodo-2’-deoxyuridine, Sigma–Aldrich, Cat# l7125-5G) for 20 min, washed with PBS 3 times, labeled with 100 μM CldU (5-chloro-2’-deoxyuridine, Sigma–Aldrich, Cat# C6891-100MG) for 40 min, and washed with PBS. The labeled cells were trypsinized and pelleted at 800 x g for 5 min at 4 °C and were then resuspended in 200 μl of PBS.
The S1 fiber assay was performed as previously described with modifications45. Labeled cells were permeabilized with CSK100 (10 mM MOPS (pH 7.0), 100 mM NaCl, 3 mM MgCl2, 300 mM sucrose, 0.5% Triton X-100) for 10 min at RT, gently washed twice with S1 buffer (30 mM sodium acetate (pH 4.6), 50 mM NaCl, 10 mM zinc acetate, 5% glycerol), treated with 10 U/ml S1 nuclease (Invitrogen, Cat# 18001-016) in S1 buffer for 30 min at 37 °C, and collected in 500 μl of PBS containing 1% BSA (Nacalai Tesque, Cat# 01860-07) with a cell scraper. Cell nuclei were pelleted at 4900 x g for 5 min at 4 °C, and 300 μl of the supernatant was then removed, leaving 200 µl. Cell nuclei were carefully resuspended by pipetting.
For both the standard DNA fiber assay and S1 fiber assay, 100 μl of fixative solution (3:1 MtOH:acetic acid) was added dropwise to 200 μl of cell suspension. This step was repeated 3 times, and 500 μl of fixative solution was then added; This step was repeated 2 times. Cells/nuclei were pelleted at 800 x g for 5 min at 4 °C and resuspended in fixative solution to a final concentration of ~1000 cells/μl.
Fifty microliters of fixed cells were spotted onto slides (Matsunami, Cat# S2112). After drying for 3 min, the slides were immersed in lysis solution (200 mM Tris-HCl (pH 7.5), 50 mM EDTA, 0.5% SDS) for ~20 min at 37 °C. The DNA fibers released from the cells were extended by tilting the slides in a high-humidity chamber for at least 30 min. The slides were immersed in fixative solution for 20 min and were then dried overnight (O/N) at 4 °C.
For immunostaining of DNA fibers, DNA fibers were rehydrated in PBS twice for 5 min each, immersed in 2.5 M HCl for 1 h to denature the DNA and washed with PBS 3 times for 5 min each. After blocking (5% BSA and 0.1% Tween 20 in PBS) for 1 h, the slides were incubated for ~2 h at 37 °C with anti-IdU (1:25, BD Biosciences, 347580) and anti-CldU (1:100, Abcam, ab6326) antibodies diluted in blocking solution to label the DNA, washed 3 times with PBST (PBS containing 0.1% Tween 20) and incubated for 1 h with Alexa Fluor 488-conjugated anti-rat IgG (1:500, Invitrogen, Cat# A11006) and Alexa Fluor 555-conjugated anti-mouse IgG (1:250, Invitrogen, Cat# A21422) in blocking solution. The stained slides were washed with PBST 3 times and mounted with ProLong Gold Antifade (Invitrogen, Cat# P36980).
Images were acquired with a fluorescence microscope (Celldiscoverer 7, Zeiss) and analyzed using ImageJ software (v2.0.0-rc-69, National Institutes of Health). Continuous red (IdU) and green (CldU) tracks were measured, and the distance in micrometers was converted to kilobase pairs (1 μm = 3.5 kb). Finally, the fork speed was calculated from the labeling time. A total of 200 ~ 500 fibers were measured in each sample, and 200 randomly selected fibers are shown in the figures.
Immunofluorescence staining
A total of 1 × 104 cells were plated in collagen-coated 8-well chamber dishes. Following treatments, cells were prefixed with ice-cold PBS for 5 min on ice, and a pre-extraction step was performed with pre-extraction buffer (20 mM MOPS (pH 7.0) and 0.05% Triton X-100 in PBS) for 5 min on ice. Cells were fixed with 4% paraformaldehyde (PFA) in PBS for 10 min prior to post-extraction with 0.25% Triton X-100 in PBS for 10 min. To detect incorporated native BrdU, cells were further fixed by ice-cold MtOH for 10 min. After blocking with wash buffer (3% BSA and 0.05% Tween 20 in PBS) for 1 h at 37 °C, the cells were incubated for 1 h at 37 °C or O/N at 4 °C with the following primary antibodies in wash buffer: anti-BrdU (1:1000, GE health, Cat# RPN20AB) anti-RPA32 (1:200, Abcam, Cat# ab2175), anti-phospho-RPA32 (Ser4/8) (1:200, Bethyl, # A300-245A), anti-phospho-RPA32 (Ser33) (1:200, Bethyl, # A300-246A), anti-γH2AX (1:500, Cell Signaling Technology, Cat# 9718), anti-H3K27me3 (1:500, Cell Signaling Technology, Cat# 9733), H3K9me3 (1:500, Abcam, Cat# ab8898), anti-PICH (1:250, Abnova, Cat# H00054821-B01P), and anti-phospho-HistoneH3 (Ser10) (1:500, Millipore, Cat# 06-570). The cells were washed with wash buffer 3 times and incubated with the following Alexa Fluor-conjugated secondary antibodies for 1 h at RT: Alexa 488 donkey anti-rabbit or anti-mouse IgG (H + L) (1:500, Jackson ImmunoResearch, Cat# 711-545-152 or Cat# 715-545-151) and Alexa 594 donkey anti-rabbit or anti-mouse IgG (H + L) (1:500, Jackson ImmunoResearch, Cat# 711-585-152 or Cat# 715-585-151). After 3 washes, the nuclei were stained with 1 μg/ml DAPI and mounted with VECTASHIELD. Immunofluorescence images were acquired with a fluorescence microscope (Celldiscoverer 7, Zeiss) for intensity analysis (Zen v2.6, Zeiss).
PrimPol phosphoproteomics assay
Following treatments, 1.5 × 107 cells were washed with PBS supplemented with PhosSTOP and were then lysed in 8 ml of lysis buffer (30 mM Tris-HCl (pH 7.5), 150 mM NaCl, 2.5 mM MgCl2, 0.5% NP40, PhosSTOP, cOmplete, 1 mM dithiothreitol (DTT)). After treatment with 100 U/ml benzonase for 1 h at 4 °C, the insoluble fraction was pelleted at 20000 × g for 10 min at 4 °C. The cell lysate was collected, and the protein concentration was estimated by the Bradford assay. Samples containing equal amounts of protein were incubated with an anti-myc tag antibody (15 μg/1.5 × 107 cells, MBL, Cat# M192-3) conjugated to Protein-G Sepharose beads (Cytiva, Cat# 17061802) for 4 h at 4 °C, and the beads were washed 3 times with lysis buffer. Immunoprecipitated proteins were denatured with sampling buffer and were then separated by SDS‒PAGE. Proteins were stained using SYPRO Ruby Protein Gel Stain (Invitrogen, Cat# S12000) according to the manufacturer’s instructions.
Protein bands were excised and subjected to in-gel tryptic digestion essentially as described previously90. In brief, the gel pieces were destained and washed, and after DTT reduction and iodoacetamide alkylation, the proteins were digested with trypsin O/N at 37 °C. The resulting tryptic peptides were extracted from the gel pieces by sequential treatment with 30% acetonitrile, 0.3% trifluoroacetic acid and 100% acetonitrile. The extracts were evaporated in a vacuum centrifuge to remove the organic solvent and were then desalted and concentrated with reversed-phase C18 StageTips as previously described91. Immobilized metal affinity chromatography (IMAC) enrichment of phosphopeptides was performed as previously described92.
LC‒MS/MS was performed by coupling an UltiMate 3000 Nano LC system (Thermo Fisher Scientific) and an HTC-PAL autosampler (CTC Analytics) to a Q Exactive hybrid quadrupole-Orbitrap mass spectrometer (Thermo Fisher Scientific). Peptides were delivered to an analytical column (75 μm × 30 cm, packed in-house with ReproSil-Pur C18-AQ, 1.9 μm resin, Dr. Maisch, Ammerbuch, Germany) and separated at a flow rate of 280 nl/min using an 85-min gradient from 5% to 30% of solvent B (solvent A, 0.1% formic acid (FA) and 2% acetonitrile; solvent B, 0.1% FA and 90% acetonitrile). The Q Exactive instrument was operated in data-dependent mode. Survey full-scan MS spectra (m/z 350 to 1800) were acquired in the Orbitrap at a resolution of 70,000 after ion accumulation to a target value of 3 × 106. The dynamic exclusion time was set to 20 s. The 12 most intense multiply charged ions (z ≥ 2) were sequentially accumulated to a target value of 1 × 105 and fragmented in the collision cell by higher-energy collisional dissociation (HCD) with a maximum injection time of 120 ms and a resolution of 35,000. Typical mass spectrometric conditions were as follows: spray voltage, 2 kV; heated capillary temperature, 250 °C; normalized HCD collision energy, 25%. The MS/MS ion selection threshold was set to 2.5 × 104 counts. A 2.0 Da isolation width was chosen.
Raw MS data were processed with MaxQuant (version 1.6.14.0) supported by the Andromeda search engine. The MS/MS spectra were searched against the UniProt human database (https://www.uniprot.org) with the following search parameters: full tryptic specificity, up to two missed cleavage sites, carbamidomethylation of cysteine residues as a fixed modification, and serine, threonine, and tyrosine phosphorylation, N-terminal protein acetylation and methionine oxidation as variable modifications. The false discovery rates (FDRs) of protein groups, peptides, and phosphosites were <0.01. The quantitative values of the phosphorylation sites across the fractions were automatically integrated and summarized in “Phospho (STY) Sites.txt” by MaxQuant. Peptides identified from the reversed database or identified as potential contaminants were not used subsequent analysis.
Chromatin fractionation
The fractionation method was previously described93. A total of 2 × 105 cells were plated in a 60 mm dish. Following treatments, cells were harvested with a scraper, pelleted at 800 × g for 5 min at 4 °C, and resuspended in 120 μl of PBS. Approximately 15% of the cells were collected as the input sample. The cells were pelleted again and were then resuspended in 50 μl of Solution A (10 mM HEPES-KOH (pH 7.9), 10 mM KCl, 1.5 mM MgCl2, 0.34 M sucrose, 10% glycerol, PhosSTOP, cOmplete, 1 mM DTT) supplemented with 0.2% Triton X-100. After incubation on ice for 5 min, the nuclear fraction was precipitated at 2500 x g for 5 min at 4 °C, and the supernatant was then collected as the cytosolic fraction. After washing with Solution A, the nuclear fraction was resuspended in 100 μl of Solution B (3 mM EDTA, 0.2 mM EGTA, 1 mM DTT) and incubated on ice for 20 min. The chromatin fraction was precipitated and washed with Solution B and finally pelleted at 20,000 × g for 5 min at 4 °C. The fraction samples were denatured with sampling buffer, and the protein concentration was estimated by the Bradford assay. Samples containing equal amounts of protein were resolved by SDS‒PAGE and analyzed by Western blotting.
EU incorporation assay
EU incorporation assays were performed with a Click-iT RNA Alexa Fluor 594 Imaging Kit (Invitrogen, Cat# C10330). Following treatments, cells were labeled with 1 mM EU for 1 h or 23 h and were then fixed with 4% PFA in PBS for 15 min. Fixed cells were permeabilized with 0.5% Triton X-100 for 15 min. The Click-iT reaction was performed according to the manufacturer’s instructions. Nuclei were stained with 1 µg/ml DAPI, and slides were mounted with VECTASHIELD (Vector Laboratories, Cat# H-1000). Images were acquired with a fluorescence microscope (Celldiscoverer 7, Zeiss) for intensity analysis (Zen v2.6, Zeiss).
Slot blot assay
The DNA‒RNA hybrid detection method was previously described (Matos et al., 2020). A total of 1 × 105 cells were plated in 6-well plates. Following treatments, cells were trypsinized, pelleted at 800 × g for 5 min at 4 °C, and resuspended in 200 μl of PBS. Total DNA and RNA were purified using a QIAamp DNA Mini Kit (QIAGEN, Cat# 51106) according to the manufacturer’s instructions and were then resuspended in 10 mM TE buffer (pH 8.0). To digest DNA‒RNA hybrids, the nucleic acid solution was treated with 50 mU/μl RNaseH1 (NEB, Cat# M0297) for 1 h at 37 °C, and the reaction was then terminated with 5 mM EDTA. The DNA concentration was estimated using a NanoDrop 1000 spectrophotometer (Thermo Scientific). A total of 125 ng of nucleic acids was spotted on nylon membranes (Amersham Hybond-N, GE Healthcare, Cat# RPN303N) using a BIO-DOT SF apparatus (Bio-Rad) according to the manufacturer’s instructions, and the membrane was subjected to UV crosslinking (120 mJ/cm2). As mentioned in the Western blot method, the membrane was blocked and probed with S9.6 (1:500, Millipore, Cat# MABE1095) and anti-dsDNA (1:5000, Abcam, Cat# ab27156) primary antibodies diluted in Block Ace (DS Pharma Biomedical) prior to incubation with HRP-conjugated secondary antibodies and blot development with Western Lightning Plus ECL reagent. The S9.6 signals were normalized based on the dsDNA signals to determine the relative amount of DNA:RNA hybrids.
MNase sensitivity assay
Cells (8 × 104) were plated on a collagen-coated 35 mm dish. Following treatments, cells were permeabilized with CSK100 for 10 min at RT, gently washed with MNase buffer (50 mM Tris-HCl pH 8.0, 5 mM NaCl, 5 mM CaCl2), and then treated with the 75 gelU/ml of MNase (New England Biolab, Cat# M0247S) in MNase buffer for 5 ~ 20 min at 25 °C. The MNase-treated cells were lysed in 250 μl of TES buffer (10 mM Tris-HCl pH 7.5, 10 mM EDTA, 0.5% SDS). The cell lysates were sequentially treated with 1 mg/ml RNaseA (NipponGene, Cat# 318-06391) for 45 min and with 200 μg/ml Proteinase K (QIAGEN, Cat# 19133) for 90 min at 50 °C, after which an equal volume of phenol/chloroform (NipponGene, Cat# 311-90151) was added. The insoluble fraction was pelleted at 20,000 x g, the supernatant was mixed with 0.5 M of NaCl, and an EtOH wash was performed. The extracted DNA was resuspended in TE buffer (pH 8.0), resolved by 1.2% agarose gel (Fast Gene, Cat# NE-AG02) electrophoresis at 1 μg per lane, and stained with EtBr (NipponGene, Cat# 315-90051). Images were captured with a LAS 3000 luminescent image analyzer (FujiFilm) for intensity analysis with ImageJ.
PLA
A total of 1 × 104 cells were plated in collagen-coated 8-well chamber dishes. To monitor the proximity between ssDNA gaps and H3K27me3, the cells were cultured with 10 μM BrdU (Sigma–Aldrich, Cat# B5002-1G) for 48 h and then incubated without BrdU for 24 h. S-phase cells were pulse labeled with 10 μM EdU for 30 min. As mentioned in the immunofluorescence method, cells were pre-extracted, fixed and processed with the Click-iT Plus EdU Alexa Fluor 488 Imaging Kit (Invitrogen, Cat# C10337) according to the manufacturer’s instructions prior to blocking and primary antibody incubation O/N at 4 °C. PLA was performed using Duolink In Situ PLA kits according to the manufacturer’s instructions (Sigma–Aldrich, Cat# DUO92001, Cat# DUO92005 and Cat# DUO92008) with the following primary antibodies: anti-BrdU (1:1000, GE Healthcare, Cat# RPN202) and anti-H3K27me3 (1:1600, Cell Signaling Technology, Cat# 9733) or anti-PCNA (1:500, Santa Cruz, Cat# sc-56) and anti-phospho-POLII (Ser2) (1:10000, Novus, Cat# NB100-1805). Nuclei were stained with 1 μg/ml DAPI, and slides were mounted with VECTASHIELD. Images were acquired with a fluorescence microscope (Celldiscoverer 7, Zeiss) for PLA focus counting (Zen v2.6, Zeiss) or a confocal laser scanning microscope for representative nucleus imaging (TCS SP8, Leica).
Whole-transcriptome RNA-seq
Following treatments, 1 × 106 cells were washed with PBS, and total RNA was then purified using an RNeasy Plus Mini Kit (QIAGEN, Cat# 74134) according to the manufacturer’s instructions. Paired-end sequencing with a read length of 150 bases was performed on the DNBSEQ-G400 (MGI tech) platform following the manufacturer’s instructions. Raw sequence data quality was checked with FastQC software (ver. 0.11.9). Alignment and TPM calculations were performed using Kallisto with a human cDNA reference. Output files of Kallisto (ver. 0.46.0) were converted to an expression matrix with the ‘tximport (ver. 1.18.0)’ package in R software (ver. 3.6.3). Scaled TPM counts were used for further analysis. The distributions of H3K27me3 ChIP-seq signals within gene loci were visualized with the scale-regions function of deepTools94. H3K27me3 ChIP-seq data of SAECs were provided by Suzuki, A., et al. (2014)49.
Micronucleus assay
Following treatments, cells were fixed with 4% PFA in PBS and stained with an anti-phospho-HistoneH3 (Ser10) antibody (1:500, Millipore, Cat# 06-570) as described above. After imaging with a fluorescence microscope (Celldiscoverer 7, Zeiss), the number of cells with MN or blebs was determined. Phospho-HistoneH3 (Ser10)-positive cells were omitted because of nuclear membrane disappearance. At least 200 cells were analyzed per sample (Zen v3.1, Zeiss).
Plate colony-forming efficiency assay
A total of 1 × 104 cells were plated in 6-well plates and treated with 4OHT and/or berzosertib for 6 days. The medium was changed 3 days after cell seeding. Cells were then stained with 0.5% crystal violet dissolved in 20% MtOH for 30 min and rinsed 3 times with distilled water. After drying, images of each whole 6-well plate were acquired using a LAS-3000 imager (Fujifilm). The mean gray value of each well was quantified using ImageJ.
WGS
Following treatments, ~2 × 105 cells were lysed in 250 μl of TES buffer (10 mM Tris-HCl (pH 7.5), 10 mM EDTA, 0.5% SDS). Cell lysates were sequentially treated with 1 mg/ml RNaseA (NipponGene, Cat# 318-06391) for 45 min and 200 μg/ml proteinase K (QIAGEN, Cat# 19133) for 90 min at 50 °C prior to the addition of 260 μl phenol/chloroform (NipponGene, Cat# 311-90151). The insoluble fraction was pelleted at 20,000 x g, and the supernatant was mixed with 0.5 M NaCl and washed with EtOH 2 times. Extracted DNA was resuspended in 50 μl of TE buffer (pH 8.0).
Genome library preparation was performed with a TruSeq DNA PCR-Free Library Prep Kit (Illumina). Sequencing with 150 bp paired-end reads was performed on the NovaSeq 6000 system (Illumina) at a sequencing depth of ~120x, and the sequence reads were aligned to the human reference genome (hg38) with BWA-MEM (version 0.7.17)95 according to the GATK Best Practices workflow. SAEC cells were defined as the normal-equivalent specimen, and Control-ER-Kras-G12V, RSTC#2, RSTC#5 and RSTC#7 cells were defined as tumor-equivalent specimens. Somatic SNVs and small insertions and deletions (INDELs) were called with Mutect2 (gatk version 4.1.2.0)96 with tumor-normal mode, subtracting variants in normal specimens from those in tumor specimens. These variants were filtered with FilterMutectCalls and FilterAlignmentArtifacts, and those with a variant allele frequency (VAF) of <0.1 were removed.
Structural variations were detected by the Manta program (version 1.6.0)97, which identified the break-end junction by discordant paired and split reads. To identify high-confidence rearrangements, we selected them with the following criteria: PASS in the FILTER field, VAF > 0.1 and split read >0. Circos plots were generated with the shinyCircos program (ver 1)98. Allele-specific copy numbers, tumor purity and ploidy were estimated with FACETS (version 0.6.2)99 with the recommended parameter “cval = 400”.
Single-base substitutions, excluding repeat regions, were categorized into trinucleotide contexts with SigprofilerMatrixGenerator (version 1.2.12)100, and de novo signatures were extracted with SigprofilerExtracter (version 1.1.2)101,102 using nonnegative matrix factorization (NMF) with default parameters. These signatures were decomposed to COSMIC signatures (version 3.3) using cosine similarity and assigned to each sample.
Statistical analysis
Boxplot data are presented as the biological means ± SEM for biological repetition or ± SD for technical repetition. For two-group experiments, statistical significance was determined by a t-test (see the figure legends). For multiple-group experiments, statistical significance was determined by multiple comparison test (see the figure legends). All statistical analysis were performed using Prism 9 software, with p < 0.05 considered statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
RNA-seq data from SAEC cell samples generated in this study have been deposited in the NCBI GEO database under accession code “GSE223027”. ChIP-seq data from SAEC cell samples used in this study have been deposited in the DDBJ data base under accession code “DRA002311”. Proteomics data generated in this study have been deposited in JPOST, a public proteome database certified by the ProteomeXchange Consortium, under accession code “PXD043419 [https://repository.jpostdb.org/preview/1810070348649e9ff3ac6da](Access key:2101)”. WGS data generated in this study have been deposited in DDBJ under accession code “PRJDB16238”. UniProt human database is available at (https://www.uniprot.org). Human genome (GRCh38) data is available at (https://gdc.cancer.gov/about-data/gdc-data-processing/gdc-reference-files).The Whole Transcriptome Sequencing (WTS) data from NCCJ-cohort that support the findings of this study are not publicly available and restrictions apply to the availability of these data. Such WTS data are available through to the corresponding authors (Bunsyo Shiotani: bshiotan@ncc.go.jp) for academic non-commercial research purposes upon reasonable request, and subject to review of a project proposal that will be evaluated by PRISM data access committee, entering into an appropriate data access agreement and subject to any applicable ethical approvals. The data supporting the findings of this study are available within the Article and its supplementary information files or Source Data file. Source data are provided with this paper.
References
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Negrini, S., Gorgoulis, V. G. & Halazonetis, T. D. Genomic instability–an evolving hallmark of cancer. Nat. Rev. Mol. Cell Biol. 11, 220–228 (2010).
Macheret, M. & Halazonetis, T. D. DNA replication stress as a hallmark of cancer. Annu. Rev. Pathol. 10, 425–448 (2015).
Kotsantis, P., Petermann, E. & Boulton, S. J. Mechanisms of oncogene-induced replication stress: Jigsaw falling into place. Cancer Discov. 8, 537–555 (2018).
Halazonetis, T. D., Gorgoulis, V. G. & Bartek, J. An oncogene-induced DNA damage model for cancer development. Science 319, 1352–1355 (2008).
Kakiuchi, N. & Ogawa, S. Clonal expansion in non-cancer tissues. Nat. Rev. Cancer 21, 239–256 (2021).
Pylayeva-Gupta, Y., Grabocka, E. & Bar-Sagi, D. RAS oncogenes: weaving a tumorigenic web. Nat. Rev. Cancer 11, 761–774 (2011).
Prior, I. A., Lewis, P. D. & Mattos, C. A comprehensive survey of Ras mutations in cancer. Cancer Res. 72, 2457–2467 (2012).
Huang, L., Guo, Z., Wang, F. & Fu, L. KRAS mutation: from undruggable to druggable in cancer. Signal Transduct. Target Ther. 6, 386 (2021).
Downward, J. RAS synthetic lethal screens revisited: still seeking the elusive prize. Clin. Cancer Res. 21, 1802–1809 (2015).
Luo, J., Solimini, N. L. & Elledge, S. J. Principles of cancer therapy: oncogene and non-oncogene addiction. Cell 136, 823–837 (2009).
Kotsantis, P. et al. Increased global transcription activity as a mechanism of replication stress in cancer. Nat. Commun. 7, 13087 (2016).
Shiotani, B. et al. Two distinct modes of ATR activation orchestrated by Rad17 and Nbs1. Cell Rep. 3, 1651–1662 (2013).
Saldivar, J. C. et al. An intrinsic S/G2 checkpoint enforced by ATR. Science 361, 806–810 (2018).
Buisson, R. et al. Coupling of Homologous Recombination and the Checkpoint by ATR. Mol. Cell 65, 336–346 (2017).
Matos, D. A. et al. ATR protects the genome against R loops through a MUS81-triggered feedback loop. Mol. Cell 77, 514–527.e514 (2020).
Cimprich, K. A. & Cortez, D. ATR: an essential regulator of genome integrity. Nat. Rev. Mol. Cell Biol. 9, 616–627 (2008).
Liu, Q. et al. Chk1 is an essential kinase that is regulated by Atr and required for the G(2)/M DNA damage checkpoint. Genes Dev. 14, 1448–1459 (2000).
Bartek, J. & Lukas, J. Chk1 and Chk2 kinases in checkpoint control and cancer. Cancer Cell 3, 421–429 (2003).
Brown, E. J. & Baltimore, D. ATR disruption leads to chromosomal fragmentation and early embryonic lethality. Genes Dev. 14, 397–402 (2000).
Fang, Y. et al. ATR functions as a gene dosage-dependent tumor suppressor on a mismatch repair-deficient background. EMBO J. 23, 3164–3174 (2004).
Niida, H. et al. Cooperative functions of Chk1 and Chk2 reduce tumour susceptibility in vivo. EMBO J. 29, 3558–3570 (2010).
Gilad, O. et al. Combining ATR suppression with oncogenic Ras synergistically increases genomic instability, causing synthetic lethality or tumorigenesis in a dosage-dependent manner. Cancer Res. 70, 9693–9702 (2010).
Schoppy, D. W. et al. Oncogenic stress sensitizes murine cancers to hypomorphic suppression of ATR. J. Clin. Investig. 122, 241–252 (2012).
Lopez-Contreras, A. J., Gutierrez-Martinez, P., Specks, J., Rodrigo-Perez, S. & Fernandez-Capetillo, O. An extra allele of Chk1 limits oncogene-induced replicative stress and promotes transformation. J. Exp. Med. 209, 455–461 (2012).
Berti, M., Cortez, D. & Lopes, M. The plasticity of DNA replication forks in response to clinically relevant genotoxic stress. Nat. Rev. Mol. Cell Biol. 21, 633–651 (2020).
Bianchi, J. et al. PrimPol bypasses UV photoproducts during eukaryotic chromosomal DNA replication. Mol. Cell 52, 566–573 (2013).
Mouron, S. et al. Repriming of DNA synthesis at stalled replication forks by human PrimPol. Nat. Struct. Mol. Biol. 20, 1383–1389 (2013).
Garcia-Gomez, S. et al. PrimPol, an archaic primase/polymerase operating in human cells. Mol. Cell 52, 541–553 (2013).
Bailey, L. J. et al. PLK1 regulates the PrimPol damage tolerance pathway during the cell cycle. Sci. Adv. 7, eabh1004 (2021).
Simoneau, A., **ong, R. & Zou, L. The trans cell cycle effects of PARP inhibitors underlie their selectivity toward BRCA1/2-deficient cells. Genes Dev. 35, 1271–1289 (2021).
Quinet, A., Tirman, S., Cybulla, E., Meroni, A. & Vindigni, A. To skip or not to skip: choosing repriming to tolerate DNA damage. Mol. Cell 81, 649–658 (2021).
Mehta, K. P. M. et al. CHK1 phosphorylates PRIMPOL to promote replication stress tolerance. Sci. Adv. 8, eabm0314 (2022).
Otsubo, C. et al. TSPAN2 is involved in cell invasion and motility during lung cancer progression. Cell Rep. 7, 527–538 (2014).
Dendo, K. et al. Induction of non-apoptotic programmed cell death by oncogenic RAS in human epithelial cells and its suppression by MYC overexpression. Carcinogenesis 39, 202–213 (2018).
Shi, L. et al. KRAS induces lung tumorigenesis through microRNAs modulation. Cell Death Dis. 9, 219 (2018).
Shi, L. et al. A KRAS-responsive long non-coding RNA controls microRNA processing. Nat. Commun. 12, 2038 (2021).
Zhao, Y. et al. K-RAS acts as a critical regulator of CD44 to promote the invasiveness and stemness of GBM in response to ionizing radiation. Int J. Mol. Sci. 22, 10923 (2021).
Wang, J. et al. Repression of ATR pathway by miR-185 enhances radiation-induced apoptosis and proliferation inhibition. Cell Death Dis. 4, e699 (2013).
Jackson, D. A. & Pombo, A. Replicon clusters are stable units of chromosome structure: evidence that nuclear organization contributes to the efficient activation and propagation of S phase in human cells. J. Cell Biol. 140, 1285–1295 (1998).
Quinet, A., Carvajal-Maldonado, D., Lemacon, D. & Vindigni, A. in Methods in Enzymology (ed Brandt FE) Ch. 3 (Academic Press, 2017).
Buisson, R., Boisvert, J. L., Benes, C. H. & Zou, L. Distinct but concerted roles of ATR, DNA-PK, and Chk1 in countering replication stress during S phase. Mol. Cell 59, 1011–1024 (2015).
Couch, F. B. et al. ATR phosphorylates SMARCAL1 to prevent replication fork collapse. Genes Dev. 27, 1610–1623 (2013).
Moiseeva, T. et al. ATR kinase inhibition induces unscheduled origin firing through a Cdc7-dependent association between GINS and And-1. Nat. Commun. 8, 1392 (2017).
Quinet, A. et al. PRIMPOL-mediated adaptive response suppresses replication fork reversal in BRCA-deficient cells. Mol. Cell 77, 461–474.e469 (2020).
Tirman, S. et al. Temporally distinct post-replicative repair mechanisms fill PRIMPOL-dependent ssDNA gaps in human cells. Mol. Cell 81, 4026–4040.e4028 (2021).
Quinet, A. et al. Translesion synthesis mechanisms depend on the nature of DNA damage in UV-irradiated human cells. Nucleic Acids Res. 44, 5717–5731 (2016).
Hamperl, S., Bocek, M. J., Saldivar, J. C., Swigut, T. & Cimprich, K. A. Transcription-replication conflict orientation modulates R-loop levels and activates Distinct DNA Damage Responses. Cell 170, 774–786.e719 (2017).
Suzuki, A. et al. Aberrant transcriptional regulations in cancers: genome, transcriptome and epigenome analysis of lung adenocarcinoma cell lines. Nucleic acids Res. 42, 13557–13572 (2014).
Long, Y. et al. RNA is essential for PRC2 chromatin occupancy and function in human pluripotent stem cells. Nat. Genet. 52, 931–938 (2020).
Quinodoz, S. A. et al. RNA promotes the formation of spatial compartments in the nucleus. Cell 184, 5775–5790.e5730 (2021).
Lambert, S. & Carr, A. M. Impediments to replication fork movement: stabilisation, reactivation and genome instability. Chromosoma 122, 33–45 (2013).
Kurashima, K. et al. SMARCA4 deficiency-associated heterochromatin induces intrinsic DNA replication stress and susceptibility to ATR inhibition in lung adenocarcinoma. NAR Cancer 2, zcaa005 (2020).
Krajewski, W. A. Alterations in the internucleosomal DNA helical twist in chromatin of human erythroleukemia cells in vivo influences the chromatin higher-order folding. FEBS Lett. 361, 149–152 (1995).
Catalano, R. et al. A drug repurposing screening reveals a novel epigenetic activity of hydroxychloroquine. Eur. J. Med. Chem. 183, 111715 (2019).
Ziv, Y. et al. Chromatin relaxation in response to DNA double-strand breaks is modulated by a novel ATM- and KAP-1 dependent pathway. Nat. Cell Biol. 8, 870–876 (2006).
Hawkins, R. D. et al. Distinct epigenomic landscapes of pluripotent and lineage-committed human cells. Cell Stem Cell 6, 479–491 (2010).
Boros, J., Arnoult, N., Stroobant, V., Collet, J.-F. & Decottignies, A. Polycomb repressive complex 2 and H3K27me3 cooperate with H3K9 methylation to maintain heterochromatin protein 1α at chromatin. Mol. Cell. Biol. 34, 3662–3674 (2014).
Piberger, A. L. et al. PrimPol-dependent single-stranded gap formation mediates homologous recombination at bulky DNA adducts. Nat. Commun. 11, 5863 (2020).
Guo, X. et al. The molecular origins and pathophysiological consequences of micronuclei: new insights into an age-old problem. Mutat. Res. Rev. Mutat. Res. 779, 1–35 (2019).
Chan, K. L., Palmai-Pallag, T., Ying, S. M. & Hickson, I. D. Replication stress induces sister-chromatid bridging at fragile site loci in mitosis. Nat. Cell Biol. 11, 753–U120 (2009).
Bielski, C. M. et al. Genome doubling shapes the evolution and prognosis of advanced cancers. Nat. Genet. 50, 1189–1195 (2018).
Zeng, J., Hills, S. A., Ozono, E. & Diffley, J. F. X. Cyclin E-induced replicative stress drives p53-dependent whole-genome duplication. Cell 186, 528–542.e14 (2023).
Singh, V. K., Rastogi, A., Hu, X., Wang, Y. & De, S. Mutational signature SBS8 predominantly arises due to late replication errors in cancer. Commun. Biol. 3, 421 (2020).
Bartkova, J. et al. DNA damage response as a candidate anti-cancer barrier in early human tumorigenesis. Nature 434, 864–870 (2005).
Bartkova, J. et al. Oncogene-induced senescence is part of the tumorigenesis barrier imposed by DNA damage checkpoints. Nature 444, 633–637 (2006).
Bartek, J., Mistrik, M. & Bartkova, J. Thresholds of replication stress signaling in cancer development and treatment. Nat. Struct. Mol. Biol. 19, 5–7 (2012).
Bianco, J. N. et al. Overexpression of Claspin and timeless protects cancer cells from replication stress in a checkpoint-independent manner. Nat. Commun. 10, 910 (2019).
Yan, Y. et al. The deubiquitinase USP36 regulates DNA replication stress and confers therapeutic resistance through PrimPol stabilization. Nucleic Acids Res. 48, 12711–12726 (2020).
Guilliam, T. A. et al. Human PrimPol is a highly error-prone polymerase regulated by single-stranded DNA binding proteins. Nucleic Acids Res. 43, 1056–1068 (2015).
Crossley, M. P., Bocek, M. & Cimprich, K. A. R-loops as cellular regulators and genomic threats. Mol. Cell 73, 398–411 (2019).
Macheret, M. & Halazonetis, T. D. Intragenic origins due to short G1 phases underlie oncogene-induced DNA replication stress. Nature 555, 112–116 (2018).
Xu, J. et al. Super-resolution imaging reveals the evolution of higher-order chromatin folding in early carcinogenesis. Nat. Commun. 11, 1899 (2020).
Poole, L. A. & Cortez, D. Functions of SMARCAL1, ZRANB3, and HLTF in maintaining genome stability. Crit. Rev. Biochem. Mol. Biol. 52, 696–714 (2017).
Bansbach, C. E., Betous, R., Lovejoy, C. A., Glick, G. G. & Cortez, D. The annealing helicase SMARCAL1 maintains genome integrity at stalled replication forks. Genes Dev. 23, 2405–2414 (2009).
Middleton, M. R. et al. Phase 1 study of the ATR inhibitor berzosertib (formerly M6620, VX-970) combined with gemcitabine +/− cisplatin in patients with advanced solid tumours. Br. J. Cancer 125, 510–519 (2021).
Yap, T. A. et al. Ceralasertib (AZD6738), an Oral ATR Kinase inhibitor, in combination with carboplatin in patients with advanced solid tumors: a Phase I study. Clin. Cancer Res. 27, 5213–5224 (2021).
Yap, T. A. et al. First-in-human trial of the oral Ataxia Telangiectasia and RAD3-Related (ATR) inhibitor BAY 1895344 in patients with advanced solid tumors. Cancer Discov. 11, 80–91 (2021).
Lecona, E., Fernandez-Capetillo, O. & Targeting, A. T. R. Targeting ATR in cancer. Nat. Rev. Cancer 18, 586–595 (2018).
Kim, H. et al. Targeting the ATR/CHK1 Axis with PARP inhibition results in tumor regression in BRCA-mutant ovarian cancer models. Clin. Cancer Res. 23, 3097–3108 (2017).
Yazinski, S. A. et al. ATR inhibition disrupts rewired homologous recombination and fork protection pathways in PARP inhibitor-resistant BRCA-deficient cancer cells. Genes Dev. 31, 318–332 (2017).
Kim, H. et al. Combining PARP with ATR inhibition overcomes PARP inhibitor and platinum resistance in ovarian cancer models. Nat. Commun. 11, 3726 (2020).
Berti, M. et al. Human RECQ1 promotes restart of replication forks reversed by DNA topoisomerase I inhibition. Nat. Struct. Mol. Biol. 20, 347–354 (2013).
Genois, M. M. et al. CARM1 regulates replication fork speed and stress response by stimulating PARP1. Mol. Cell 81, 784–800.e788 (2021).
Cong, K. et al. Replication gaps are a key determinant of PARP inhibitor synthetic lethality with BRCA deficiency. Mol. Cell 81, 3128–3144.e3127 (2021).
Konstantinopoulos, P. A. et al. Berzosertib plus gemcitabine versus gemcitabine alone in platinum-resistant high-grade serous ovarian cancer: a multicentre, open-label, randomised, phase 2 trial. Lancet Oncol. 21, 957–968 (2020).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Frankish, A. et al. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 47, D766–D773 (2019).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Wilm, M. et al. Femtomole sequencing of proteins from polyacrylamide gels by nano-electrospray mass spectrometry. Nature 379, 466–469 (1996).
Rappsilber, J., Ishihama, Y. & Mann, M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal. Chem. 75, 663–670 (2003).
Adachi, J. et al. Improved proteome and phosphoproteome analysis on a cation exchanger by a combined acid and salt gradient. Anal. Chem. 88, 7899–7903 (2016).
Yang, X. H. & Zou, L. Recruitment of ATR-ATRIP, Rad17, and 9-1-1 complexes to DNA damage. Methods Enzymol. 409, 118–131 (2006).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Li, H. et al. The sequence alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Chen, X. et al. Manta: rapid detection of structural variants and indels for germline and cancer sequencing applications. Bioinformatics 32, 1220–1222 (2016).
Yu, Y., Ouyang, Y. & Yao, W. shinyCircos: an R/Shiny application for interactive creation of Circos plot. Bioinformatics 34, 1229–1231 (2018).
Shen, R. & Seshan, V. E. FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing. Nucleic Acids Res. 44, e131 (2016).
Bergstrom, E. N. et al. SigProfilerMatrixGenerator: a tool for visualizing and exploring patterns of small mutational events. BMC Genom. 20, 685 (2019).
Alexandrov, L. B., Nik-Zainal, S., Wedge, D. C., Campbell, P. J. & Stratton, M. R. Deciphering signatures of mutational processes operative in human cancer. Cell Rep. 3, 246–259 (2013).
Alexandrov, L. B. et al. The repertoire of mutational signatures in human cancer. Nature 578, 94–101 (2020).
Acknowledgements
We thank Dr. Tohru Kiyono for the retroviral plasmid constructs and the SAECs, Dr. Helfrid Hochegger for the RPE-1-puro cells, Dr. Yoshihisa Kobayashi for the H3122 cells, Dr. Makoto Nakanishi for the lentivirus plasmid constructs, Dr. Yusuke Yamamoto for the helpful advice on RNA-seq analysis, and members of the Shiotani lab for discussions. This work was supported by the Princess Takamatsu Cancer Research Fund (17-24912), the National Cancer Center Research and Development Fund (29-Seeds-4), and JSPS KAKENHI (18KK0235 and 18H03378) to B.S. and in part by the Japan Science and Technology Agency (JST) CREST (JPMJCR1689) to R.H. and AIP-PRISM (JPMJCR18Y4) to R.H.
Author information
Authors and Affiliations
Contributions
T.I., M.M., and B.S. designed the project. T.I., M.M., and K.Y. performed the cellular and biochemical experiments and data analysis. T.I., T.Ya., M.N., T. Yo., Y.Y., K.S., H.H., T.K., R.H., and B.S. performed clinical data analysis. T.I. and J.A. performed proteomic experiments and data analysis. T.I., A.M., K.S., T.K., R.H., and B.S. performed whole-genome sequencing experiments and data analysis. N.T., A.Y., L.Z., and B.S. supervised the experiments and data analysis. T.I. and B.S. wrote the manuscript with contributions from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Igarashi, T., Mazevet, M., Yasuhara, T. et al. An ATR-PrimPol pathway confers tolerance to oncogenic KRAS-induced and heterochromatin-associated replication stress. Nat Commun 14, 4991 (2023). https://doi.org/10.1038/s41467-023-40578-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-023-40578-2
- Springer Nature Limited
This article is cited by
-
A perspective on tumor radiation resistance following high-LET radiation treatment
Journal of Cancer Research and Clinical Oncology (2024)