Abstract
Background
Histone deacetylases (HDACs) have been shown to be involved in tumorigenesis, but their precise role and molecular mechanisms in gastric cancer (GC) have not yet been fully elucidated.
Methods
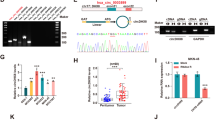
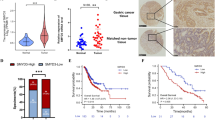
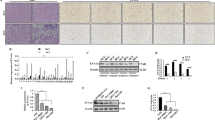
Bioinformatics screening analysis, qRT-PCR, and immunohistochemistry (IHC) were used to identify the expression of HDAC4 in GC. In vitro and in vivo functional assays illustrated the biological function of HDAC4. RNA-seq, GSEA pathway analysis, and western blot revealed that HDAC4 activated p38 MAPK signalling. Immunofluorescence, western blot, and IHC verified the effect of HDAC4 on autophagy. ChIP and dual-luciferase reporter assays demonstrated that the transcriptional regulation mechanism of HDAC4 and ATG4B.
Results
HDAC4 is upregulated in GC and correlates with poor prognosis. In vitro and in vivo assays showed that HDAC4 contributes to the malignant phenotype of GC cells. HDAC4 inhibited the MEF2A-driven transcription of ATG4B and prevented MEKK3 from p62-dependent autophagic degradation, thus activating p38 MAPK signalling. Reciprocally, the downstream transcription factor USF1 enhanced HDAC4 expression by regulating HDAC4 promoter activity, forming a positive feedback loop and continuously stimulating HDAC4 expression and p38 MAPK signalling activation.
Conclusion
HDAC4 plays an oncogenic role in GC, and HDAC4-based targeted therapy would represent a novel strategy for GC treatment.
Similar content being viewed by others
Background
Gastric cancer (GC) is now the fifth most common malignant cancer globally, and the number of GC cases in China accounts for >40% of all new cases of GC in the world [1]. Most patients have a definite diagnosis of GC only in the advanced phase and miss the chance to undergo radical surgical treatment [2]. The prognosis of patients with GC remains poor. Therefore, it has become an urgent need to conduct in-depth research on the pathogenesis of GC and to identify effective therapeutic targets.
Mitogen-activated protein kinase (MAPK) is a class of serine/threonine protein kinases that can be activated by various intracellular and extracellular stimuli, including growth factors, hormones, oxidative stress, and endoplasmic reticulum stress [3]. The MAPK signal transduction pathway consists of three types of sequentially activated protein members: MAP kinase kinase kinase (MAPKKK or MEKK), MAP kinase kinase (MAPKK or MEK), and MAPK, which play a role in enhancing the expression of target genes or directly acting on cytoplasmic downstream kinases, regulating cell proliferation, differentiation, stress response and cell apoptosis, and other physiological activities [4]. Each MEK can be activated by at least one MEKK, and each MAPK can be activated by different MEKs, forming a complex regulatory network of MAPK [5, 6]. MAPK consists of three main subgroups: extracellular signal-regulated kinase (ERK), c-Jun amino-terminal kinase (JNK), and p38 [7, 8]. Abnormal expression or overexpression of MAPK members plays an important role in the malignant transformation and evolution of cells.
Histone deacetylases (HDACs) are a hotspot in the field of cancer drug development. Inhibition of histone deacetylation has become a recognised approach for tumour therapy [9,10,11]. HDACs are involved in the regulation of tumour proliferation, invasion, and migration [12, 13]. Until now, 18 HDAC subtypes have been found in the human body, which can be further subdivided into four categories: Class I HDACs (HDAC1–3 and 8) mainly exist in the nucleus, and their main function is the deacetylation of histones. Class II HDACs are further divided into Class IIA (HDAC4, 5, 7, and 9) and Class IIB (HDAC6 and 10). Class IV HDAC11 is only expressed in the brain, kidney, and testes. Class III HDAC (SIRT1–7) is associated with the yeast protein SIR2 [14, 15]. Different types of HDACs have different structures, and their effects are also different. Different subtypes of HDACs also have great differences in their baseline expression levels as well as the mechanism of action in different tumour tissues [16,17,18]. The role of HDACs in GC development has been studied but the mechanisms are inadequately understood [19, 29]. We demonstrate the same results using real clinical data. Kang et al. found that HDAC4 promotes GC progression via p21 repression [30]. We found a different mechanism: HDAC4 facilitates the progression of GC mainly by activating the p38 MAPK pathway. Our results suggest that high expression of HDAC4 may be a poor predictor of GC.
As an important pathway of intracellular protein degradation, the autophagy-lysosomal system plays an important role in both nutrient cycling and scavenging and maintenance of stability [31]. Target proteins degraded by the autophagy-lysosomal system, such as WNT and Keap1, first bind to the key autophagy protein p62/LC3B and are recognised by the receptor proteins [32, 33]. Then they are wrapped by the autophagosome with a bilayer membrane structure, after which they enter the autophagy lysosomes to complete the autophagic degradation of the proteins. We found that HDAC4 knockdown enhances the autophagic degradation of MEKK3 and reduces the expression of MEKK3 in cells, thus inhibiting the activation of the MAPK pathway and the proliferation, migration, and invasion of GC cells.
HDAC4 plays different roles by regulating autophagy. After HDAC4 interacts with autophagy-related microtubule-associated protein 1S (MAP1s), the acetylation level of MAP1s decreases, and it becomes unstable. This inhibits autophagy and promotes the accumulation of MHTT aggregates, causing the occurrence of Huntington’s disease. The polyamine spermidine can improve MAP1s instability induced by HDAC4 and inhibit the occurrence of cirrhosis and hepatocellular carcinoma by promoting autophagy [34]. In diabetic nephropathy, HDAC4 promotes the deacetylation of signal transduction and transcriptional activator 1 (STAT1), and activated STAT1 inhibits podocyte autophagy, thereby inducing podocyte injury [35, 36]. However, during vascular inflammation, the increased expression of HDAC4 can reduce the acetylation of FoxO3a in vascular endothelial cells, and activated FoxO3a can promote the transcription of autophagy-related genes ATG5 and LC3B, thereby inducing the autophagy of vascular endothelial cells [37]. In our study, HDAC4 inhibited the transcription of the autophagy-related gene ATG4B and consequently autophagy in GC.
Traditional HDACs contain the amino acid tyrosine in their enzyme active region; however, for type II HDACs, the tyrosine is replaced by histidine, so that their activity is >1000 times lower than that of type I HDACs. Class II HDACs have a type of protein structure that has a specific amino acid sequence targeting the acetyl modification of lysine and can recruit HDAC3. HDAC3 can perform the deacetylase activity in case of class II HDAC deletion and can continue to bind to the NCoR/SMRT transcription co-inhibitory complex, remove the acetyl groups of histones and non-histone proteins, and inhibit DNA transcription [38]. Non-histone proteins studied in recent years mainly include runt-associated transcription factor 2, hypoxic-inducible factor-1α, and STAT1 [39,40,41]. Class II HDACs can also bind to transcription factors such as MEF2s, thereby inhibiting the transcription of genes regulated by these transcription factors [42]. Our study also confirmed that HDAC4 in GC cells inhibits the expression level of ATG4B by inhibiting the effect of MEF2A on the transcription of ATG4B, thus inhibiting the autophagy of GC cells.
In conclusion, our study confirmed that HDAC4 plays an important role in the development of GC, and high HDAC4 expression can be used as an independent predictor of poor prognosis of GC. High expression of HDAC4 inhibits the transcriptional activity of MEF2A, which in turn inhibits the transcription of ATG4B, thereby inhibiting the autophagy of GC cells, reducing the degradation of MEKK3, activating p38, and promoting the growth and metastasis of GC. Therefore, HDAC4 can be used as a new potential GC therapeutic target.
Data availability
The data that support the findings of this study are available on request from the corresponding author.
References
Yan C, Zhu M, Ding Y, Yang M, Wang M, Li G, et al. Meta-analysis of genome-wide association studies and functional assays decipher susceptibility genes for gastric cancer in Chinese populations. Gut. 2020;69:641–51.
Ahn JR, Jung M, Kim C, Hong MH, Chon HJ, Kim HR, et al. Prognosis of pN3 stage gastric cancer. Cancer Res Treat. 2009;41:73–9.
Guo YJ, Pan WW, Liu SB, Shen ZF, Xu Y, Hu LL. ERK/MAPK signalling pathway and tumorigenesis (Review). Exp Ther Med. 2020;19:1997–2007.
Kim EK, Choi EJ. Pathological roles of MAPK signaling pathways in human diseases. Biochim Biophys Acta. 2010;1802:396–405.
Fan X, Wang C, Shi P, Gao W, Gu J, Geng Y, et al. Platelet MEKK3 regulates arterial thrombosis and myocardial infarct expansion in mice. Blood Adv. 2018;2:1439–48.
Deacon K, Blank JL. MEK kinase 3 directly activates MKK6 and MKK7, specific activators of the p38 and c-Jun NH2-terminal kinases. J Biol Chem. 1999;6:16604–10.
Lee S, Rauch J, Kolch W. Targeting MAPK signaling in cancer: mechanisms of drug resistance and sensitivity. Int J Mol Sci. 2020;21:1102.
Cicenas J, Zalyte E, Rimkus A, Dapkus D, Noreika R, Urbonavicius S. JNK, p38, ERK, and SGK1 Inhibitors in Cancer. Cancers. 2017;10:1.
Camphausen K, Tofilon PJ. Inhibition of histone deacetylation: a strategy for tumor radiosensitization. J Clin Oncol. 2007;25:4051–6.
Huang Y, Vasilatos SN, Boric L, Shaw PG, Davidson NE. Inhibitors of histone demethylation and histone deacetylation cooperate in regulating gene expression and inhibiting growth in human breast cancer cells. Breast Cancer Res Treat. 2012;131:777–89.
Relles D, Chipitsyna GI, Gong Q, Yeo CJ, Arafat HA. Thymoquinone promotes pancreatic cancer cell death and reduction of tumor size through combined inhibition of histone deacetylation and induction of histone acetylation. Adv Prev Med. 2016;2016:1407840.
Li Y, Wang K, Wei Y, Yao Q, Zhang Q, Qu H, et al. lncRNA-MIAT regulates cell biological behaviors in gastric cancer through a mechanism involving the miR-29a-3p/HDAC4 axis. Oncol Rep. 2017;38:3465–72.
Wang X, Chen X, Tian Y, Jiang D, Song Y. Long noncoding RNA RGMB-AS1 acts as a microRNA-574 sponge thereby enhancing the aggressiveness of gastric cancer via HDAC4 upregulation. Onco Targets Ther. 2020;13:1691–704.
Seto E, Yoshida M. Erasers of histone acetylation: the histone deacetylase enzymes. Cold Spring Harb Perspect Biol. 2014;6:a018713.
Haberland M, Montgomery RL, Olson EN. The many roles of histone deacetylases in development and physiology: implications for disease and therapy. Nat Rev Genet. 2009;10:32–42.
Wang YF, Liu F, Sherwin S, Farrelly M, Yan XG, Croft A, et al. Cooperativity of HOXA5 and STAT3 is critical for HDAC8 inhibition-mediated transcriptional activation of PD-L1 in human melanoma cells. J Invest Dermatol. 2018;138:922–32.
Yu Y, Chen L, Zhao G, Li H, Guo Q, Zhu S, et al. RBBP8/CtIP suppresses P21 expression by interacting with CtBP and BRCA1 in gastric cancer. Oncogene. 2020;39:1273–89.
Wang W, Zhao M, Cui L, Ren Y, Zhang J, Chen J, et al. Characterization of a novel HDAC/RXR/HtrA1 signaling axis as a novel target to overcome cisplatin resistance in human non-small cell lung cancer. Mol Cancer. 2020;19:134.
Li Y, Zhang M, Dorfman RG, Pan Y, Tang D, Xu L, et al. SIRT2 promotes the migration and invasion of gastric cancer through RAS/ERK/JNK/MMP-9 pathway by increasing PEPCK1-related metabolism. Neoplasia. 2018;20:745–56.
**ong K, Zhang H, Du Y, Tian J, Ding S. Identification of HDAC9 as a viable therapeutic target for the treatment of gastric cancer. Exp Mol Med. 2019;51:1–15.
Schader T, Lowe O, Reschke C, Malacarne P, Hahner F, Muller N, et al. Oxidation of HDAC4 by Nox4-derived H2O2 maintains tube formation by endothelial cells. Redox Biol. 2020;36:101669.
Fu Y, Hong L, Xu J, Zhong G, Gu Q, Gu Q, et al. Discovery of a small molecule targeting autophagy via ATG4B inhibition and cell death of colorectal cancer cells in vitro and in vivo. Autophagy. 2019;15:295–311.
Tanc M, Cleenewerck M, Kurdi A, Roelandt R, Declercq W, De Meyer G, et al. Synthesis and evaluation of novel benzotropolones as Atg4B inhibiting autophagy blockers. Bioorg Chem. 2019;87:163–8.
Yu Z, Kong J, Pan B, Sun H, Lv T, Zhu J, et al. Islet-1 may function as an assistant factor for histone acetylation and regulation of cardiac development-related transcription factor Mef2c expression. PLoS ONE. 2013;8:e77690.
Lemercier C, Verdel A, Galloo B, Curtet S, Brocard MP, Khochbin S. mHDA1/HDAC5 histone deacetylase interacts with and represses MEF2A transcriptional activity. J Biol Chem. 2000;275:15594–9.
Zhu K, Wang H, Gul Y, Zhao Y, Wang W, Liu S, et al. Expression characterization and the promoter activity analysis of zebrafish hdac4. Fish Physiol Biochem. 2012;38:585–93.
Galibert MD, Carreira S, Goding CR. The Usf-1 transcription factor is a novel target for the stress-responsive p38 kinase and mediates UV-induced Tyrosinase expression. EMBO J. 2001;9:5022–31.
Wang Z, Qin G, Zhao TC. HDAC4: mechanism of regulation and biological functions. Epigenomics. 2014;2:139–50.
Spaety ME, Gries A, Badie A, Venkatasamy A, Romain B, Orvain C, et al. HDAC4 levels control sensibility toward cisplatin in gastric cancer via the p53-p73/BIK pathway. Cancers. 2019;11:1747.
Kang ZH, Wang CY, Zhang WL, Zhang JT, Yuan CH, Zhao PW, et al. Histone deacetylase HDAC4 promotes gastric cancer SGC-7901 cells progression via p21 repression. PLoS ONE. 2014;9:e98894.
**e C, Li N, Wang H, He C, Hu Y, Peng C, et al. Inhibition of autophagy aggravates DNA damage response and gastric tumorigenesis via Rad51 ubiquitination in response to H. pylori infection. Gut Microbes. 2020;11:1567–89.
Jiang T, Harder B, Rojo de la Vega M, Wong PK, Chapman E, Zhang DD. p62 links autophagy and Nrf2 signaling. Free Radic Biol Med. 2015;88:199–204.
Lorzadeh S, Kohan L, Ghavami S, Azarpira N. Autophagy and the Wnt signaling pathway: a focus on Wnt/beta-catenin signaling. Biochim Biophys Acta Mol Cell Res. 2021;1868:118926.
Yue F, Li W, Zou J, Jiang X, Xu G, Huang H, et al. Spermidine prolongs lifespan and prevents liver fibrosis and hepatocellular carcinoma by activating MAP1S-mediated autophagy. Cancer Res. 2017;77:2938–51.
Wang X, Liu J, Zhen J, Zhang C, Wan Q, Liu G, et al. Histone deacetylase 4 selectively contributes to podocyte injury in diabetic nephropathy. Kidney Int. 2014;86:712–25.
Wei Q, Dong Z. HDAC4 blocks autophagy to trigger podocyte injury: non-epigenetic action in diabetic nephropathy. Kidney Int. 2014;86:666–8.
Yang D, **ao C, Long F, Su Z, Jia W, Qin M, et al. HDAC4 regulates vascular inflammation via activation of autophagy. Cardiovasc Res. 2018;114:1016–28.
Bourque S, Jeandroz S, Grandperret V, Lehotai N, Aime S, Soltis DE, et al. The evolution of HD2 proteins in green plants. Trends Plant Sci. 2016;21:1008–16.
Byun Y, Choi YC, Jeong Y, Lee G, Yoon S, Jeong Y, et al. MiR-200c downregulates HIF-1alpha and inhibits migration of lung cancer cells. Cell Mol Biol Lett. 2019;24:28.
Tang M, Liu Y, Zhang QC, Zhang P, Wu JK, Wang JN, et al. Antitumor efficacy of the Runx2-dendritic cell vaccine in triple-negative breast cancer in vitro. Oncol Lett. 2018;16:2813–22.
Zhang J, Wang F, Liu F, Xu G. Predicting STAT1 as a prognostic marker in patients with solid cancer. Ther Adv Med Oncol. 2020;12:1758835920917558.
Clocchiatti A, Di Giorgio E, Ingrao S, Meyer-Almes FJ, Tripodo C, Brancolini C. Class IIa HDACs repressive activities on MEF2-depedent transcription are associated with poor prognosis of ER(+) breast tumors. FASEB J. 2013;27:942–54.
Funding
This work was supported by the National Natural Science Foundation of China (81672409), the Foundation of Nantong science and technology bureau (MSZ19204), the Foundation of Nantong science and technology bureau (MS12019026), and the Postgraduate Research and Practice Innovation Program of Jiangsu Province (Grant No. KYCX20_2844 and KYCX21_3113).
Author information
Authors and Affiliations
Contributions
W-JX and Y-ZZ designed the study; W-JZ and Y-LH carried out the experiments and prepared the manuscript; C-YQ and YF processed the data; J-ZL and J-LY performed statistical analysis; HH assisted in analysis and collection of clinical samples. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was approved by the Human Research Ethics Committee of the Affiliated Hospital of Nantong University (Nantong, China, approval 2020-L145), and each patient provided written informed consent. The study is compliant with all relevant ethical regulations involving human participants. Animal experiments were approved by Animal Center of Medical College of Nantong University.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zang, WJ., Hu, YL., Qian, CY. et al. HDAC4 promotes the growth and metastasis of gastric cancer via autophagic degradation of MEKK3. Br J Cancer 127, 237–248 (2022). https://doi.org/10.1038/s41416-022-01805-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-022-01805-7
- Springer Nature Limited
This article is cited by
-
ECHDC2 inhibits the proliferation of gastric cancer cells by binding with NEDD4 to degrade MCCC2 and reduce aerobic glycolysis
Molecular Medicine (2024)
-
β-hydroxybutyrate impairs nasopharyngeal carcinoma cell aggressiveness via histone deacetylase 4 inhibition
Molecular & Cellular Toxicology (2024)
-
Natural products targeting the MAPK-signaling pathway in cancer: overview
Journal of Cancer Research and Clinical Oncology (2024)
-
Ginsenoside Rh4 inhibits inflammation-related hepatocellular carcinoma progression by targeting HDAC4/IL-6/STAT3 signaling
Molecular Genetics and Genomics (2023)