Abstract
NRAS and KRAS activating point mutations are present in 10–30% of myeloid malignancies and are often associated with a proliferative phenotype. RAS mutations harbor allele-specific structural and biochemical properties depending on the hotspot mutation, contributing to variable biological consequences. Given their subclonal nature in most myeloid malignancies, their clonal architecture, and patterns of cooperativity with other driver genetic alterations may potentially have a direct, causal influence on the prognosis and treatment of myeloid malignancies. RAS mutations overall tend to be associated with poor clinical outcome in both chronic and acute myeloid malignancies. Several recent prognostic scoring systems have incorporated RAS mutational status. While RAS mutations do not always act as independent prognostic factors, they significantly influence disease progression and survival. However, their clinical significance depends on the type of mutation, disease context, and treatment administered. Recent evidence also indicates that RAS mutations drive resistance to targeted therapies, particularly FLT3, IDH1/2, or JAK2 inhibitors, as well as the venetoclax-azacitidine combination. The investigation of novel therapeutic strategies and combinations that target multiple axes within the RAS pathway, encompassing both upstream and downstream components, is an active field of research. The success of direct RAS inhibitors in patients with solid tumors has brought renewed optimism that this progress will be translated to patients with hematologic malignancies. In this review, we highlight key insights on RAS mutations across myeloid malignancies from the past decade, including their prevalence and distribution, cooperative genetic events, clonal architecture and dynamics, prognostic implications, and therapeutic targeting.
Similar content being viewed by others
Introduction
RAS proteins are a family of 21-kDa proteins that are at the heart of signaling pathways controlling various biological processes such as cell proliferation, differentiation, and survival. This family of proteins are specialized guanine nucleotide-binding and hydrolyzing molecules that belong to the small G-protein (GTP-ase) superfamily. They are encoded by highly related RAS genes, namely, KRAS (Kirsten rat sarcoma viral oncogene homolog), NRAS (neuroblastoma RAS viral oncogene homolog), and HRAS (Harvey rat sarcoma viral oncogene homolog), encoding 4 homologous proteins (sharing 85% sequence homology); H-RAS, K-RAS4A and K-RAS4B (two splice variants of K-RAS), and N-RAS [1]. Oncogenic mutations in RAS GTPases render the proteins constitutively GTP bound and active, promoting oncogenesis. However, the level of expression and activation of each specific RAS protein leads to different cellular responses and oncogenic phenotypes [2, 3]. Three well-studied RAS effectors are PI3-kinase (PI3K), Raf, and Ral-GDS proteins. Among these, the abnormal activation of the Raf/MEK/ERK pathway and the PI3K/Akt/mTOR cascade are strongly implicated in the development and maintenance of RAS-mutated cancers [4, 5].
RAS activating point mutations are found in nearly 20% of human cancers [5] and are highly prevalent in myeloid malignancies where they are often associated with a more proliferative phenotype [6, 7] and a more aggressive disease [8, 9]. While RAS mutation status has long been integrated into clinical decision making in patients with solid tumors, the clinical significance of RAS mutations in myeloid malignancies has only recently begun to be fully appreciated. Although considered as ‘undruggable’ in the past decade [10], significant progress in understanding RAS biology has brought us a step closer to identifying novel strategies for targeting RAS-mutated cancers, particularly in the context of myeloid malignancies.
This review aims to provide a detailed exploration of RAS mutations in myeloid malignancies including prevalent occurrences in acute myeloid leukemia (AML), myelodysplastic syndromes (MDS), chronic myelomonocytic leukemia (CMML), juvenile myelomonocytic leukemia (JMML), and myeloproliferative neoplasms (MPN). We address various challenges that have remained unanswered throughout the past decade. First, is how RAS mutations are not all equal; the type of the RAS mutated protein, the amino acid position, as well as the type of substitutions, varies across human cancers, including myeloid malignancies. Second, we decipher the cooperating genetic events with RAS mutations which modulate the resulting phenotype in mouse models. Third, we cover the clonal architecture and dynamics of RAS mutations. Lastly, we discuss variable prognostic implications depending on disease context, mutational type and the type of treatment administered. This highlights the challenge of RAS targeting therapies; due to their structural and biochemical properties, oncogenic RAS remains difficult targets for drug discovery.
Prevalence and type of oncogenic mutations
RAS mutations are prevalent in 10–30% of myeloid malignancies, with higher frequency in pediatric than adult diseases [11]. Among which, NRAS mutations are the most frequent, followed by KRAS mutations, whereas HRAS mutations are negligible in hematologic malignancies. The prevalence of N/KRAS mutations varies across different types of myeloid malignancies. According to recent studies using high-throughput sequencing technologies and covering the entire N/KRAS coding sequences (that is, exons 2, 3 and 4) [5, 12,13,14,15,16,17], MDS/MPN, notably CMML and JMML, harbor the highest incidence of N/KRAS mutations, ranging from 15% to 20% of cases for each gene. In adult MDS, both N- and KRAS mutations are identified in 2–3% of cases [15]. In AML, the overall prevalence of RAS mutations ranges between 15% and 20%. RAS mutations are particularly enriched in specific subsets of AML, such as AML with inv(3)/t(3;3) and AML with inv(16)/t(16;16), where these mutations are identified in around 30% and 35–40% of cases, respectively [15, 18,19,20]. In MPN, the prevalence of RAS mutations is very low in polycythemia vera (PV) and essential thrombocythemia (ET) (<1%), but can reach 6–8% in primary myelofibrosis (PMF) [21, 22] (Fig. 1).
Percentage of mutated cases based on recent studies using high-throughput sequencing technologies and covering the entire N/KRAS coding sequence. AML: 1105 patients [12, 16]; MDS: 2957 patients [15]; CMML: 1540 patients (399 patients [15] and 1141 patients from unpublished personal data; JMML: 117 patients [13, 14, 17]. AML acute myeloid leukemia, MDS myelodysplastic syndromes, CMML chronic myelomonocytic leukemia, JMML juvenile myelomonocytic leukemia.
Although more than 150 mutation sites have been reported in RAS genes, the most prevalent mutational hotspots are G12, G13, and Q61, accounting for approximately 80–95% of NRAS and 40–95% of KRAS mutations [5, 12,13,14,15,16,17]. NRAS and KRAS exhibit different hotspot preferences for G12, G13, Q61, and other non-canonical codons, such as T58, G60, K117, and A146, as illustrated in Fig. 2. Recent experimental evidence supports the notion that RAS mutations harbor allele-specific structural and biochemical properties, contributing to variability in biological consequences [23, 24].
Distribution of the most frequently mutated codons in CMML (n = 1540), AML (n = 1105), JMML (n = 117) and MDS (n = 2975). The color code of each hotspot mutation is indicated on the right of each pie chart. The data are derived from the same patient cohorts as in Fig. 1.
RAS mutation patterns vary across different types of myeloid malignancies and even across disease subtypes. In AML, NRAS mutations equally affect G12, G13, and Q61 codons, each of the 3 amino acids representing about one third of all mutations, while KRAS mutation distribution displays less Q61 mutations and more rare variants, such as those involving K117 and Q146 codons. In core binding factor (CBF) AML particularly in AML with inv(16)/t(16;16), codon Q61 is much more frequently mutated than other codons. In contrast, NPM1 mutations preferentially associate with NRAS G12/13 but not with NRAS Q61 mutations. In adult chronic myeloid disorders, including CMML and MDS, NRAS mutations are predominantly found at G12 codon, accounting for 50–70% of cases, while KRAS mutations show much more diversity in terms of amino acid positions. In JMML, more than 80% of N/KRAS mutations affect G12 and G13 codons (Fig. 2). Altogether, RAS mutation patterns in myeloid malignancies are likely shaped by quantitative and qualitative differences in the activation of downstream signaling pathways, as suggested in the “sweet spot” model proposed by Li et al. [3]. Although the underlying biological mechanisms of codon-preferential RAS mutations in specific types of myeloid malignancies remain poorly understood, it is likely that distinct mutagenesis and/or selection processes are involved in different clinical settings. Properly identifying and understanding the roles and nuances of different RAS mutations could potentially guide targeted therapies and ultimately improve patient outcomes.
Modeling Ras mutations and their cooperation with other gene mutations
Ras mouse models have been extensively employed in hematologic malignancies with the aim of conducting in vivo experiments for disease understanding and preclinical trials. Table 1 provides an updated compilation of the recent Ras mouse models and their respective phenotype. Distinct Ras mutations exhibit variable behavior, for instance, the induction of heterozygous KrasG12D/+ expression in the hematopoietic system alone through Mx1-Cre leads to a rapid and highly penetrant myeloproliferative disease (MPD) modeling human MDS/MPN, but does not lead to AML progression [25]. In parallel, induction of the heterozygous KrasA146T/+ mutation in the hematopoietic compartment also led to an MDS/MPN phenotype similar to KrasG12D mice, but with a significantly delayed onset [23]. In contrast, endogenous heterozygous NrasG12D expression exhibits a modest and variable myeloid phenotype, although mice that are homozygous for a conditional NrasG12D knock-in allele model aggressive MPN [26]. When expressed in the hematopoietic compartment, NrasG12D alone induces a MPD similar to KrasG12D but with significantly longer disease latency and lower penetrance [27,28,29,30]. These findings collectively suggest that discrepancies in the mutation type, and/or expression levels of distinct Ras proteins influence the severity of myeloid growth dysregulation [4]. The cooperation of Ras mutations with other genetic alterations, such as Tet2, Dnmt3a, or Tp53 mutations, has also been recently investigated in mouse models (Table 1). Complete Dnmt3a loss enhances self-renewal in hematopoietic stem cells (HSCs), impairs differentiation, but is not sufficient to drive leukemogenesis in mice; specific disease progression depends on additional genetic alterations, such as Ras mutations [31, 32] Thus, complete loss of Dnmt3a synergizes with KrasG12D, expediting disease progression and culminating in approximately 30% of mice develo** AML [30] (Table 1). Concurrently, NrasG12D, in conjunction with heterozygous Dnmt3a loss, promotes AML onset in one-third of the induced mice, providing a potentially more biologically pertinent representation given the prevalent heterozygosity of DNMT3A mutations in human disease. Alternatively, hotspot Dnmt3aR878H mutation with NrasG12D led to a much earlier onset in mice, shorter lifespan, and more severe AML-like disease [33]. This suggests that the type of DNMT3A mutation, along with acquisition of RAS mutations, could significantly promote the leukemogenic transformation and proliferation of HSCs.
Tet2−/− and NrasG12D in hematopoietic cells synergize in vivo, engendering a lethal CMML-like disease with elevated self-renewal potential compared to mice harboring either mutation alone. Upon acquisition of the Nras mutation, clonal expansion is observed, precipitating leukemia progression and heightened sensitivity to GM-CSF [34]. These findings were validated in the context of Tet2 haploinsufficiency and Ras mutations where they collaborate to disrupt hematopoietic stem and progenitor cells (HSPCs), inducing a lethal and significantly penetrant CMML-like disorder. The concurrent Nras and Tet2 mutations also evoke cytokine hypersensitivity in HSPCs [35].
In the context of NrasG12D/+ associated with p53 mutations, NrasG12D/+ x p53−/− mice developed mixed AML and T-cell malignancy, whereas NrasG12D/+; p53R172H/+ mice rapidly developed a lethal AML with full penetrance and a median survival of ~80 days. Additionally, NrasG12D/+; p53R172H/+ HSPCs show imbalanced myelopoiesis and lymphopoiesis. It has also been reported that mutant p53 and oncogenic Nras cooperatively dysregulate hematopoietic transcription factor networks and promote inflammation via NfkB [36]. This demonstrates that Nras mutations cooperate with p53 mutants to promote AML in a much more important manner than either mutation alone.
BCOR mutations have been identified in various hematologic malignancies including MDS and AML. Bcor inactivation in aged mice was not sufficient for leukemogenesis but was associated with a significant increase in the absolute number of bone marrow myeloid progenitors. In contrast, BcorKO mice in cooperation with KrasG12D/+ developed a leukemia-like phenotype (Table 1). Additionally, the survival of BcorKO;KrasG12D/+ mice was significantly reduced compared with KrasG12D/+ controls indicating that Bcor inactivation resulted in functional co-operation with oncogenic Kras to initiate leukemia in vivo [37].
CUX1 mutations are common in myeloid neoplasms and significantly co-occur with oncogenic mutations in RAS, PTPN11, or CBL [38]. In murine models, Cux1 deficiency gives rise to MDS-like phenotype but falls short of driving AML independently. However, mice bearing NrasG12D and Cux1 knockdown concurrently exhibited AML development, an outcome absent in mice with either mutation alone. The oncogenic influence of Ras drives an increase in self-renewal in Cux1-deficient HSPCs. Conversely, Cux1 knockdown intensifies Ras signaling by mitigating negative regulators of RAS/PI3K signaling. Table 1 describes the phenotype of the resulting Cux1low;NrasG12D mice, which mimic an AML-like disease compared to Cux1mid;NrasG12D mice which are more MDS/MPN similar to that of JMML/CMML, indicating that the further decrease of Cux1 expression drives a more penetrating phenotype in cooperation with NrasG12D to drive AML [39]. Of note, all double mutant mice in Table 1 have significantly reduced survival as compared to mice harboring each mutation alone.
Taken together, murine models in myeloid malignancies have resulted in highly significant advancements in understanding how Ras mutations serve as cooperating mutations with other disease-initiating mutations. While Ras mutations alone do contribute to a significant myeloproliferative phenotype, they require cooperation with other mutations, more particularly those of tumor suppressor genes to drive leukemogenesis. The above examples underscore that the specific type of the cooperating mutation, in conjunction with Ras mutations, can yield diverse pathologic outcomes.
Clonal architecture and evolution
The consequence of the type and order of mutation acquired leads to the HSC being more or less likely to facilitate subsequent acquisition of mutations and leukemia development. Recent research prompted inquiry into how such clones facilitate the acquisition of other mutations in signaling pathways, such as RAS, to enhance their clonal fitness [40, 41]. In the context of age-associated myeloid malignancies, RAS mutations tend to emerge exclusively in the context of other clonal hematopoiesis mutations suggesting that these late events may cooperate with founder mutations to drive the progression of clonal hematopoiesis toward malignancy, aligning with a stepwise model of leukemogenesis [40, 42].
In clonal hematopoiesis of indeterminate potential (CHIP), RAS mutations only occur secondarily in the presence of other mutations strongly correlated with the apparition of a hematologic malignancy. Unlike most age-associated MDS/MPN where RAS mutations are often observed to be subclonal, JMML is essentially a RASopathy arising through the acquisition of de novo signaling mutations or in the context of germline predisposition syndromes. Recent evidence suggests that very few somatic events are required for JMML leukemogenesis and confirmed the predominant role of RAS pathway alterations in disease initiation. RAS-activating mutations might have distinct effects on epigenome remodeling possibly correlated with disease aggressiveness [42,43,44,45]. CMML however, is a RASopathy of the elderly often found in a background of epigenetic and splicing alterations. In the context of AML, N/KRAS mutations may function as an early/initiating event but mostly as cooperating mutations acquired during disease progression. Regarding MDS and MPN, RAS mutations mainly appear as a late event, driving progression and transformation to AML.
Various studies have attempted to replicate the sequential addition of Ras mutations within different mutational contexts. For instance, HSCs acquiring Runx1::Runx1T1 gain a competitive advantage, which leads to an expansion in the number of HSCs, thereby increasing the pool of cells capable of acquiring additional mutations like Kras. Ultimately, this promotes the development of leukemia and mimics the disease phenotype in mice. Conversely, HSCs that only acquire a Kras mutation, whether alone or in combination with Runx1::Runx1T1, are depleted due to loss of quiescence and self-renewal. This observation may elucidate why signaling mutations like RAS are not typically detected in pre-leukemic HSCs in AML patients; they tend to manifest as a later event in the leukemogenesis process. Ras mutations may necessitate cooperation with other mutations to confer this particular phenotype and are insufficient to do so as a solitary mutation [46]. This leads to the conclusion that the timing of emergence of Ras mutations in the clonal evolution is vital for the cell’s fate to transformation. However, earlier studies argue that Ras mutations alone partially enhance competitiveness of the HSC and promote pre-leukemic clonal expansion. It has been reported that NrasG12D/+ has a bimodal effect on HSCs in mice, increasing self-renewal potential and reducing division in one HSC subset while increasing division and reducing self-renewal in another HSC subset. Short-lived but rapidly dividing NrasG12D/+ HSCs presumably outcompete wild-type HSCs and are replenished over time by quiescent NrasG12D/+ HSCs [47]. Given that heterogeneity within HSCs is likely governed by various mechanisms of gene expression control, epigenetics and RNA splicing, variations in methylation levels and patterns give rise to stochastic transcriptional heterogeneity among genetically identical cells which may or may not protect the cell from external stress and the potential of acquiring further mutations. This heterogeneity could also elucidate the differing outcomes observed when HSCs are transformed by the same oncogenic event such as N/KRAS mutations. This suggests that the expansion of an NRAS mutant clone may be contingent on a specific cellular state [48], or possibly a chromatin state depending on which epigenetic factors are mutated [40].
Dormancy may be another factor influencing the emergence of RAS subclones. Dormant HSCs are normally in a quiescent state and are resistant to acute stress, but chronic stress such as infections, metabolic stress, or cytokine-related inflammation can exhaust them. Leukemic HSC are reported to co-opt physiological mechanisms of HSC sustenance to overcome this exhaustion, dominating normal HSCs in the niche and rendering them more fit. Moreover, mutant HSCs such as Tet2−/− or Dnmt3a−/− are also reported to secrete IL1-β and IFN-γ, allowing mutated clones to outcompete non-mutated clones. The niche thus becomes predominantly mutated, giving rise to both dormant, and active HSCs, which are more susceptible to proliferative signals [48]. Taken together, this raises the following hypotheses: does the exposure of mutant HSCs to chronic stress lead to epigenetic modifications rendering the clone more susceptible to acquire a RAS mutation? Does the harsh inflammatory milieu lead to a selection pressure of the RAS-mutated clone to evolve and expand? Could this explain why in JMML, a single initiating driver event is sufficient to drive leukemogenesis? Perhaps the cellular state, and cytokine milieu in utero is favorable for the competitive phenotype of the mutation, and the environment was predisposed to an infection or inflammation which rendered the clone to expand.
The literature currently presents a contradictory perspective regarding whether RAS mutations can induce clonal expansion independently or if they require a predisposed mutational context to manifest. Nevertheless, the arrangement of clonal populations may potentially have a direct, causal influence on the prognosis and treatment of myeloid malignancies [49]. Conversely, it is plausible that clonal architecture and the microenvironment might serve as a surrogate for an underlying process that itself contributes to chemoresistance or relapse. This highlights the need for more comprehensive research of the time-dependent consequences of Ras mutation emergence in the clonal architecture of leukemogenesis. Novel knock-in models facilitating the sequential introduction of mutations hold significant promise for future advancements. Additionally, deciphering the unique molecular signatures linked to pre-leukemic mutations in HSCs could pave the way for potential therapeutic advances aimed at selectively targeting the expansion of preleukemic stem cells. Exploring evolutionary dynamics through single-cell technologies and mathematical modeling holds the potential to enhance our comprehension of leukemic transformation and treatment resistance. This approach may also pave the way for the development of innovative therapeutic strategies and the identification of valuable biomarkers.
Prognostic implications
N/KRAS mutations are significant contributors to the pathogenesis, progression, and often prognosis of myeloid malignancies. They are quite infrequent in the context of CHIP, with relatively low variant allele frequencies at 1% and 2%, respectively. The late emergence of a RAS-mutated clone however, is associated with a 12-fold elevated risk of develo** a myeloid malignancy (Table 2). While further research is needed to strengthen this finding, it implies that individuals harboring a RAS-mutated CHIP clone require careful clinical monitoring due to their high-risk profile [40].
In MDS, RAS mutations are correlated with more aggressive disease subtypes, higher IPSS-M risk, and reduced event-free survival (EFS) and overall survival (OS). RAS-mutated MDS patients are also reported to have an OS of only 16 months versus 92 months in non-RAS-mutated MDS patients [8]. This has been validated in a separate cohort where patients have an increased risk of leukemic transformation, primarily associated with NRAS rather than KRAS mutations [15] (Table 2). One hypothesis is that the rarer occurrence of KRAS mutations may make their prognostic impact more challenging to determine. These two mutations may possess distinct biochemical properties and functional consequences giving rise to distinct prognostic implications. Nevertheless, the presence of both NRAS and KRAS mutations appears to exert a substantial toll on OS, as supported by various independent studies [50,51,52,53]. This emphasizes the importance of screening for RAS mutations both at diagnosis and during follow-up, enabling the identification of high-risk patients and the personalization of therapeutic strategies. N/KRAS mutations do not seem to influence responses to anthracycline-based chemotherapies, as observed in AML. Knowledge is scarcer regarding their role in responses to hypomethylating agents and combination therapies such as azacitidine (AZA) and venetoclax (Ven). The elusive nature of their prognostic relevance in this context may be due to the low frequency of RAS-mutated patients in princeps studies and a lack of dedicated investigations [8, 54, 55].
In CMML, RAS mutations are more prevalent especially in the proliferative form of the disease, at around 20–30% [56]. While RAS mutations appear to play a pivotal role in the transformation of CMML to AML, only NRAS mutations seem to exhibit a significant association with adverse clinical outcomes and are included in dedicated scores such as CPSS-Mol score, as well as the CMML transplant score. NRAS-mutated CMML patients encounter reduced response rates to HMA and allogeneic hematopoietic stem cell transplantation (allo-SCT), resulting in higher relapse rates and ultimately shorter OS (Table 2) [57,58,59]. Unlike NRAS, KRAS mutations are only represented in the IPSS-M score. Although conducted on a very large patient cohort, the IPSS-M score predominantly encompasses MDS but also, to a lesser extent, CMML and other MDS/MPN. While both N/KRAS mutations increase the risk of acute transformation in CMML, only mutant NRAS has been conclusively shown to influence EFS and OS in CMML patients.
In PMF, RAS mutations remain infrequent but are associated with higher bone marrow cellularity, increased splenomegaly, elevated circulating blast percentages, and additional driver mutations. However, in multivariate analysis, RAS mutations are not retained as prognostic factors for acute transformation independently of well-established markers, such as high-risk cytogenetic abnormalities and other alterations such as mutations in ASXL1, EZH2, SRSF2, IDH1/2, or U2AF1 [21, 22, 60].
Despite their rarity, N/KRAS mutations in PMF are associated with reduced responses to ruxolitinib, necessitating the monitoring of RAS mutational status for all PMF patients. This assessment may become a routine part of monitoring to guide therapeutic decision-making. JAK2 inhibitor therapies in PMF are primarily symptom-focused and have limited impact on bone marrow fibrosis and mutation allele burden. A recent study reported that the presence of RAS and CBL mutations was linked to poorer symptom improvement and spleen size reduction, suggesting potential resistance to JAK inhibitors. This resistance may stem from two mechanisms: one study showed that a RAS mutation acquired within a JAK2V617F mutated clone confers resistance to JAK inhibition, while another study highlighted PDGF-BB’s role in maintaining MEK/ERK activation in the presence of ruxolitinib (Table 2) [21, 22, 61, 62].
In adult AML, RAS mutations do not appear to significantly influence survival in patients subjected to intensive anthracycline-based chemotherapy, and accordingly have not been included in the European LeukemiaNet genetic risk classifications [63, 64]. However, emerging evidence suggests that RAS mutations may hold prognostic significance in AML patients treated with non-intensive therapies. Indeed, RAS mutations have been associated with higher relapse risk post-HMA treatment, such as AZA, or the Ven-AZA combination [54, 65,66,67,68]. KRAS but not NRAS mutations were also found to be associated with inferior survival in AML, particularly in the context of HMA-based therapies (Table 2) [69]. Furthermore, a recent study validated a new molecular prognostic risk signature, called mPRS, tailored for AML patients treated with HMA and Ven. This mPRS, based on the mutational status of 4 genes (NRAS, KRAS, FLT3, and TP53), can accurately segregate 3 groups of AML patients with distinct outcomes. Notably, N/KRAS mutations appear to negatively impact patient outcomes [70].
In JMML, a subset of RAS-mutated cases, combined with favorable prognostic factors; normal fetal hemoglobin levels for age and high platelet counts, have long-term survival without the need for allo-SCT. However, NRAS mutations in JMML are often associated with higher relapse rates, warranting adjusted post-transplant treatment strategies, including low-intensity graft versus host disease (GVHD) prophylaxis to enhance the graft versus leukemia (GVL) effect and reduce the risk of relapse. Conversely, KRAS-mutated JMML exhibit lower relapse rates, necessitating classical high-intensity GVHD prophylaxis (Table 2).
In pediatric AML, there is limited data regarding the potential influence of RAS mutations on clinical outcomes. The frequently altered tyrosine kinase and RAS/MAPK/MEK pathways, identified in 30-90% of pediatric AML patients, contributes to around 20% of relapses in this group [71, 72]. The prognostic impact of RAS mutations in pediatric AML has not been systematically investigated, but RAS-mutated pediatric AML seem to exhibit greater chemosensitivity compared to non-RAS-mutated AML [73, 74].
RAS mutations appear to negatively influence treatment responses following non-intensive therapies. In AML, RAS mutational status plays a pivotal role in the response to FLT3 inhibitors. For instance, RAS/MAPK pathway mutations emerge in approximately one third of AML patients experiencing disease progression on gilteritinib therapy [75]. A parallel study on crenolanib therapy in relapsed/refractory FLT3-mutated AML also identified epigenetic and genetic alterations, including NRAS mutations, associated with resistance. This resistance may be due to mutant RAS facilitating downstream ERK signaling reactivation in the presence of FLT3 inhibitors. RAS mutations also affect responses to venetoclax therapy by activating the Ras/Raf/MEK/ERK pathway, leading to increased MCL-1 compared to BCL2, thereby conferring resistance to BCL2 inhibitors [67, 68]. Collectively, this emphasizes the importance of early monitoring for RAS mutations upon initiating FLT3 inhibitor therapy, which could provide a crucial window for proactive intervention. It also suggests that targeting both MCL1 and BCL2 with venetoclax could be an alternative approach [9, 76].
In the context of IDH inhibitors such as ivosidenib and enasidenib, it is established that the existence of a RAS co-mutation is linked to inherent [77], but also acquired resistance [78]. Several hypotheses that might explain this resistance include the potent oncogenic signals of RAS activation diminishing 2-HG dependency and the contribution of RAS pathway-activating mutations to a sustained differentiation block following drug initiation. RAS mutations may also activate alternative pathways, change cellular metabolism, and induce epigenetic alterations, all of which may lead to resistance against IDH inhibitors by promoting cell survival and reducing drug sensitivity [77, 79,80,81].
Altogether, the impact of RAS mutations is far from uniform, with its significance heavily contingent upon the specific hematologic malignancy and the treatment modalities employed. Nonetheless, the presence of a RAS mutation correlates with an increased relapse risk in patients receiving non-intensive or targeted therapies.
RAS targeting therapeutic strategies
Over the last decade, significant progress has been made in the development of targeted therapies in myeloid malignancies. However, molecularly targeted therapies with clinical efficacy are still lacking for RAS-mutant myeloid malignancies. The investigation of novel therapeutic strategies and combinations targeting the RAS pathway, encompassing both upstream and downstream components, is an active field of research.
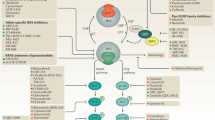
MEK, the downstream effector of the RAS-MAPK pathway, has recently been the primary therapeutic focus. Trametinib, a MEK1/2 inhibitor, showed promise by inhibiting ERK phosphorylation, resulting in reduced proliferation of NRAS-mutated AML cells in preclinical studies [82, 83]. Other clinical trials of MEK inhibitors are currently ongoing including trametinib, cobimetinib, selumetinib, and binimetinib, in various hematologic malignancies (Table 3). A recent study also revealed that RAS pathway mutations are associated with a unique gene expression profile enriched in mitotic kinases, such as polo-like kinase 1 (PLK1). Pharmacologic inhibition of PLK1 in RAS mutant patient-derived xenografts yielded promising results [9]. Onvansertib, a PLK1 inhibitor, is currently undergoing phase 1 trials for relapsed/refractory RAS-mutated CMML patients (Table 3).
Novel inhibitor molecules are currently being explored in solid malignancies, offering future therapeutic promise. For example, KRASG12C inhibitors, such as sotorasib and adagrasib, have shown encouraging results in clinical trials of non-small cell lung cancer and colorectal cancer. Recent studies also investigated other inhibitory molecules such as MRTX1133 and JAB-23000 that are selective inhibitors of KRAS G12D and KRAS G12V, respectively. A phase 1 trial testing RMC-6236, a triple inhibitor of KRAS (G12V, G12D, G13C, G13D, Q61H), NRAS (Q61X) and HRAS mutants, is also underway, with optimistic outcomes [84, 85]. Nonetheless, the applicability of such molecules in the field of hematology has still been restricted due to various factors. First, given that specific NRAS inhibitors are not readily available for all mutation types, the relatively higher prevalence of NRAS mutations in myeloid malignancies as compared to solid tumors represents a limitation. Second, patients may harbor multiple subclones, each carrying a distinct RAS mutation. This genetic heterogeneity renders the therapeutic targeting of RAS mutations even more challenging. In this context, the use of inhibitors designed to target multiple mutations, such as RMC-6236, may be a more suitable approach. Such broad-spectrum inhibitors have the potential to address the diversity of RAS mutations and offer a more comprehensive strategy for tackling these genetic alterations in myeloid malignancies.
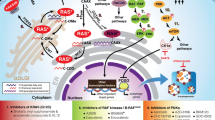
Beyond targeting the mutated RAS protein directly, alternative strategies aim to prevent its activation by inhibiting upstream signaling molecules such as SOS1. In addition, the development of PROTACs (Proteolysis-Targeting Chimeras), bi-functional molecules designed to induce proteasomal degradation of specific target proteins, are under investigation and have shown to be efficient in preclinical studies targeting RAS-mutant proteins [86,87,88,89]. RNA-based approaches, such as small interfering RNAs (siRNAs), are yet another strategy to silence the expression of mutated RAS at the mRNA level (Fig. 3). The siRNA inhibition strategy is a technological challenge, given the heterogeneous distribution of K/NRAS mutations in myeloid malignancies, and refers more towards ultra-personalized medicine than to a global management strategy [89].
The binding of growth factors to the tyrosine kinase receptor leads to its phosphorylation and the binding to the Grb2/Sos complex. RAS is controlled by a loop of an inactive, GDP-bound state and an active, GTP-bound state. Activation of RAS occurs by the binding Guanine Nucleotide Exchange Factor (GEF) proteins, including SOS, which initiate the exchange of GDP for GTP. The GTP-bound RAS activates a cascade mechanism of downstream signaling molecules including RAF and PI3K, which regulates different cellular functions such as cell proliferation, differentiation, and cell growth/survival. This figure summarizes the druggable pathways and targets in clinical trials or that are potential therapies for future use. Four critical therapeutic axes are highlighted: direct RAS inhibitors, MEK inhibitors, PI3K inhibitors and PLK1 inhibitors. Targets labeled in green are those currently in clinical trials in hematologic diseases. Targets labeled in blue are FDA-approved in the oncology field. Preclinical and clinical drugs targeting RAS-mutant in solid tumors are labeled respectively in black and red. RTK receptor tyrosine kinase, PROTACs proteolysis-targeting chimeras, siRNA small interfering RNA.
Besides targeting the RAS-mutated clones, the development of therapies targeting the inflammatory mediators may also be beneficial to improve survival, symptoms, and quality of life for patients with RAS-mutated myeloid malignancies. This has been recently illustrated in CMML where KRAS-mutated monocytes showed constitutive activation of the NLRP3 inflammasome, increased IL-1β release, and a specific inflammatory cytokine signature. Treatment of a CMML patient with a KRASG12D mutation using the IL-1 receptor blocker anakinra inhibited NLRP3 inflammasome activation, reduced monocyte count, and improved patient clinical status, allowing bridging to allo-SCT [90].
Given the fact that dormancy and the pro-inflammatory microenvironment of mutant-HSCs impact the subclonal emergence of RAS-mutated clones, it is likely that the proliferation of a RAS-mutated clone may be contingent upon a distinct cellular state and microenvironment [48], indicating that a more effective therapeutic strategy could involve targeting both the microenvironment and the RAS-mutated clone. This dual approach should be considered in the future to prevent the emergence of resistance and reduce the risk of relapse. In summary, several innovative therapies and strategies are being explored either in preclinical studies or in early clinical development in solid tumors (Fig. 3). The potential for these targeted therapies to transform the treatment of myeloid malignancies remains to be investigated in combination with other molecules.
Conclusion
This review highlights key insights on RAS mutations in myeloid malignancies from the past decade, encompassing their pivotal role in disease pathogenesis, prognosis, and therapy. While they may not always act as independent prognostic factors, they significantly influence clinical outcomes, disease progression, and survival. Different RAS proteins (NRAS vs. KRAS) may also differentially impact prognosis, in addition to their presence along concurrent mutations. Recent evidence indicates that RAS mutations also drive resistance to targeted therapies, especially FLT3, IDH1/2, or JAK2 inhibitors, as well as the venetoclax-azacitidine combination, necessitating early monitoring for intervention and exploring the clonal evolution of such subclones. While mouse models, despite limitations, offer vital platforms for studying Ras mutations and their interplay with other driver genetic alterations, our understanding of the intricate relationship between leukemic clones, the emergence of the RAS subclone, along the inflammatory microenvironment warrants further exploration. Advances in pharmacologic strategies have paved the way for potential therapeutic interventions targeting such mutations. Nonetheless, there is an uncharted territory regarding the application of solid tumor therapies to hematologic malignancies, promising novel trials in the future. The heterogeneity of RAS mutations emphasizes the need for personalized treatments and a meticulous screening of individual mutation profiles for effective therapeutic approaches.
References
Simanshu DK, Nissley DV, McCormick F. RAS Proteins and Their Regulators in Human Disease. Cell. 2017;170:17–33.
Haigis KM, Kendall KR, Wang Y, Cheung A, Haigis MC, Glickman JN, et al. Differential effects of oncogenic K-Ras and N-Ras on proliferation, differentiation and tumor progression in the colon. Nat Genet. 2008;40:600–8.
Li S, Balmain A, Counter CM. A model for RAS mutation patterns in cancers: finding the sweet spot. Nat Rev Cancer. 2018;18:767–77.
Ward AF, Braun BS, Shannon KM. Targeting oncogenic Ras signaling in hematologic malignancies. Blood. 2012;120:3397–406.
Prior IA, Hood FE, Hartley JL. The Frequency of Ras Mutations in Cancer. Cancer Res. 2020;80:2969–74.
Patnaik MM, Lasho T. Evidence-Based Minireview: Myelodysplastic syndrome/myeloproliferative neoplasm overlap syndromes: a focused review. Hematology. 2020;2020:460–4.
Kurata M, Antony ML, Noble-Orcutt KE, Rathe SK, Lee Y, Furuno H, et al. Proliferation and Self-Renewal Are Differentially Sensitive to NRASG12V Oncogene Levels in an Acute Myeloid Leukemia Cell Line. Mol Cancer Res. 2022;20:1646–58.
Ren Y, Lang W, Mei C, Luo Y, Ye L, Wang L, et al. Co‐mutation landscape and clinical significance of RAS pathway related gene mutations in patients with myelodysplastic syndrome. Hematological Oncol. 2023;41:159–66.
Carr RM, Vorobyev D, Lasho T, Marks DL, Tolosa EJ, Vedder A, et al. RAS mutations drive proliferative chronic myelomonocytic leukemia via a KMT2A-PLK1 axis. Nat Commun. 2021;12:2901.
Scott AJ, Lieu CH, Messersmith WA. Therapeutic Approaches to RAS Mutation. Cancer J. 2016;22:165–74.
Pikman Y, Stieglitz E. Targeting the Ras pathway in pediatric hematologic malignancies. Curr Opin Pediatrics. 2021;33:49–58.
Bottomly D, Long N, Schultz AR, Kurtz SE, Tognon CE, Johnson K, et al. Integrative analysis of drug response and clinical outcome in acute myeloid leukemia. Cancer Cell. 2022;40:850–864.e9.
Lipka DB, Witte T, Toth R, Yang J, Wiesenfarth M, Nöllke P, et al. RAS-pathway mutation patterns define epigenetic subclasses in juvenile myelomonocytic leukemia. Nat Commun. 2017;8:2126.
Caye A, Strullu M, Guidez F, Cassinat B, Gazal S, Fenneteau O, et al. Juvenile myelomonocytic leukemia displays mutations in components of the RAS pathway and the PRC2 network. Nat Genet. 2015;47:1334–40.
Bernard E, Tuechler H, Greenberg PL, Hasserjian RP, Arango Ossa JE, Nannya Y, et al. Molecular International Prognostic Scoring System for Myelodysplastic Syndromes. NEJM Evid. 2022;1. https://evidence.nejm.org/doi/10.1056/EVIDoa2200008.
The Cancer Genome Atlas Research Network. Genomic and Epigenomic Landscapes of Adult De Novo Acute Myeloid Leukemia. N Engl J Med. 2013;368:2059–74.
Stieglitz E, Taylor-Weiner AN, Chang TY, Gelston LC, Wang YD, Mazor T, et al. The genomic landscape of juvenile myelomonocytic leukemia. Nat Genet. 2015;47:1326–33.
Papaemmanuil E, Gerstung M, Bullinger L, Gaidzik VI, Paschka P, Roberts ND, et al. Genomic Classification and Prognosis in Acute Myeloid Leukemia. N. Engl J Med. 2016;374:2209–21.
Duployez N, Marceau-Renaut A, Boissel N, Petit A, Bucci M, Geffroy S, et al. Comprehensive mutational profiling of core binding factor acute myeloid leukemia. Blood. 2016;127:2451–9.
Itzykson R, Duployez N, Fasan A, Decool G, Marceau-Renaut A, Meggendorfer M, et al. Clonal interference of signaling mutations worsens prognosis in core-binding factor acute myeloid leukemia. Blood. 2018;132:187–96.
Coltro G, Rotunno G, Mannelli L, Mannarelli C, Fiaccabrino S, Romagnoli S, et al. RAS/CBL mutations predict resistance to JAK inhibitors in myelofibrosis and are associated with poor prognostic features. Blood Adv. 2020;4:3677–87.
Santos FPS, Getta B, Masarova L, Famulare C, Schulman J, Datoguia TS, et al. Prognostic impact of RAS-pathway mutations in patients with myelofibrosis. Leukemia. 2020;34:799–810.
Poulin EJ, Bera AK, Lu J, Lin YJ, Strasser SD, Paulo JA, et al. Tissue-Specific Oncogenic Activity of KRASA146T. Cancer Discov. 2019;9:738–55.
Hunter JC, Manandhar A, Carrasco MA, Gurbani D, Gondi S, Westover KD. Biochemical and Structural Analysis of Common Cancer-Associated KRAS Mutations. Mol Cancer Res. 2015;13:1325–35.
Chan IT, Kutok JL, Williams IR, Cohen S, Kelly L, Shigematsu H, et al. Conditional expression of oncogenic K-ras from its endogenous promoter induces a myeloproliferative disease. J Clin Invest. 2004;113:528–38.
Wang J, Liu Y, Li Z, Wang Z, Tan LX, Ryu MJ, et al. Endogenous oncogenic Nras mutation initiates hematopoietic malignancies in a dose- and cell type-dependent manner. Blood. 2011;118:368–79.
Wang J, Kong G, Liu Y, Du J, Chang YI, Tey SR, et al. NrasG12D/+ promotes leukemogenesis by aberrantly regulating hematopoietic stem cell functions. Blood. 2013;121:5203–7.
Chang YI, You X, Kong G, Ranheim EA, Wang J, Du J, et al. Loss of Dnmt3a and endogenous KrasG12D/+ cooperate to regulate hematopoietic stem and progenitor cell functions in leukemogenesis. Leukemia. 2015;29:1847–56.
Kurtz KJ, Conneely SE, O’Keefe M, Wohlan K, Rau RE. Murine Models of Acute Myeloid Leukemia. Front Oncol. 2022;12:854973.
Kurtz KJ, Conneely SE, O’Keefe M, Wohlan K, Rau RE. Corrigendum: Murine models of acute myeloid leukemia. Front Oncol. 2023;12:1089874.
Mayle A, Yang L, Rodriguez B, Zhou T, Chang E, Curry CV, et al. Dnmt3a loss predisposes murine hematopoietic stem cells to malignant transformation. Blood. 2015;125:629–38.
Challen GA, Sun D, Jeong M, Luo M, Jelinek J, Berg JS, et al. Dnmt3a is essential for hematopoietic stem cell differentiation. Nat Genet. 2011;44:23–31.
Shi X, Yang Y, Shang S, Wu S, Zhang W, Peng L, et al. Cooperation of Dnmt3a R878H with Nras G12D promotes leukemogenesis in knock-in mice: a pilot study. BMC Cancer. 2019;19:1072.
Kunimoto H, Meydan C, Nazir A, Whitfield J, Shank K, Rapaport F, et al. Cooperative Epigenetic Remodeling by TET2 Loss and NRAS Mutation Drives Myeloid Transformation and MEK Inhibitor Sensitivity. Cancer Cell. 2018;33:44–59.e8.
** X, Qin T, Zhao M, Bailey N, Liu L, Yang K, et al. Oncogenic N-Ras and Tet2 haploinsufficiency collaborate to dysregulate hematopoietic stem and progenitor cells. Blood Adv. 2018;2:1259–71.
Rajagopalan A, Feng Y, Gayatri MB, Ranheim EA, Klungness T, Matson DR, et al. A gain-of-function p53 mutant synergizes with oncogenic NRAS to promote acute myeloid leukemia in mice. J Clin Investig. 2023;133:e173116.
Kelly MJ, So J, Rogers AJ, Gregory G, Li J, Zethoven M, et al. Bcor loss perturbs myeloid differentiation and promotes leukaemogenesis. Nat Commun. 2019;10:1347.
Aly M, Ramdzan ZM, Nagata Y, Balasubramanian SK, Hosono N, Makishima H, et al. Distinct clinical and biological implications of CUX1 in myeloid neoplasms. Blood Adv. 2019;3:2164–78.
An N, Khan S, Imgruet MK, Jueng L, Gurbuxani S, McNerney ME. Oncogenic RAS promotes leukemic transformation of CUX1-deficient cells. Oncogene. 2023;42:881–93.
van Zeventer IA, de Graaf AO, Salzbrunn JB, Nolte IM, Kamphuis P, Dinmohamed A, et al. Evolutionary landscape of clonal hematopoiesis in 3,359 individuals from the general population. Cancer Cell. 2023;41:1017–1031.e4.
Bolton KL, Ptashkin RN, Gao T, Braunstein L, Devlin SM, Kelly D, et al. Cancer therapy shapes the fitness landscape of clonal hematopoiesis. Nat Genet. 2020;52:1219–26.
Deininger MWN, Tyner JW, Solary E. Turning the tide in myelodysplastic/myeloproliferative neoplasms. Nat Rev Cancer. 2017;17:425–40.
Niemeyer CM, Flotho C. Juvenile myelomonocytic leukemia: who’s the driver at the wheel? Blood. 2019;133:1060–70.
Fiñana C, Gómez-Molina N, Alonso-Moreno S, Belver L. Genomic and Epigenomic Landscape of Juvenile Myelomonocytic Leukemia. Cancers. 2022;14:1335.
Schönung M, Meyer J, Nöllke P, Olshen AB, Hartmann M, Murakami N, et al. International Consensus Definition of DNA Methylation Subgroups in Juvenile Myelomonocytic Leukemia. Clin Cancer Res. 2021;27:158–68.
Di Genua C, Norfo R, Rodriguez-Meira A, Wen WX, Drissen R, Booth CAG, et al. Cell-intrinsic depletion of Aml1-ETO-expressing pre-leukemic hematopoietic stem cells by K-Ras activating mutation. Haematologica. 2019;104:2215–24.
Li Q, Bohin N, Wen T, Ng V, Magee J, Chen SC, et al. Oncogenic Nras has bimodal effects on stem cells that sustainably increase competitiveness. Nature. 2013;504:143–7.
Zhao HG, Deininger MW. Always stressed but never exhausted: how stem cells in myeloid neoplasms avoid extinction in inflammatory conditions. Blood. 2023;141:2797–812.
Duchmann M, Laplane L, Itzykson R. Clonal Architecture and Evolutionary Dynamics in Acute Myeloid Leukemias. Cancers. 2021;13:4887.
Nazha A, Komrokji R, Meggendorfer M, Jia X, Radakovich N, Shreve J, et al. Personalized Prediction Model to Risk Stratify Patients With Myelodysplastic Syndromes. JCO. 2021;39:3737–46.
Badar T, Patel KP, Thompson PA, DiNardo C, Takahashi K, Cabrero M, et al. Detectable FLT3-ITD or RAS mutation at the time of transformation from MDS to AML predicts for very poor outcomes. Leuk Res. 2015;39:1367–74.
Makishima H, Yoshizato T, Yoshida K, Sekeres MA, Radivoyevitch T, Suzuki H, et al. Dynamics of clonal evolution in myelodysplastic syndromes. Nat Genet. 2017;49:204–12.
Shiozawa Y, Malcovati L, Gallì A, Pellagatti A, Karimi M, Sato-Otsubo A, et al. Gene expression and risk of leukemic transformation in myelodysplasia. Blood. 2017;130:2642–53.
Park HS, Son BR, Shin KS, Byeon S, Kim HK, Yang Y, et al. The Prognostic Ability of RAS Pathway-Related Gene Mutations in Patients with Myeloid Neoplasms Treated with Hypomethylating Agents. Acta Haematol. 2021;144:649–59.
Feld J, Tremblay D, Dougherty M, Czaplinska T, Sanchez G, Brady C, et al. Safety and Efficacy: Clinical Experience of Venetoclax in Combination With Hypomethylating Agents in Both Newly Diagnosed and Relapsed/Refractory Advanced Myeloid Malignancies. HemaSphere. 2021;5:e549.
Patnaik MM. How I diagnose and treat chronic myelomonocytic leukemia. haematol. 2022;107:1503–17.
Elena C, Gallì A, Such E, Meggendorfer M, Germing U, Rizzo E, et al. Integrating clinical features and genetic lesions in the risk assessment of patients with chronic myelomonocytic leukemia. Blood. 2016;128:1408–17.
Coston T, Pophali P, Vallapureddy R, Lasho TL, Finke CM, Ketterling RP, et al. Suboptimal response rates to hypomethylating agent therapy in chronic myelomonocytic leukemia; a single institutional study of 121 patients. Am J Hematol. 2019;94:767–79.
Gagelmann N, Badbaran A, Beelen DW, Salit RB, Stölzel F, Rautenberg C, et al. A prognostic score including mutation profile and clinical features for patients with CMML undergoing stem cell transplantation. Blood Adv. 2021;5:1760–9.
Loscocco GG, Rotunno G, Mannelli F, Coltro G, Gesullo F, Pancani F, et al. The prognostic contribution of CBL, NRAS, KRAS, RUNX1, and TP53 mutations to mutation-enhanced international prognostic score systems (MIPSS70/plus/plus v2.0) for primary myelofibrosis. Am J Hematol. 2024;99:68–78.
Winter PS, Sarosiek KA, Lin KH, Meggendorfer M, Schnittger S, Letai A, et al. RAS signaling promotes resistance to JAK inhibitors by suppressing BAD-mediated apoptosis. Sci Signal. 2014;7. Available from: https://www.science.org/doi/10.1126/scisignal.2005301.
Stivala S, Codilupi T, Brkic S, Baerenwaldt A, Ghosh N, Hao-Shen H, et al. Targeting compensatory MEK/ERK activation increases JAK inhibitor efficacy in myeloproliferative neoplasms. J Clin Investig. 2019;129:1596–611.
Döhner H, Estey E, Grimwade D, Amadori S, Appelbaum FR, Büchner T, et al. Diagnosis and management of AML in adults: 2017 ELN recommendations from an international expert panel. Blood. 2017;129:424–47.
Döhner H, Wei AH, Appelbaum FR, Craddock C, DiNardo CD, Dombret H, et al. Diagnosis and management of AML in adults: 2022 recommendations from an international expert panel on behalf of the ELN. Blood. 2022;140:1345–77.
DiNardo CD, Tiong IS, Quaglieri A, MacRaild S, Loghavi S, Brown FC, et al. Molecular patterns of response and treatment failure after frontline venetoclax combinations in older patients with AML. Blood. 2020;135:791–803.
Maiti A, Rausch CR, Cortes JE, Pemmaraju N, Daver NG, Ravandi F, et al. Outcomes of relapsed or refractory acute myeloid leukemia after frontline hypomethylating agent and venetoclax regimens. haematol. 2020;106:894–8.
Zhang Q, Riley-Gillis B, Han L, Jia Y, Lodi A, Zhang H, et al. Activation of RAS/MAPK pathway confers MCL-1 mediated acquired resistance to BCL-2 inhibitor venetoclax in acute myeloid leukemia. Sig Transduct Target Ther. 2022;7:51.
Dhakal P, Bates M, Tomasson MH, Sutamtewagul G, Dupuy A, Bhatt VR. Acute myeloid leukemia resistant to venetoclax-based therapy: What does the future hold? Blood Rev. 2023;59:101036.
Mustafa Ali MK, Williams MT, Corley EM, AlKaabba F, Niyongere S. Impact of KRAS and NRAS mutations on outcomes in acute myeloid leukemia. Leuk Lymphoma. 2023;64:962–71.
Bataller A, Bazinet A, DiNardo CD, Maiti A, Borthakur G, Daver NG, et al. Prognostic risk signature in patients with acute myeloid leukemia treated with hypomethylating agents and venetoclax. Blood Adv. 2024;8:927–35.
Farrar JE, Schuback HL, Ries RE, Wai D, Hampton OA, Trevino LR, et al. Genomic Profiling of Pediatric Acute Myeloid Leukemia Reveals a Changing Mutational Landscape from Disease Diagnosis to Relapse. Cancer Res. 2016;76:2197–205.
Ney GM, Anderson B, Bender J, Kumar-Sinha C, Wu YM, Vats P, et al. Mutations predictive of hyperactive Ras signaling correlate with inferior survival across high-risk pediatric acute leukemia. Transl Pediatr. 2020;9:43–50.
Bachas C, Schuurhuis GJ, Hollink IHIM, Kwidama ZJ, Goemans BF, Zwaan CM, et al. High-frequency type I/II mutational shifts between diagnosis and relapse are associated with outcome in pediatric AML: implications for personalized medicine. Blood. 2010;116:2752–8.
Bachas C, Schuurhuis GJ, Reinhardt D, Creutzig U, Kwidama ZJ, Zwaan CM, et al. Clinical relevance of molecular aberrations in paediatric acute myeloid leukaemia at first relapse. Br J Haematol. 2014;166:902–10.
Perrone S, Ottone T, Zhdanovskaya N, Molica M. How acute myeloid leukemia (AML) escapes from FMS-related tyrosine kinase 3 (FLT3) inhibitors? Still an overrated complication? Cancer Drug Resist. 2023;6:223–38.
Han L, Zhang Q, Dail M, Shi C, Cavazos A, Ruvolo VR, et al. Concomitant targeting of BCL2 with venetoclax and MAPK signaling with cobimetinib in acute myeloid leukemia models. Haematologica. 2020;105:697–707.
Choe S, Wang H, DiNardo CD, Stein EM, de Botton S, Roboz GJ, et al. Molecular mechanisms mediating relapse following ivosidenib monotherapy in IDH1-mutant relapsed or refractory AML. Blood Adv. 2020;4:1894–905.
Wang F, Morita K, DiNardo CD, Furudate K, Tanaka T, Yan Y, et al. Leukemia stemness and co-occurring mutations drive resistance to IDH inhibitors in acute myeloid leukemia. Nat Commun. 2021;12:2607.
DiNardo CD, Stein EM, de Botton S, Roboz GJ, Altman JK, Mims AS, et al. Durable Remissions with Ivosidenib in IDH1-Mutated Relapsed or Refractory AML. N Engl J Med. 2018;378:2386–98.
Amatangelo MD, Quek L, Shih A, Stein EM, Roshal M, David MD, et al. Enasidenib induces acute myeloid leukemia cell differentiation to promote clinical response. Blood. 2017;130:732–41.
Intlekofer AM, Shih AH, Wang B, Nazir A, Rustenburg AS, Albanese SK, et al. Acquired resistance to IDH inhibition through trans or cis dimer-interface mutations. Nature. 2018;559:125–9.
Burgess MR, Hwang E, Firestone AJ, Huang T, Xu J, Zuber J, et al. Preclinical efficacy of MEK inhibition in Nras-mutant AML. Blood. 2014;124:3947–55.
Decroocq J, Birsen R, Montersino C, Chaskar P, Mano J, Poulain L, et al. RAS activation induces synthetic lethality of MEK inhibition with mitochondrial oxidative metabolism in acute myeloid leukemia. Leukemia. 2022;36:1237–52.
Zhou X, Ji Y, Zhou J. Multiple Strategies to Develop Small Molecular KRAS Directly Bound Inhibitors. Molecules. 2023;28:3615.
Harwood SJ, Smith CR, Lawson JD, Ketcham JM. Selected Approaches to Disrupting Protein–Protein Interactions within the MAPK/RAS Pathway. IJMS. 2023;24:7373.
Escher TE, Satchell KJF. RAS degraders: The new frontier for RAS-driven cancers. Mol Ther. 2023;31:1904–19.
Lu X, ** J, Wu Y, Liu X, Liang X, Lin J, et al. Progress in RAS-targeted therapeutic strategies: From small molecule inhibitors to proteolysis targeting chimeras. Med Res Rev. 2024;44:812–32.
Steffen CL, Kaya P, Schaffner-Reckinger E, Abankwa D. Eliminating oncogenic RAS: back to the future at the drawing board. Biochemical Soc Trans. 2023;51:447–56.
Yang H, Zhou X, Fu D, Le C, Wang J, Zhou Q, et al. Targeting RAS mutants in malignancies: successes, failures, and reasons for hope. Cancer Commun. 2023;43:42–74.
Hurtado-Navarro L, Cuenca-Zamora EJ, Zamora L, Bellosillo B, Such E, Soler-Espejo E, et al. NLRP3 inflammasome activation and symptom burden in KRAS-mutated CMML patients is reverted by IL-1 blocking therapy. Cell Rep Med. 2023;4:101329.
Funding
DA was supported by a scholarship from the Ministère de l’Enseignement supérieur et de la Recherche. LCF was supported by a scholarship from the Agence Régionale de Santé (ARS).
Author information
Authors and Affiliations
Contributions
DA, LCF, and AR conceptualized the review and wrote the manuscript. CW critically reviewed and edited the manuscript. All authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Alawieh, D., Cysique-Foinlan, L., Willekens, C. et al. RAS mutations in myeloid malignancies: revisiting old questions with novel insights and therapeutic perspectives. Blood Cancer J. 14, 72 (2024). https://doi.org/10.1038/s41408-024-01054-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41408-024-01054-2
- Springer Nature Limited