Abstract
Circular RNA (circRNAs) is a covalently closed circular non-coding RNA formed by reverse back-splicing from precursor messenger RNA. It is found widely in eukaryotic cells and can be released to the surrounding environment and captured by other cell types. This, circRNAs serve as connections between different cell types for the mediation of multiple signaling pathways. CircRNAs reshape the tumor microenvironment (TME), a key factor involved in all stages of cancer development, by regulating epithelial-stromal transformation, tumor vascularization, immune cell function, and inflammatory responses. Immune cells are the most abundant cellular TME components, and they have profound toxicity to cancer cells. This review summarizes circRNA regulation of immune cells, including T cells, natural killer cells, and macrophages; highlights the impact of circRNAs on tumor progression, treatment, and prognosis; and indicates new targets for tumor immunotherapy.
Similar content being viewed by others
Facts
-
The expression of circRNAs is frequently dysregulated in human cancers.
-
CircRNAs play different roles during tumorigenesis and cancer progression.
-
CircRNAs regulate T cells, NK cells, and macrophages to reshape the tumor microenvironment.
-
CircRNA regulation of the tumor microenvironment provides potential therapeutic opportunities for cancer treatment.
Questions
-
Why circRNAs have multiple functions in the same or different human cancers. What are the underlining molecular determinants of this specificity?
-
Does dose-dependent targeting of circRNAs work in mouse models, at least in three-dimensional tumor organoid models?
-
Is circRNA targeting applicable in clinical trials?
-
Can we design prophylactic or therapeutic anti-cancer approaches based on genetic of polymorphisms of circRNAs?
Introduction
Circular RNAs (circRNA) is a closed circular molecule that is resistant to exonucleases, and is thus stable and widespread in animals and plants. CircRNA was discovered in 1976 when the Sanger team studied virus-like RNAs [1]. In 1991, Nigro et al. [2] accidentally discovered a normal novel RNA product. Due to its low expression and the limitations of detection technology, circRNA was originally considered to be an aberrant product of RNA splicing. Recently, with advances in high-throughput sequencing technology, increasing numbers of circRNAs have been characterized and their roles and mechanisms have become active areas of investigation [3,4,5].
The immune system maintains homeostasis through immunomodulation, surveillance, and the prevention of pathogen invasion. The immune response coordinates a variety of immune cells and has antiviral, antibacterial, and antitumor functions. With rapid developments in oncology, immunology, molecular biology, and related disciplines, immunotherapies such as immune checkpoint inhibitors, tumor vaccines, and adoptive cell therapy have revolutionized cancer treatment. However, therapeutic responses, especially those of solid tumors, have been unsatisfactory in clinical trials and clinical applications. Recent studies have demonstrated that circRNAs are involved in cancer development [6,7,8] and immune responses [9,10,11,12]. In this review, we discuss the roles of circRNAs in the regulation of immune cells, immune-related molecules, and tumor immunity. We anticipate that this summary of current knowledge will facilitate the development of strategies to target circRNAs in the immune microenvironments of human cancers.
Biogenesis and function of circRNAs
CircRNA is a class of non-coding RNA generated from precursor messenger RNA (mRNA). Most circRNAs originate from exons in gene coding regions; others originate from 3′–untranslated regions (UTRs), 5'-UTRs, introns, intergenic regions, and antisense RNA [13, 14]. CircRNAs can be divided into four categories based on their sequence origin: (1) exonic circular RNAs (EciRNAs) derived from exons of the parent gene; (2) lasso-type or circular intronic RNAs (ciRNAs) derived from introns; (3) exonic–intronic circular RNAs (EIciRNAs) derived from both exons and introns; and (4) other circRNAs, including those derived from antisense strand transcripts (antisense circRNAs) and those derived from intergenic sequences or other unannotated genomic sequences (intergenic circRNAs) [15]. About 80% of circRNAs are EciRNAs localized mainly to the cytoplasm, whereas ciRNAs and EIciRNAs are often localized to the nucleus. CircRNAs are relatively evolutionarily conserved in different species. Jeck et al. [16] used the genome-wide RNase R enrichment method to detect >25,000 circRNAs in fibroblasts. Wang et al. [17] observed circRNA expression in fungi, plants, and prokaryotes, reflecting a high degree of conservation and widespread distribution among species. The expression of the same circRNA varies greatly under diseased and non-diseased conditions, among tissues, and during different time periods. The half-life of circRNAs exceeds that of the associated linear mRNA, as the covalent closed-loop structure lacks 5′ and 3′ends, which makes circRNAs more resistant to the exonuclease RNase R [18].
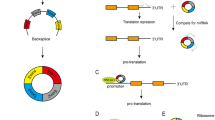
CircRNAs have four main biological functions (Fig. 1). (1) As they contain a large number of micro-RNA (miRNA) binding sites, they serve as molecular sponges and compete for miRNA binding to target mRNAs, thereby upregulating the expression of target genes [19,38,39,40]. Tumor cell-derived circRNAs have recently been reported to play a vital and direct role in tumor immune escape (Table 1). Mechanically, circRNAs enhance the interaction between the immunosuppressive molecule programmed death receptor 1 (PD-1) and its ligand (PD-L1) by upregulating PD-1 expression in T cells, suppressing T-cell activation and cytokine secretion. Exosomes derived from different tumor cells deliver various circRNAs to T cells to inhibit their killing ability via PD-1 upregulation. Those derived from ovarian cancer cells were found to deliver circ-0001068 into T cells, increasing PD-1 expression via miR-28-5p sponging and thereby causing T-cell exhaustion [41]. In lung adenocarcinoma, circRNA-002178 was found to enter CD8+ T cells via exosomes and upregulate PD-1 expression by absorbing miR-34a [42]. circRNA can also upregulate the expression of the immune checkpoint molecules PD-L1 and CD73 on tumor cell surfaces via miRNA sponging, which helps tumor cells to escape recognition and death by T cells. Multiple studies have shown that circRNAs regulate PD-L1 expression via the circRNA–miRNA–mRNA axis, for instance, the circRNA of vimentin, CDR1-AS, hsa_circ_0003288, hsa_circ_0000190, hsa_circ_0046523, circ-CPA4, hsa-circRNA-002178, circ_0000284, circ_001678, circ-HSP90A, and circIGF2BP3 (Table 1) [43,44,114,115].
TAMs form the most abundant immune cell population in the TME. CircRNAs in tumor cells can regulate macrophage polarization through multiple pathways in the TME (Table 3, Fig. 4): (1) they induce crosstalk between tumor cells and macrophages (Fig. 4A), (2) they promote chemokine secretion from tumor cells (Fig. 4B), (3) tumor cell-derived circRNAs in exosomes enter macrophages to play a regulatory role (Fig. 4C), and (4) they promote tumor-cell expression of cytokines such as IL-4 and PD-L1 (Fig. 4D). In-depth investigation of the mechanisms underlying these roles and preclinical studies are urgently needed.
A CircRNAs mediate crosstalk between tumor cells and macrophages, induce M2 macrophage polarization, and impair T cell function, resulting in the formation of an immunosuppressive tumor microenvironment (TME). B CircRNAs upregulate chemokine expression in tumor cells and induce M2 macrophage polarization. C Tumor cells release exosomes containing circRNAs into macrophages to enhance M2 macrophage polarization. D CircRNAs promote the secretion of inflammatory factors and immunosuppressive molecules in tumor cells to recruit and induce M2 macrophage polarization and disable T cells.
CircRNAs regulate neutrophils, myeloid-derived suppressor cells, and cancer-associated fibroblast
The neutrophils are also an important component in the TME, participating in different stages of tumor development and progression such as tumorigenesis, proliferation and metastasis. Neutrophils could play dual roles as a pro-tumor(N2) or tumor suppressor (N1) in the tumor microenvironment due to heterogeneous phenotypes and functional diversity. Recently, mounting evidence show that circular RNA affects tumor development by regulating the function of neutrophils. In bladder cancer, circDHTKD1 recruited and activated neutrophils by inducing CXCL5 expression, and then neutrophils participated in lymphangiogenesis by secreting VEGF-C, facilitating lymphatic metastasis of bladder cancer cells [116]. But in CRC, circPACRGL mainly promoted differentiation of N1 to N2 neutrophils by sponging miR-142-3p/miR-506-3p, N2 neutrophils increased the expression of transforming growth factor-β1 (TGF-β1), which promoted CRC cell proliferation, migration and invasion [117]. Although the underlying mechanism is not very clear, but the diversity and plasticity of neutrophils maybe act as a potential and promising immunotherapy target in clinical treatment.
Myeloid-derived suppressor cells (MDSC) are also another key player in TME. In addition to the immunosuppressive effect, MDSC can also exert tumor-promoting effects by promoting angiogenesis, invasion and metastasis. More details about non-coding RNAs including circRNAs modulate MDSCs in TME have been summarized elsewhere [118].
Cancer-associated fibroblasts (CAFs), also named as tumor-associated fibroblast, are a key factor in tumor microenvironment. It plays important role in tumor growth and metastasis due to diverse functions, such as interactions with cancer cells and crosstalk with infiltrating leukocytes and so on. In pancreatic cancer, Hu et al found that circFARP1 upregulated the expression and secretion of LIF via CAV1 in CAFs to induce chemoresistance [51,52,53, 57]. Stromal cells, such as cancer-associated fibroblasts, endothelial cells, and pericytes, are important TME components, and much more research is warranted to explore their potential regulation. Although TME reprogramming is considered to be a potentially effective strategy for tumor eradication and the improvement of tumor immunotherapy efficacy, there is still a long way to go before we can conquer cancer. For example, does dose targeting of circRNAs work in mouse models, at least in three-dimensional tumor organoid models? Is circRNA targeting applicable in clinical trials? Thorough investigations of circRNAs using animal models would help to accelerate the translation of basic research into clinical practice. We believe that an improved understanding of circRNA functions and mechanisms related to tumorigenesis and immunotherapy would certainly contribute to the development of new therapeutic strategies for cancer.
References
Sanger HL, Klotz G, Riesner D, Gross HJ, Kleinschmidt AK. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc Natl Acad Sci USA. 1976;73:3852–6.
Nigro JM, Cho KR, Fearon ER, Kern SE, Ruppert JM, Oliner JD, et al. Scrambled exons. Cell. 1991;64:607–13.
Chen LL, Yang L. Regulation of circRNA biogenesis. RNA Biol. 2015;12:381–8.
Chen LL. The expanding regulatory mechanisms and cellular functions of circular RNAs. Nat Rev Mol Cell Biol. 2020;21:475–90.
Liu CX, Li X, Nan F, Jiang S, Gao X, Guo SK, et al. Structure and degradation of circular RNAs regulate PKR activation in innate immunity. Cell. 2019;177:865–880.
Han J, Zhao G, Ma X, Dong Q, Zhang H, Wang Y, et al. CircRNA circ-BANP-mediated miR-503/LARP1 signaling contributes to lung cancer progression. Biochem Biophys Res Commun. 2018;503:2429–35.
Yang X, Ye T, Liu H, Lv P, Duan C, Wu X, et al. Expression profiles, biological functions and clinical significance of circRNAs in bladder cancer. Mol Cancer. 2021;20:4.
Li YF, Zhang J, Yu L. Circular RNAs regulate cancer onset and progression via Wnt/β-Catenin signaling pathway. Yonsei Med J. 2019;60:1117–28.
Duan S, Wang S, Huang T, Wang J, Yuan X. circRNAs: Insight into their role in tumor-associated macrophages. Front Oncol. 2021;11:780744.
Diaz-Munoz MD, Turner M. Uncovering the role of RNA-binding proteins in gene expression in the immune system. Front Immunol. 2018;9:1094.
Li Z, Cheng Y, Wu F, Wu L, Cao H, Wang Q, et al. The emerging landscape of circular RNAs in immunity: breakthroughs and challenges. Biomark Res. 2020;8:25.
Taheri M, Barth DA, Kargl J, Rezaei O, Ghafouri-Fard S, Pichler M. Emerging role of non-coding RNAs in regulation of T-lymphocyte function. Front Immunol. 2021;12:756042.
Kristensen LS, Andersen MS, Stagsted LVW, Ebbesen KK, Hansen TB, Kjems J. The biogenesis, biology and characterization of circular RNAs. Nat Rev Genet. 2019;20:675–91.
Ashwal-Fluss R, Meyer M, Pamudurti NR, Ivanov A, Bartok O, Hanan M, et al. circRNA biogenesis competes with pre-mRNA splicing. Mol Cell. 2014;56:55–66.
Li X, Yang L, Chen LL. The biogenesis, functions, and challenges of circular RNAs. Mol Cell. 2018;71:428–42.
Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J, et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA. 2013;19:141–57.
Wang PL, Bao Y, Yee MC, Barrett SP, Hogan GJ, Olsen MN, et al. Circular RNA is expressed across the eukaryotic tree of life. PLoS One. 2014;9:e90859.
Wang Y, Liu J, Ma J, Sun T, Zhou Q, Wang W, et al. Exosomal circRNAs: biogenesis, effect and application in human diseases. Mol Cancer. 2019;18:116.
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell. 2011;146:353–8.
Shuwen H, Qing Z, Yan Z, ** Y. Competitive endogenous RNA in colorectal cancer: a systematic review. Gene. 2018;645:157–62.
Zhang S. The characteristics of circRNA as competing endogenous RNA in pathogenesis of acute myeloid leukemia. BMC Cancer. 2021;21:277.
** Y, Yu LL, Zhang B, Liu CF, Chen Y. Circular RNA hsa_circ_0000523 regulates the proliferation and apoptosis of colorectal cancer cells as miRNA sponge. Braz J Med Biol Res. 2018;51:e7811.
Li Z, Huang C, Bao C, Chen L, Lin M, Wang X, et al. Corrigendum: Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol. 2017;24:194.
Zang J, Lu D, Xu A. The interaction of circRNAs and RNA binding proteins: an important part of circRNA maintenance and function. J Neurosci Res. 2020;98:87–97.
Li Q, Wang Y, Wu S, Zhou Z, Ding X, Shi R, et al. CircACC1 regulates assembly and activation of AMPK complex under metabolic stress. Cell Metab. 2019;30:157–173.
Pan Z, Cai J, Lin J, Zhou H, Peng J, Liang J, et al. A novel protein encoded by circFNDC3B inhibits tumor progression and EMT through regulating Snail in colon cancer. Mol Cancer. 2020;19:71.
Zheng X, Chen L, Zhou Y, Wang Q, Zheng Z, Xu B, et al. A novel protein encoded by a circular RNA circPPP1R12A promotes tumor pathogenesis and metastasis of colon cancer via Hippo-YAP signaling. Mol Cancer. 2019;18:47.
Liu X, Liu Y, Liu Z, Lin C, Meng F, Xu L, et al. CircMYH9 drives colorectal cancer growth by regulating serine metabolism and redox homeostasis in a p53-dependent manner. Mol Cancer. 2021;20:114.
Chen J, Liu G, Wu Y, Ma J, Wu H, **e Z, et al. Correction to: CircMYO10 promotes osteosarcoma progression by regulating miR-370-3p/RUVBL1 axis to enhance the transcriptional activity of beta-catenin/LEF1 complex via effects on chromatin remodeling. Mol Cancer. 2020;19:75.
Restifo NP, Dudley ME, Rosenberg SA. Adoptive immunotherapy for cancer: harnessing the T cell response. Nat Rev Immunol. 2012;12:269–81.
Forman J. T cells, the MHC, and function. Immunol Rev. 1984;81:203–19.
Petrova G, Ferrante A, Gorski J. Cross-reactivity of T cells and its role in the immune system. Crit Rev Immunol. 2012;32:349–72.
Pittet MJ, Speiser DE, Valmori D, Rimoldi D, Lienard D, Lejeune F, et al. Ex vivo analysis of tumor antigen specific CD8+ T cell responses using MHC/peptide tetramers in cancer patients. Int Immunopharmacol. 2001;1:1235–47.
Villa-Morales M, Fernandez-Piqueras J. Targeting the Fas/FasL signaling pathway in cancer therapy. Expert Opin Ther Targets. 2012;16:85–101.
Wajant H. CD95L/FasL and TRAIL in tumour surveillance and cancer therapy. Cancer Treat Res. 2006;130:141–65.
Law LW, Appella E, Henriksen O, Rogers M. Some biologic and biochemical properties of soluble tumor antigens. Ann N. Y Acad Sci. 1976;276:11–25.
Zusman I. Soluble tumor-associated antigens in cancer detection, prevention and therapy. Med Sci Monit. 2004;10:RA317–24.
Kok VC, Yu CC. Cancer-derived exosomes: their role in cancer biology and biomarker development. Int J Nanomed. 2020;15:8019–36.
He L, Zhu W, Chen Q, Yuan Y, Wang Y, Wang J, et al. Ovarian cancer cell-secreted exosomal miR-205 promotes metastasis by inducing angiogenesis. Theranostics. 2019;9:8206–20.
Milane L, Singh A, Mattheolabakis G, Suresh M, Amiji MM. Exosome mediated communication within the tumor microenvironment. J Control Release. 2015;219:278–94.
Wang X, Yao Y, ** M. Circ-0001068 is a novel biomarker for ovarian cancer and inducer of PD1 expression in T cells. Aging (Albany NY). 2020;12:19095–106.
Wang J, Zhao X, Wang Y, Ren F, Sun D, Yan Y, et al. circRNA-002178 act as a ceRNA to promote PDL1/PD1 expression in lung adenocarcinoma. Cell Death Dis. 2020;11:32.
Xu YJ, Zhao JM, Gao C, Ni XF, Wang W, Hu WW, et al. Hsa_circ_0136666 activates Treg-mediated immune escape of colorectal cancer via miR-497/PD-L1 pathway. Cell Signal. 2021;86:110095.
Gao C, Xu YJ, Qi L, Bao YF, Zhang L, Zheng L. CircRNA VIM silence synergizes with sevoflurane to inhibit immune escape and multiple oncogenic activities of esophageal cancer by simultaneously regulating miR-124/PD-L1 axis. Cell Biol Toxicol. 2022;38:825–45.
Zou Y, Zheng S, Deng X, Yang A, **e X, Tang H, et al. The role of circular RNA CDR1as/ciRS-7 in regulating tumor microenvironment: a pan-cancer analysis. Biomolecules. 2019;9:429.
Xu G, Zhang P, Liang H, Xu Y, Shen J, Wang W, et al. Circular RNA hsa_circ_0003288 induces EMT and invasion by regulating hsa_circ_0003288/miR-145/PD-L1 axis in hepatocellular carcinoma. Cancer Cell Int. 2021;21:212.
Luo YH, Yang YP, Chien CS, Yarmishyn AA, Adekunle Ishola A, Chien Y, et al. Circular RNA hsa_circ_0000190 facilitates the tumorigenesis and immune evasion by upregulating the expression of soluble PD-L1 in non-small-cell lung cancer. Int J Mol Sci. 2021;23:64.
Fu X, Sun G, Tu S, Fang K, **ong Y, Tu Y, et al. Hsa_circ_0046523 mediates an immunosuppressive tumor microenvironment by regulating MiR-148a-3p/PD-L1 axis in pancreatic cancer. Front Oncol. 2022;12:877376.
Hong W, Xue M, Jiang J, Zhang Y, Gao X. Circular RNA circ-CPA4/ let-7 miRNA/PD-L1 axis regulates cell growth, stemness, drug resistance and immune evasion in non-small cell lung cancer (NSCLC). J Exp Clin Cancer Res. 2020;39:149.
Li L, Zhang Q, Lian K. Circular RNA circ_0000284 plays an oncogenic role in the progression of non-small cell lung cancer through the miR-377-3p-mediated PD-L1 promotion. Cancer Cell Int. 2020;20:247.
Tian Q, Wu T, Zhang X, Xu K, Yin X, Wang X, et al. Immunomodulatory functions of the circ_001678/miRNA-326/ZEB1 axis in non-small cell lung cancer via the regulation of PD-1/PD-L1 pathway. Hum Mol Genet. 2022;31:4094–106.
Lei J, Zhu J, Hui B, Jia C, Yan X, Jiang T, et al. Circ-HSP90A expedites cell growth, stemness, and immune evasion in non-small cell lung cancer by regulating STAT3 signaling and PD-1/PD-L1 checkpoint. Cancer Immunol Immunother. 2023;72:101–24.
Liu Z, Wang T, She Y, Wu K, Gu S, Li L, et al. N(6)-methyladenosine-modified circIGF2BP3 inhibits CD8(+) T-cell responses to facilitate tumor immune evasion by promoting the deubiquitination of PD-L1 in non-small cell lung cancer. Mol Cancer. 2021;20:105.
Yang Z, Chen W, Wang Y, Qin M, Ji Y. CircKRT1 drives tumor progression and immune evasion in oral squamous cell carcinoma by sponging miR‐495‐3p to regulate PDL1 expression. Cell Biol Int. 2021;45:1423–35.
Ghalamfarsa G, Kazemi MH, Raoofi Mohseni S, Masjedi A, Hojjat-Farsangi M, Azizi G, et al. CD73 as a potential opportunity for cancer immunotherapy. Expert Opin Ther Targets. 2019;23:127–42.
Allard D, Chrobak P, Allard B, Messaoudi N, Stagg J. Targeting the CD73-adenosine axis in immuno-oncology. Immunol Lett. 2019;205:31–39.
Xu YP, Dong ZN, Wang SW, Zheng YM, Zhang C, Zhou YQ, et al. circHMGCS1-016 reshapes immune environment by sponging miR-1236-3p to regulate CD73 and GAL-8 expression in intrahepatic cholangiocarcinoma. J Exp Clin Cancer Res. 2021;40:290.
Li P, Ren X, Zheng Y, Sun J, Ye G. Tumor promoting effect of circ_002172 associates with induced immune escape in breast cancer via the miR-296-5p/CXCL12 axis. Int Immunopharmacol. 2022;106:108530.
Garg B, Giri B, Modi S, Sethi V, Castro I, Umland O, et al. NFkappaB in pancreatic Stellate cells reduces infiltration of tumors by cytotoxic T cells and killing of cancer cells, via up-regulation of CXCL12. Gastroenterology. 2018;155:880–891.
Chen CD, Wang CS, Huang YH, Chien KY, Liang Y, Chen WJ, et al. Overexpression of CLIC1 in human gastric carcinoma and its clinicopathological significance. Proteomics. 2007;7:155–67.
**a J, Wang Q, Ju F, Luo X, Wang F, Zhou Y, et al. Chloride intracellular channel 1 is a potential biomarker for breast cancer. Breast Cancer. 2022;14:247–58.
Yasuda Y, Nagano T, Jimbo N, Kiriu T, Suraya R, Hazama D, et al. Chloride intracellular channel 1 expression is associated with poor prognosis of lung adenocarcinoma. Anticancer Res. 2022;42:271–7.
Zhang S, Wang XM, Yin ZY, Zhao WX, Zhou JY, Zhao BX, et al. Chloride intracellular channel 1 is overexpression in hepatic tumor and correlates with a poor prognosis. APMIS. 2013;121:1047–53.
Li B, Liang L, Chen Y, Liu J, Wang Z, Mao Y, et al. Circ_0008287 promotes immune escape of gastric cancer cells through impairing microRNA-548c-3p-dependent inhibition of CLIC1. Int Immunopharmacol. 2022;111:108918.
Yang C, Wu S, Mou Z, Zhou Q, Dai X, Ou Y, et al. Exosome-derived circTRPS1 promotes malignant phenotype and CD8+ T cell exhaustion in bladder cancer microenvironments. Mol Ther. 2022;30:1054–70.
Zhao X, Lu C, Chu W, Zhang B, Zhen Q, Wang R, et al. MicroRNA-124 suppresses proliferation and glycolysis in non-small cell lung cancer cells by targeting AKT-GLUT1/HKII. Tumour Biol. 2017;39:1010428317706215.
Guo XY, Chen JN, Sun F, Wang YQ, Pan Q, Fan JG. circRNA_0046367 prevents hepatoxicity of lipid peroxidation: an inhibitory role against hepatic steatosis. Oxid Med Cell Longev. 2017;2017:3960197.
Tanaka A, Sakaguchi S. Targeting Treg cells in cancer immunotherapy. Eur J Immunol. 2019;49:1140–6.
Chen Y, Li Z, Liang J, Liu J, Hao J, Wan Q, et al. CircRNA has_circ_0069313 induced OSCC immunity escape by miR-325-3p-Foxp3 axes in both OSCC cells and Treg cells. Aging (Albany NY). 2022;14:4376–89.
Huang M, Huang X, Huang N. Exosomal circGSE1 promotes immune escape of hepatocellular carcinoma by inducing the expansion of regulatory T cells. Cancer Sci. 2022;113:1968–83.
Fang Z, Jiang C, Li S. The potential regulatory roles of circular RNAs in tumor immunology and immunotherapy. Front Immunol. 2021;11:617583.
Liu H, Wang S, **n J, Wang J, Yao C, Zhang Z. Role of NKG2D and its ligands in cancer immunotherapy. Am J Cancer Res. 2019;9:2064–78.
Guillerey C. NK Cells in the Tumor Microenvironment. Adv Exp Med Biol. 2020;1273:69–90.
Zhang PF, Gao C, Huang XY, Lu JC, Guo XJ, Shi GM, et al. Cancer cell-derived exosomal circUHRF1 induces natural killer cell exhaustion and may cause resistance to anti-PD1 therapy in hepatocellular carcinoma. Mol Cancer. 2020;19:110.
Li S, Chen Z, Zhou R, Wang S, Wang W, Liu D, et al. Hsa_circ_0048674 facilitates hepatocellular carcinoma progression and natural killer cell exhaustion depending on the regulation of miR-223-3p/PDL1. Histol Histopathol. 2022;37:1185–1199.
Ma Y, Zhang C, Zhang B, Yu H, Yu Q. circRNA of AR-suppressed PABPC1 91 bp enhances the cytotoxicity of natural killer cells against hepatocellular carcinoma via upregulating UL16 binding protein 1. Oncol Lett. 2019;17:388–97.
Shi M, Li ZY, Zhang LM, Wu XY, **ang SH, Wang YG, et al. Hsa_circ_0007456 regulates the natural killer cell-mediated cytotoxicity toward hepatocellular carcinoma via the miR-6852-3p/ICAM-1 axis. Cell Death Dis. 2021;12:94.
Zhang W, Zhong W, Wang B, Yang J, Yang J, Yu Z, et al. ICAM-1-mediated adhesion is a prerequisite for exosome-induced T cell suppression. Dev Cell. 2022;57:329–343.
Dieterich LC. Mechanisms of extracellular vesicle-mediated immune evasion in melanoma. Front Immunol. 2022;13:1002551.
Ou ZL, Luo Z, Wei W, Liang S, Gao TL, Lu YB. Hypoxia-induced shedding of MICA and HIF1A-mediated immune escape of pancreatic cancer cells from NK cells: role of circ_0000977/miR-153 axis. RNA Biol. 2019;16:1592–603.
Li M, Zhang M, Chen M, **ao J, Mu X, Peng J. KLF2-induced circZKSCAN1 potentiates the tumorigenic properties of clear cell renal cell carcinoma by targeting the miR-1294/PIM1 axis. Cell Cycle. 2022;21:1376–90.
Luchtenborg AM, Metzger P, Cosenza Contreras M, Oria V, Biniossek ML, Lindner F, et al. Kruppel-like factor 7 influences translation and pathways involved in ribosomal biogenesis in breast cancer. Breast Cancer Res. 2022;24:65.
**u DH, Chen Y, Liu L, Yang HS, Liu GF. Tumor-suppressive role of Kruppel-like factor 4 (KLF-4) in colorectal cancer. Genet Mol Res. 2017;16. https://doi.org/10.4238/gmr16019272.
**ao S, **-**ang Y, Long T, **u-Rong L, Hong G, Jie-Cheng Y, et al. Kruppel-like factor 2 disturb non-small cell lung cancer energy metabolism by inhibited glutamine consumption. J Pharm Pharm. 2020;72:843–51.
Yang F, Chen Y, Luo L, Nong S, Li T. circFOXO3 induced by KLF16 modulates clear cell renal cell carcinoma growth and natural killer cell cytotoxic activity through sponging miR-29a-3p and miR-122-5p. Dis Markers. 2022;2022:6062236.
Martinez-Rojo E, Berumen LC, Garcia-Alcocer G, Escobar-Cabrera J. The role of androgens and androgen receptor in human bladder cancer. Biomolecules. 2021;11:594.
Liu Q, You B, Meng J, Huang CP, Dong G, Wang R, et al. Targeting the androgen receptor to enhance NK cell killing efficacy in bladder cancer by modulating ADAR2/circ_0001005/PD-L1 signaling. Cancer Gene Ther. 2022;29:1988–2000.
Ke H, Zhang J, Wang F, **ong Y. ZNF652-Induced circRHOT1 promotes SMAD5 expression to modulate tumorigenic properties and nature killer cell-mediated Ttoxicity in bladder cancer via targeting miR-3666. J Immunol Res. 2021;2021:7608178.
Li X, Han M, Zhang H, Liu F, Pan Y, Zhu J, et al. Structures and biological functions of zinc finger proteins and their roles in hepatocellular carcinoma. Biomark Res. 2022;10:2.
Heyliger SO, Soliman KFA, Saulsbury MD, Reams RR. The identification of zinc-finger protein 433 as a possible prognostic biomarker for clear-cell renal cell carcinoma. Biomolecules. 2021;11:1193.
Hu ZQ, Zhou SL, Li J, Zhou ZJ, Wang PC, **n HY, et al. Circular RNA sequencing identifies CircASAP1 as a key regulator in hepatocellular carcinoma metastasis. Hepatology. 2020;72:906–22.
Han S, Wang W, Wang S, Yang T, Zhang G, Wang D, et al. Tumor microenvironment remodeling and tumor therapy based on M2-like tumor associated macrophage-targeting nano-complexes. Theranostics. 2021;11:2892–916.
Niedobitek G, Barros MH, Dreyer JH, Hauck F, Al-Sheikhyaqoob D. Tumor-associated macrophages: function and differentiation. Pathologe. 2015;36:477–84.
Wu K, Lin K, Li X, Yuan X, Xu P, Ni P, et al. Redefining tumor-associated macrophage subpopulations and functions in the tumor microenvironment. Front Immunol. 2020;11:1731.
Lu JC, Zhang PF, Huang XY, Guo XJ, Gao C, Zeng HY, et al. Amplification of spatially isolated adenosine pathway by tumor-macrophage interaction induces anti-PD1 resistance in hepatocellular carcinoma. J Hematol Oncol. 2021;14:200.
Wang X, Sheng W, Xu T, Xu J, Gao R, Zhang Z. CircRNA hsa_circ_0110102 inhibited macrophage activation and hepatocellular carcinoma progression via miR-580-5p/PPARalpha/CCL2 pathway. Aging (Albany NY). 2021;13:11969–87.
Cao P, Ma B, Sun D, Zhang W, Qiu J, Qin L, et al. hsa_circ_0003410 promotes hepatocellular carcinoma progression by increasing the ratio of M2/M1 macrophages through the miR-139-3p/CCL5 axis. Cancer Sci. 2022;113:634–47.
Wang Y, Gao R, Li J, Tang S, Li S, Tong Q, et al. Downregulation of hsa_circ_0074854 suppresses the migration and invasion in hepatocellular carcinoma via interacting with HuR and via suppressing exosomes-mediated macrophage M2 polarization. Int J Nanomed. 2021;16:2803–18.
Hume DA, MacDonald KP. Therapeutic applications of macrophage colony-stimulating factor-1 (CSF-1) and antagonists of CSF-1 receptor (CSF-1R) signaling. Blood. 2012;119:1810–20.
Lin C, ** Y, Yu H, Wang Z, Chen X, Shen W. circRNA TCFL5 promote esophageal cancer progression by modulating M2 macrophage polarization via the miR-543-FMNL2 axis. J Oncol. 2022;2022:5075615.
Lu Q, Wang X, Zhu J, Fei X, Chen H, Li C. Hypoxic tumor-derived exosomal Circ0048117 facilitates M2 macrophage polarization acting as miR-140 sponge in esophageal squamous cell carcinoma. Onco Targets Ther. 2020;13:11883–97.
Yi B, Dai K, Yan Z, Yin Z. Circular RNA PLCE1 promotes epithelial mesenchymal transformation, glycolysis in colorectal cancer and M2 polarization of tumor-associated macrophages. Bioengineered. 2022;13:6243–56.
Chen B, Wu L, Tang X, Wang T, Wang S, Yu H, et al. Quercetin inhibits tumorigenesis of colorectal cancer through downregulation of hsa_circ_0006990. Front Pharm. 2022;13:874696.
Chen W, Tang D, Lin J, Huang X, Lin S, Shen G, et al. Exosomal circSHKBP1 participates in non-small cell lung cancer progression through PKM2-mediated glycolysis. Mol Ther Oncolytics. 2022;24:470–85.
Chen T, Liu Y, Li C, Xu C, Ding C, Chen J, et al. Tumor-derived exosomal circFARSA mediates M2 macrophage polarization via the PTEN/PI3K/AKT pathway to promote non-small cell lung cancer metastasis. Cancer Treat Res Commun. 2021;28:100412.
Gao J, Ao YQ, Zhang LX, Deng J, Wang S, Wang HK, et al. Exosomal circZNF451 restrains anti-PD1 treatment in lung adenocarcinoma via polarizing macrophages by complexing with TRIM56 and FXR1. J Exp Clin Cancer Res. 2022;41:295.
Hegde S, Leader AM, Merad M. MDSC: Markers, development, states, and unaddressed complexity. Immunity. 2021;54:875–84.
Bruger AM, Dorhoi A, Esendagli G, Barczyk-Kahlert K, van der Bruggen P, Lipoldova M, et al. How to measure the immunosuppressive activity of MDSC: assays, problems and potential solutions. Cancer Immunol Immunother. 2019;68:631–44.
Katopodi T, Petanidis S, Domvri K, Zarogoulidis P, Anestakis D, Charalampidis C, et al. Kras-driven intratumoral heterogeneity triggers infiltration of M2 polarized macrophages via the circHIPK3/PTK2 immunosuppressive circuit. Sci Rep. 2021;11:15455.
**e T, Fu DJ, Li ZM, Lv DJ, Song XL, Yu YZ, et al. CircSMARCC1 facilitates tumor progression by disrupting the crosstalk between prostate cancer cells and tumor-associated macrophages via miR-1322/CCL20/CCR6 signaling. Mol Cancer. 2022;21:173.
Gunassekaran GR, Poongkavithai Vadevoo SM, Baek MC, Lee B. M1 macrophage exosomes engineered to foster M1 polarization and target the IL-4 receptor inhibit tumor growth by reprogramming tumor-associated macrophages into M1-like macrophages. Biomaterials. 2021;278:121137.
Zheng Y, Ren S, Zhang Y, Liu S, Meng L, Liu F, et al. Circular RNA circWWC3 augments breast cancer progression through promoting M2 macrophage polarization and tumor immune escape via regulating the expression and secretion of IL-4. Cancer Cell Int. 2022;22:264.
Li H, Luo F, Jiang X, Zhang W, **ang T, Pan Q, et al. CircITGB6 promotes ovarian cancer cisplatin resistance by resetting tumor-associated macrophage polarization toward the M2 phenotype. J Immunother Cancer. 2022;10:e004029.
Huang X, Wang J, Guan J, Zheng Z, Hao J, Sheng Z, et al. Exosomal Circsafb2 resha** tumor environment to promote renal cell carcinoma progression by mediating M2 macrophage polarization. Front Oncol. 2022;12:808888.
Pan Z, Zhao R, Li B, Qi Y, Qiu W, Guo Q, et al. EWSR1-induced circNEIL3 promotes glioma progression and exosome-mediated macrophage immunosuppressive polarization via stabilizing IGF2BP3. Mol Cancer. 2022;21:16.
Lu Q, Yin H, Deng Y, Chen W, Diao W, Ding M, et al. circDHTKD1 promotes lymphatic metastasis of bladder cancer by upregulating CXCL5. Cell Death Disco. 2022;8:243.
Shang A, Gu C, Wang W, Wang X, Sun J, Zeng B, et al. Exosomal circPACRGL promotes progression of colorectal cancer via the miR-142-3p/miR-506-3p- TGF-β1 axis. Mol Cancer. 2020;19:117.
Liu X, Zhao S, Sui H, Liu H, Yao M, Su Y, et al. MicroRNAs/LncRNAs modulate MDSCs in tumor microenvironment. Front Oncol. 2022;12:772351.
Hu C, **a R, Zhang X, Li T, Ye Y, Li G, et al. CircFARP1 enables cancer-associated fibroblasts to promote gemcitabine resistance in pancreatic cancer via the LIF/STAT3 axis. Mol Cancer. 2022;21:24.
Yang K, Zhang J, Bao C. Exosomal circEIF3K from cancer-associated fibroblast promotes colorectal cancer (CRC) progression via miR-214/PD-L1 axis. BMC Cancer. 2021;21:933.
Gu C, Lu H, Qian Z. Matrine reduces the secretion of exosomal circSLC7A6 from cancer-associated fibroblast to inhibit tumorigenesis of colorectal cancer by regulating CXCR5. Biochem Biophys Res Commun. 2020;527:638–45.
Liu G, Sun J, Yang ZF, Zhou C, Zhou PY, Guan RY, et al. Cancer-associated fibroblast-derived CXCL11 modulates hepatocellular carcinoma cell migration and tumor metastasis through the circUBAP2/miR-4756/IFIT1/3 axis. Cell Death Dis. 2021;12:260.
Hsu MT, Coca-Prados M. Electron microscopic evidence for the circular form of RNA in the cytoplasm of eukaryotic cells. Nature. 1979;280:339–40.
Acknowledgements
We thank all of the members from the Han and Zhou laboratory for their critical comments and helpful suggestions. We apologize for being unable to cite many important papers in this field due to space limitations. We thank Medjaden Inc. for scientific editing of this manuscript.
Funding
This study was supported in part by the Key Science and Technology Program of Henan Province (no. 222102310098 to L.G.), the National Natural Science Foundation of China (nos. 82002731 and 82172891 to T.H., no. 82173022 to Q.H., no. 82273098 to X.Z., no. 82002731 to the Article Publishing Charge), the Henan National Science Fund for Excellent Young Scholars (no. 212300410067 to T.H.), the Doctoral Foundation of ** Guan, Qian Hao.
Authors and Affiliations
Contributions
Conceptualization, W.R., T.H. and L.G.; writing—original draft preparation, T.H., L.G., and X.Z.; writing—review and editing, L.G, Q.H., B.G., and M.W.; visualization, L.G. and Q.H.; supervision, W.R., T.H., and X.Z.; funding acquisition, T.H., L.G., Q.H., and X.Z. All authors have read the final version of the manuscript and agreed to its publication.