Abstract
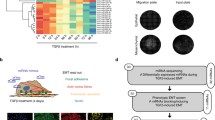
Since their discovery, microRNAs (miRNAs) have been widely studied in almost every aspect of biology and medicine, leading to the identification of important gene regulation circuits and cellular mechanisms. However, investigations are generally focused on the analysis of their downstream targets and biological functions in overexpression and knockdown approaches, while miRNAs endogenous levels and activity remain poorly understood. Here, we used the cellular plasticity-regulating process of epithelial-to-mesenchymal transition (EMT) as a model to show the efficacy of a fluorescent sensor to separate cells with distinct EMT signatures, based on miR-200b/c activity. The system was further combined with a CRISPR-Cas9 screening platform to unbiasedly identify miR-200b/c upstream regulating genes. The sensor allows to infer miRNAs fundamental biological properties, as profiling of sorted cells indicated miR-200b/c as a molecular switch between EMT differentiation and proliferation, and suggested a role for metabolic enzymes in miR-200/EMT regulation. Analysis of miRNAs endogenous levels and activity for in vitro and in vivo applications could lead to a better understanding of their biological role in physiology and disease.
Similar content being viewed by others
Introduction
MicroRNA (miRNA)s are short, highly conserved, RNAs that repress the expression of their target genes [1]. They directly/indirectly regulate many cellular processes like cell cycle, apoptosis, senescence, aging, and migration [2, 3], and have a role in both physiology and pathology. In cancer, miRNAs can act as tumor suppressors or oncomiRs [4,5,6], and have been linked with all hallmarks [6,7,8,9,10,11]. Epithelial-to-mesenchymal transition (EMT) is an embryonic development program, which many cancers hijack to increase the migratory and invasive capacities [12]. EMT is enforced by specialized transcription factor families like SNAIL, basic helix loop helix (bHLH), TWIST, and ZEB [13, 14], which can work as E-Cadherin repressors and promote the expression of EMT effector genes. MiRNAs can target EMT-inducing transcription factors [13, 15]. For example, miR-200 inhibits the transcription factors of the ZEB family in a reciprocal feedback loop [16] and miR-34 can target SNAIL [17]. The equilibrium between EMT-suppressing miRNAs and key EMT transcription factors determines the epithelial cellular plasticity, see Supplementary Fig. 1A.
MiRNAs are usually functionally investigated by overexpression or knocking-down techniques, and the downstream effects are measured by biochemical identification of target genes and by the analysis of cellular phenotypes. However, high-throughput functional assessment indicated that the majority of miRNAs expressed in cells do not show detectable targeting activity [18], suggesting the need for robust miRNA functional reporters to better investigate their physiological role. MiRNA sensors have been recently developed [18,19,20,21], and in limited cases used to isolate cells with distinct biological properties [22]. However, the systems so far proposed were based on artificial 3′UTRs containing multiple repeated perfect complementarity sequences or miRNA binding sites, instead of naturally occurring 3′UTRs, and generally lacked essential non-binding mutant controls [23].
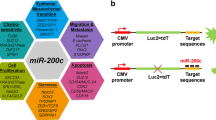
To develop a robust and controlled plasmid-based fluorescent miRNA sensor, we cloned a 3′UTR fragment of the miR-200 target ZEB2 [14], containing three strong miR-200 seed matches fused to DsRed (R) fluorescent protein, driven by a CMV promoter. As a non-targeting control, in the same plasmid we fused GFP (G) to the ZEB2 3′UTR carrying mutated seed matches, driven by an identical CMV promoter (the RwtGmut vector). Once expressed in living cells, this plasmid is designed to respond to miR-200 levels altering the intensity of the red fluorescence, while green fluorescence serves as a transfection control. A double mutant non-binding control vector (RmutGmut) was also generated, Fig. 1A. We previously showed that the RwtGmut sensor is highly specific for the miR-200 b/c/429 cluster [24]. In this study, we report that EMT biology can be explored in living cells by its transient transfection, and this approach could be exported to virtually all biologically relevant miRNAs in different cellular models, with several important applications for miRNA research.
A Schematic representation of the miR-200b/c sensor. GFP Green Fluorescent Protein, DsRed Discosoma Red Fluorescent Protein, RmutGmut and RwtGmut. wt wild-type, mut mutated, ZEB2 3′UTR 3′ untranslated region of ZEB2 gene. B Schematic representation of the separation of miR-200b/c sensor-transfected cells in a FACS plot. The X axis represents green fluorescence intensity, and the Y axis represents red fluorescence intensity. Red dots indicate miR-200b/c low cells, yellow dots are miR-200b/c high cells, and in green are cells with very high miR-200b/c levels (for instance of exogenous source). C FACS plots showing the fluorescence intensity of HCT116 cells with FITC-A (Green) and PE-A (Red) channels after transfection with sensor plasmid in the presence of either pre-control or pre-miR-200c at 100 nM concentration. Indicated are the percent of gated cells over the total amount of cells in the experiment, including un-transfected. D FACS plots showing transfection of PANC-1 cells with RmutGmut or RwtGmut plasmids and bar graphs showing the percent of inhibition. E FACS plots showing transfection of SKOV3 cells with RmutGmut or RwtGmut plasmids and bar graphs showing the percent of inhibition. F Western blot quantification of ZEB1 and E-Cadherin protein expression of HCT116 cells with miR-200c knockout (MIR200C-KO), compared to parental cells. β-Actin was used as a loading control. G FACS plots showing the transfection of sensor plasmids in MIR200C-KO HCT116 cells or in parental control cells. H Bar graphs showing the percent of inhibition of FACS analysis done in (G). In D–E p values are from Student’s t test. In H p values are from two-way ANOVA. Points are average ± SD. *<0.05, **<0.01, ***<0.001, ****<0.0001.
Results
The sensor plasmid can detect endogenous miR-200b/c activity
The dual fluorescence sensor was overexpressed in cells by transient transfection and its ability to report endogenous miR-200b/c levels was monitored by FACS. The scheme in Fig. 1B illustrates the different cell populations based on their miR-200b/c levels, as visualized in a FACS following sensor transfection. As a cellular model, we chose the colorectal cancer (CRC) cell line HCT116, which expresses high levels of miR-200s [14] and has an epithelial-like phenotype [25]. HCT116 cells were co-transfected with RmutGmut control plasmid in the presence of miR-200c mimics (pre-miR-200c) or scrambled control (pre-control), and the green and red fluorescence intensity were recorded on a flow cytometer. As a result, we found that the double mutant plasmid allowed a robust and coordinated expression of the fluorescent transgenes (cells along a diagonal line in the FACS plot), an important pre-requisite for the detection of changes of signal intensities due to the binding of endogenous miRNAs. Overexpression of pre-miR-200c produced no detectable alteration of fluorescence in these cells, Supplementary Fig. 1B, indicating the lack of binding activity. By contrast, using the signal from the non-binding RmutGmut plasmid to set the FACS gates, we could observe the appearance of a population with reduced red intensity in cells transfected with the RwtGmut sensor (with scrambled control), Fig. 1C. This result suggests that, when expressed in cells with detectable levels, the miR-200b/c sensor can report a significant binding activity in a sub-population of cells, potentially carrying the highest endogenous miR-200b/c levels. The percentage of cells in the gated area for high miR-200b/c further increased upon miR-200c overexpression, Fig. 1C, suggesting the notion that the sensor can be used to monitor cells with high miR-200b/c from endogenous and exogenous source. Similar results were obtained from another miR-200b/c positive cell line, the human bladder carcinoma RT112, Supplementary Fig. 1C, and endogenous levels were visualized also in other cancer cell lines, like PANC-1 and SKOV3, Fig. 1D, E.
The sensor reports miR-200 activity in cells with differential EMT states
The miR-200 family has a strong EMT-repressing role and is normally reduced in mesenchymal-like cells [15]. We sought to determine the activity of the sensor in cells with altered EMT status using an isogenic cellular system with high and low miR-200. For this, we tested mouse pancreatic cancer cell lines derived from a Pdx1-cre;KrasLSL.G12D/+;Tp53LSL.R172H/+ mouse model (KPC), which has been further modified to obtain the Zeb1 knockout, called KPCZ, to show a role of Zeb1 in the metastatic cascade of pancreatic tumors [26]. The mature sequence of miR-200b/c is highly conserved between human and mice [27]. Cells obtained from KPCZ tumors showed a more marked epithelial-like phenotype, Supplementary Fig. 2A, B, compared to KPC cells, with increased expression of miR-200 members b and c, Supplementary Fig. 2C. Upon transfection with miRNA sensors, KPCZ cells showed a superior inhibition rate, see Supplementary Fig. 2D, E, in line with the data obtained with the pre-miR transfection experiment.
To further test the specificity of the detection, we used a CRISPR/Cas9-based approach to permanently knockout miR-200c, the most prominent EMT-suppressing member of the miRNA family [28], taking advantage of a protospacer adjacent motif (PAM) sequence adjacent to the miRNA-200c seed match, Supplementary Fig. 3A. As a result, MIR200C-KO HCT116 cells showed a drastic reduction of miR-200c levels, as evaluated by qPCR, Supplementary Fig. 3B. In light of the high-sequence homology between the miR-200b and c, we also verified the effect of the KO on the MIR200B gene and a significant >2-fold reduction was also detected, Supplementary Fig. 3C. DNA sequencing on single-cell clones obtained from MIR200C-KO cells showed alterations around the PAM in the miR-200c, but not in the miR-200b region, Supplementary Fig. 3D, suggesting that the observed reduction in miR-200b is likely an indirect effect due to EMT induction, possibly mediated by ZEB1 increase. Phenotypic analysis of the MIR200C-KO cells, in fact, showed that they had undergone EMT, as indicated by a pronounced mesenchymal-like morphology and dispersed growth pattern, and confirmed by western blotting and immunofluorescence staining (Fig. 1F and Supplementary Fig. 4A). To check that EMT was propelled by the miR-200c knockout and not by an unwanted off-target effect [29], we reconstituted the cells with exogenous miR-200c by transient transfection and could observe an attenuation of the phenotype, Supplementary Fig. 4B. Once transfected with miR-200b/c sensors, the miR-200C-KO cells displayed a significant reduction in the percentage of cells with inhibition of red fluorescence, Fig. 1G, H, further supporting the high specificity of the miR-200b/c sensor system. Overall, these data indicated that the sensor can monitor changes in the miR-200b/c levels in living cells with distinct EMT states.
Utility of the sensor to sort cells by endogenous miR-200b/c levels/EMT status
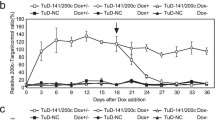
We then tested the possibility to use the sensor for FACS-sorting cells with a differential EMT status based on the endogenous high- and low-miR-200b/c expression. HCT116 cells were transfected with RwtGmut and RmutGmut plasmids on a larger scale and separated by their RwtGmut non-inhibited (miR-200b/c low) and red-inhibited (miR-200b/c high) population, Fig. 2A. Gated areas were selected to collect cells with the same intensity of green fluorescence (the mutant control) and a significant difference in red. After sorting, cells were re-plated and allowed to attach and grow for 3 days to recover, to exclude dead cells from the further characterizations and to increase in number. A western blot conducted to monitor the expression of the epithelial marker E-Cadherin identified no change in E-Cadherin abundance in RwtGmut compared to RmutGmut transfected cells (unsorted), to control that the miR-200b/c sensor itself did not alter the EMT phenotype. Additional experiments on HCT116, RT112, and PANC-1 parental unsorted cells were further performed to better rule out the possibility that the sensor was sequestering miR-200b/c and favouring EMT upon transfection, Supplementary Fig. 5A, B. Analysis of sorted cells showed that E-Cadherin levels were also unaltered in RmutGmut compared to parental cells, while a significant reduction was found in miR-200b/c low, and a relatively marked increase was detected in miR-200b/c high cells obtained from RwtGmut transfection, Fig. 2B. In a separate experiment, qPCR analysis confirmed the differential gene expression of miR-200c and EMT markers in miR-200b/c high- and low-sorted cells (Fig. 2C). Similar results were obtained by sorting sensor-transfected RT112 cells, Supplementary Fig. 5C, D. In addition, we found that sorted high and low cells had a strong propensity for quickly reverting their fluorescence after re-plating, as quantified by FACS analysis and video imaging (Supplementary Fig. 5E, F), indicating that miRNA activity can be monitored for a few days in vitro in individual sensor-transfected cells. Interestingly, despite the fact that a differential EMT status could not be evidenced by the microscopic morphological examination of sorted cells, miR-200b/c low clearly showed a significantly lower ability to attach after sorting, compared to high cells. Quantifications confirmed that they were more rounded and of smaller size when re-plated, indicating a lower adhesion capacity, before normalizing in the following days (Supplementary Fig. 5G, H). Altogether, this was the first indication that the sensor-guided FACS-sorting strategy was capable of physically separating cells with distinct miR-200b/c levels and EMT properties.
A FACS plots showing the gates used for sensor-sorting miR-200b/c low and high cells after transfection with the RwtGmut plasmid (right panel). Left panel shows RmutGmut transfected control cells and their gate for sorting control. B Western blot quantification of E-Cadherin in HCT116 cells transfected either with RmutGmut or RwtGmut plasmids (unsorted, left panel), or sorted with the gates shown in (A) and re-plated for 3 days to grow and remove dead cells (right panel). C qPCR quantification of relative mRNA levels of CDH1 (gene coding E-Cadherin), VIM (Vimentin), ZEB1 and miR-200c in miR-200b/c low or high HCT116 cells sorted as in (A). Gene set enrichment analysis of RNA-sequencing data depicting pathways D upregulated and E downregulated in sensor-sorted miR-200b/c low cells compared to miR-200b/c high cells. F qPCR validation of RNA-sequencing data. CDC25B, CDCA3, CCNF, SPC25, TNFAIP3, SERPINE1, LTBP2, ZEB1 and ZEB2 were quantified using GAPDH as a housekee** gene. p values are from Student’s t test. Points are average ± SD. *<0.05, **<0.01, ***<0.001, ****<0.0001.
miR-200b/c sensor-sorted cells can be used to molecularly characterize endogenous EMT states
EMT is a complex multifactorial process. Although a few markers like E-Cadherin, Vimentin and ZEB1 can represent a good surrogate for the determination of the EMT state [14, 30], this should be better concluded from broader, and possibly genome-wide characterizations [31]. We, therefore, repeated the sorting experiment twice independently (with duplicates) and subjected all the eight RNAs isolated from high and low cells to sequencing. Using a cut-off of twofold for defining differentially expressed genes, we identified 224 upregulated and 73 downregulated genes in the miR-200b/c low compared to high cells, as an overlap between the two distinct sorting experiments, see Supplementary Fig. 6A and Supplementary Table 1A, B. A geneset enrichment analysis revealed that the signatures most significantly upregulated in the miR-200b/c low cells were belonging to the TNF-alpha signaling via NF-κB, to EMT, and the TGF-beta signaling pathways, Fig. 2D. On the other hand, analysis of the genes downregulated in miR-200b/c low cells indicated the significant prevalence of cell-cycle-related targets, like those linked with the E2F transcription factors and genes involved in the G2/M progression through the cells division cycle, Fig. 2E. Of note, ZEB1 and ZEB2 were identified among the most differentially expressed genes, Supplementary Fig. 6B. This is particularly important as a further assay validation, since the sensor plasmid is designed as a functional readout of the miR-200b/c binding to the ZEB transcription factors. Moreover, clustering analysis of miR-sensor low and high cells with the parental HCT116 cells indicated that HCT116 cells resembled and clustered with miR-200b/c high cells, considering both differentially expressed genes (Supplementary Fig. 6C) and known miR-200b/c target genes (Supplementary Fig. 6D-F), in line with the fact that HCT116 have a predominant epithelial-like nature. Quantitative PCR was used to independently validate the RNA-sequencing results and confirmed the down- or upregulation of relevant genes in the identified pathways, like CDCA3 among the E2F target genes and TNFAIP3 for TNF-alpha signaling along with ZEB1 and ZEB2, Fig. 2F.
We next sought to functionally validate the role on EMT of the pathways associated with the gene signatures obtained from sensor-sorted cells. TGF-beta is an established master EMT inducer [32]. TNF-alpha has also been reported as an EMT-inducing cytokine in different cancer cell lines, including cells from colorectal origin, with and without the NF-κB-mediated upregulation of EMT transcription factors [24, 57], here confirmed by the analysis of endogenous levels in different cellular models and by overexpression and knockdown approaches. However, the sensitivity can be the object of further improvements, allowing to extend the assay to cells with lower miRNA endogenous expression. Sensors designed with artificial 3′UTRs with multiple fragments perfect complementary to the targeting miRNA can guarantee a higher level of sensitivity, as previously shown [58]. However, the sensor here presented has the advantage of carrying naturally occurring sequences properly controlled with non-binding mutants, in analogy with the dual reporter vectors used to validate bona fide miRNA targets. This method is preferable to better represent physiological conditions [58] by (1) making sure that the sensors are reliably reporting endogenous miRNA activity, i.e., not overestimating their effects, and by (2) minimizing or eliminating miRNA-sponge or decoy effects [18, 59], which could alter the biological properties. Another unappreciated factor is that the presence of non-natural 3′UTRs (either containing repeated seed matches or complementary sequences) carrying random spacer sequences increases the chances to introduce novel unwanted, albeit specific, miRNA binding sites. In the absence of non-binding controls, the contribution of these off-target detections can be difficult to estimate. To improve sensitivity, therefore, multiple 3′UTRs controlled with non-binding mutants in a 1:1 ratio could in future be cloned in tandem and tested. Another limiting factor is that this sensor can only be transiently introduced in the cells, thus limiting the number of cell lines to be potentially investigated and the amount of output cells per sorting round, an important factor for the downstream techniques requiring higher amount of cells. MiRNA-reporter cells with integrated sensors would allow better live tracking of miRNAs activity in experimental settings in vitro and in vivo. However, integration of this vector into the genome produced recombination events possibly due to the highly repetitive sequences [24], a fact that precluded its further use. To overcome this limitation, future studies should be conducted to improve the sensor architecture, like using single bidirectional promoter vectors [60] to minimize the presence of repetitive sequences. Once improved, this approach could be implemented for all biologically relevant miRNAs, leading to fundamental discoveries in biomedicine.
Methods
Cell culture and chemicals
HCT116 cells were cultured in McCoy’s 5A (Lonza), RT112 and COLO205 cells were cultured in RPMI1640 (Sigma), PANC-1 and HEK 293 (all from ATCC) were cultured in DMEM (Sigma), and SKOV3 cells (NCI) were cultured in RPMI1640 supplemented with 1 mM sodium pyruvate (Gibco), 1% MEM NEAA (Gibco), and 1% MEM Vitamin solution (Gibco). Media were supplemented with 10% FBS (Sigma), 1% pencillin/streptomycin (Sigma) and 1% l-glutamine (Sigma). Cells were cultured at 37 °C and 5% CO2 in a humidified incubator. Cells were STR authenticated and tested for mycoplasma regularly (Invivogen). HCT116-Cas9 cell line was generated by lentiviral transduction with lentiCas9-Blast (Addgene) and selection with 4 μg/mL of blasticidin (Sigma) for 3 days. KPC (Pdx1-cre;KrasLSL.G12D/+;Tp53LSL.R172H/+) and KPCZ (Pdx1-cre;KrasLSL.G12D/+;Tp53LSL.R172H/+; ZEBfl/fl) are mouse pancreatic cancer cells without and with ZEB1 knockout [26] and were cultured in DMEM. TNF-alpha was from Gibco, and Cdk inhibitor (CGP-60474) from Tocris.
FACS analysis
HCT116 cells were seeded in six-well plates at a density of 0.5 million per well. The next day cells were transfected with 1.5 µg of either RmutGmut or RwtGmut plasmids. After 48 h, cells were trypsinized, washed, and resuspended in FACS buffer (5 mM EDTA and 2% FBS/PBS). Samples were run on Cytoflex FACS machine (Beckman). FACS data were analyzed using FlowJo software v10.6.
CRISPR screen
HCT116-Cas9 cells were transduced with lentiviral Human GeCKO v2 library part A and part B at MOI of 0.3 in the presence of 4 μg/mL of polybrene for 24 h, then replaced the virus medium with fresh growth medium and continued to culture the cells for 48 h. The cells were selected with 4 μg/mL of puromycin for 3 days. Then combined both half libraries cells and collected 2.5 × 107 cells for genomic DNA isolation. Next-generation sequencing was performed on the Illumina HiSeq 2500 platform in Deep Sequencing Facility of TU Dresden. The raw FASTQ files were analyzed with MAGeCK-VISPR.
Additional methods are available in the Supplements.
Data availability
RNA-sequencing data have been deposited to GEO dataset GSE154429.
References
Hannon GJ. RNA interference. Nature 2002;418:244–51.
Hwang HW, Mendell JT. MicroRNAs in cell proliferation, cell death, and tumorigenesis. Br J Cancer. 2006;94:776–80.
Olivieri F, Rippo MR, Monsurrò V, Salvioli S, Capri M, Procopio AD, et al. MicroRNAs linking inflamm-aging, cellular senescence and cancer. Ageing Res Rev. 2013;12:1056–68.
Esquela-Kerscher A, Slack FJ. Oncomirs—microRNAs with a role in cancer. Nat Rev Cancer. 2006;6:259–69.
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E, et al. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci USA. 2002;99:15524–9.
Peng Y, Croce CM. The role of MicroRNAs in human cancer. Signal Transduct Target Ther. 2016;1:15004.
Lima RT, Busacca S, Almeida GM, Gaudino G, Fennell DA, Vasconcelos MH. MicroRNA regulation of core apoptosis pathways in cancer. Eur J Cancer. 2011;47:163–74.
O’Donnell KA, Wentzel EA, Zeller KI, Dang CV, Mendell JT. c-Myc-regulated microRNAs modulate E2F1 expression. Nature. 2005;435:839–43.
Fasanaro P, D’Alessandra Y, Di Stefano V, Melchionna R, Romani S, Pompilio G, et al. MicroRNA-210 modulates endothelial cell response to hypoxia and inhibits the receptor tyrosine kinase ligand Ephrin-A3. J Biol Chem. 2008;283:15878–83.
Kong W, Yang H, He L, Zhao JJ, Coppola D, Dalton WS, et al. MicroRNA-155 is regulated by the transforming growth factor beta/Smad pathway and contributes to epithelial cell plasticity by targeting RhoA. Mol Cell Biol. 2008;28:6773–84.
Gregory PA, Bert AG, Paterson EL, Barry SC, Tsykin A, Farshid G, et al. The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol. 2008;10:593–601.
Thiery JP, Acloque H, Huang RY, Nieto MA. Epithelial-mesenchymal transitions in development and disease. Cell. 2009;139:871–90.
Díaz-López A, Moreno-Bueno G, Cano A. Role of microRNA in epithelial to mesenchymal transition and metastasis and clinical perspectives. Cancer Manag Res. 2014;6:205–16.
Park SM, Gaur AB, Lengyel E, Peter ME. The miR-200 family determines the epithelial phenotype of cancer cells by targeting the E-cadherin repressors ZEB1 and ZEB2. Genes Dev. 2008;22:894–907.
Ceppi P, Peter ME. MicroRNAs regulate both epithelial-to-mesenchymal transition and cancer stem cells. Oncogene. 2014;33:269–78.
Brabletz S, Brabletz T. The ZEB/miR-200 feedback loop-a motor of cellular plasticity in development and cancer? EMBO Rep. 2010;11:670–7.
Siemens H, Jackstadt R, Hünten S, Kaller M, Menssen A, Götz U, et al. miR-34 and SNAIL form a double-negative feedback loop to regulate epithelial-mesenchymal transitions. Cell Cycle. 2011;10:4256–71.
Mullokandov G, Baccarini A, Ruzo A, Jayaprakash AD, Tung N, Israelow B, et al. High-throughput assessment of microRNA activity and function using microRNA sensor and decoy libraries. Nat Methods 2012;9:840–6.
Tian W, Dong X, Liu X, Wang G, Dong Z, Shen W, et al. High-throughput functional microRNAs profiling by recombinant AAV-based microRNA sensor arrays. PLoS ONE. 2012;7:e29551.
Brustikova K, Sedlak D, Kubikova J, Skuta C, Solcova K, Malik R, et al. Cell-based reporter system for high-throughput screening of MicroRNA pathway inhibitors and its limitations. Front Genet. 2018;9:45.
Bosson AD, Zamudio JR, Sharp PA. Endogenous miRNA and target concentrations determine susceptibility to potential ceRNA competition. Mol Cell. 2014;56:347–59.
Belur Nagaraj A, Joseph P, Ponting E, Fedorov Y, Singh S, Cole A, et al. A miRNA-mediated approach to dissect the complexity of tumor-initiating cell function and identify miRNA-targeting drugs. Stem Cell Rep. 2019;12:122–34.
Thomson DW, Bracken CP, Goodall GJ. Experimental strategies for microRNA target identification. Nucleic Acids Res. 2011;39:6845–53.
Ceppi P, Hadji A, Kohlhapp FJ, Pattanayak A, Hau A, Liu X, et al. CD95 and CD95L promote and protect cancer stem cells. Nat Commun. 2014;5:5238.
Siddiqui A, Vazakidou ME, Schwab A, Napoli F, Fernandez-Molina C, Rapa I, et al. Thymidylate synthase is functionally associated with ZEB1 and contributes to the epithelial-to-mesenchymal transition of cancer cells. J Pathol. 2017;242:221–33.
Krebs AM, Mitschke J, Lasierra Losada M, Schmalhofer O, Boerries M, Busch H, et al. The EMT-activator Zeb1 is a key factor for cell plasticity and promotes metastasis in pancreatic cancer. Nat Cell Biol. 2017;19:518–29.
Trumbach D, Prakash N. The conserved miR-8/miR-200 microRNA family and their role in invertebrate and vertebrate neurogenesis. Cell Tissue Res. 2015;359:161–77.
Ceppi P, Mudduluru G, Kumarswamy R, Rapa I, Scagliotti GV, Papotti M, et al. Loss of miR-200c expression induces an aggressive, invasive, and chemoresistant phenotype in non-small cell lung cancer. Mol Cancer Res. 2010;8:1207–16.
Zhang XH, Tee LY, Wang XG, Huang QS, Yang SH. Off-target effects in CRISPR/Cas9-mediated genome engineering. Mol Ther Nucleic Acids. 2015;4:e264.
Schwab A, Siddiqui A, Vazakidou ME, Napoli F, Bottcher M, Menchicchi B, et al. Polyol pathway links glucose metabolism to the aggressiveness of cancer cells. Cancer Res. 2018;78:1604–18.
Yang J, Antin P, Berx G, Blanpain C, Brabletz T, Bronner M, et al. Guidelines and definitions for research on epithelial-mesenchymal transition. Nat Rev Mol Cell Biol. 2020;21:341–52.
Moustakas A, Heldin CH. Induction of epithelial-mesenchymal transition by transforming growth factor beta. Semin Cancer Biol. 2012;22:446–54.
Li CW, **a W, Huo L, Lim SO, Wu Y, Hsu JL, et al. Epithelial-mesenchymal transition induced by TNF-alpha requires NF-kappaB-mediated transcriptional upregulation of Twist1. Cancer Res. 2012;72:1290–300.
Shi G, Zheng X, Zhang S, Wu X, Yu F, Wang Y, et al. Kanglaite inhibits EMT caused by TNF-α via NF-κΒ inhibition in colorectal cancer cells. Oncotarget. 2018;9:6771–9.
Wang H, Wang HS, Zhou BH, Li CL, Zhang F, Wang XF, et al. Epithelial-mesenchymal transition (EMT) induced by TNF-α requires AKT/GSK-3β-mediated stabilization of snail in colorectal cancer. PLoS ONE. 2013;8:e56664.
Pan Y, Liang H, Chen W, Zhang H, Wang N, Wang F, et al. microRNA-200b and microRNA-200c promote colorectal cancer cell proliferation via targeting the reversion-inducing cysteine-rich protein with Kazal motifs. RNA Biol. 2015;12:276–89.
Carter JV, O’Brien SJ, Burton JF, Oxford BG, Stephen V, Hallion J, et al. The microRNA-200 family acts as an oncogene in colorectal cancer by inhibiting the tumor suppressor RASSF2. Oncol Lett. 2019;18:3994–4007.
Sanjana NE, Shalem O, Zhang F. Improved vectors and genome-wide libraries for CRISPR screening. Nat Methods. 2014;11:783–4.
Behan FM, Iorio F, Picco G, Goncalves E, Beaver CM, Migliardi G, et al. Prioritization of cancer therapeutic targets using CRISPR-Cas9 screens. Nature. 2019;568:511–6.
Pastushenko I, Brisebarre A, Sifrim A, Fioramonti M, Revenco T, Boumahdi S, et al. Identification of the tumour transition states occurring during EMT. Nature. 2018;556:463–8.
Kroger C, Afeyan A, Mraz J, Eaton EN, Reinhardt F, Khodor YL, et al. Acquisition of a hybrid E/M state is essential for tumorigenicity of basal breast cancer cells. Proc Natl Acad Sci USA. 2019;116:7353–62.
Bartel DP. Metazoan microRNAs. Cell. 2018;173:20–51.
Mandal M, Ghosh B, Anura A, Mitra P, Pathak T, Chatterjee J. Modeling continuum of epithelial mesenchymal transition plasticity. Integr Biol. 2016;8:167–76.
Davis FM, Stewart TA, Thompson EW, Monteith GR. Targeting EMT in cancer: opportunities for pharmacological intervention. Trends Pharmacol Sci. 2014;35:479–88.
Oft M, Heider KH, Beug H. TGFbeta signaling is necessary for carcinoma cell invasiveness and metastasis. Curr Biol. 1998;8:1243–52.
Huber MA, Azoitei N, Baumann B, Grunert S, Sommer A, Pehamberger H, et al. NF-kappaB is essential for epithelial-mesenchymal transition and metastasis in a model of breast cancer progression. J Clin Investig. 2004;114:569–81.
Bates RC, Mercurio AM. Tumor necrosis factor-alpha stimulates the epithelial-to-mesenchymal transition of human colonic organoids. Mol Biol Cell. 2003;14:1790–800.
Hugo HJ, Pereira L, Suryadinata R, Drabsch Y, Gonda TJ, Gunasinghe NP, et al. Direct repression of MYB by ZEB1 suppresses proliferation and epithelial gene expression during epithelial-to-mesenchymal transition of breast cancer cells. Breast Cancer Res. 2013;15:R113.
Spaderna S, Schmalhofer O, Hlubek F, Berx G, Eger A, Merkel S, et al. A transient, EMT-linked loss of basement membranes indicates metastasis and poor survival in colorectal cancer. Gastroenterology. 2006;131:830–40.
Rubio CA. Further studies on the arrest of cell proliferation in tumor cells at the invading front of colonic adenocarcinoma. J Gastroenterol Hepatol. 2007;22:1877–81.
Savagner P. The epithelial-mesenchymal transition (EMT) phenomenon. Ann Oncol. 2010;21:vii89–92.
Pastushenko I, Blanpain C. EMT transition states during tumor progression and metastasis. Trends Cell Biol. 2019;29:212–26.
Ramesh V, Brabletz T, Ceppi P. Targeting EMT in cancer with repurposed metabolic inhibitors. Trends Cancer. 2020;6:942–50.
Dong C, Yuan T, Wu Y, Wang Y, Fan TW, Miriyala S, et al. Loss of FBP1 by Snail-mediated repression provides metabolic advantages in basal-like breast cancer. Cancer Cell. 2013;23:316–31.
Nguyen-Vu T, Wang J, Mesmar F, Mukhopadhyay S, Saxena A, McCollum CW, et al. Estrogen receptor beta reduces colon cancer metastasis through a novel miR-205 - PROX1 mechanism. Oncotarget. 2016;7:42159–71.
Sun Y, Wang BE, Leong KG, Yue P, Li L, Jhunjhunwala S, et al. Androgen deprivation causes epithelial-mesenchymal transition in the prostate: implications for androgen-deprivation therapy. Cancer Res. 2012;72:527–36.
Feng X, Wang Z, Fillmore R **Y. MiR-200, a new star miRNA in human cancer. Cancer Lett. 2014;344:166–73.
Mukherji S, Ebert MS, Zheng GX, Tsang JS, Sharp PA, van Oudenaarden A. MicroRNAs can generate thresholds in target gene expression. Nat Genet. 2011;43:854–9.
Ebert MS, Sharp PA. Emerging roles for natural microRNA sponges. Curr Biol. 2010;20:R858–61.
Amendola M, Venneri MA, Biffi A, Vigna E, Naldini L. Coordinate dual-gene transgenesis by lentiviral vectors carrying synthetic bidirectional promoters. Nat Biotechnol. 2005;23:108–16.
Acknowledgements
We acknowledge the work of Prof. Marcus Peter and Dr. Sun Mi Park for the early development of the sensor plasmids. We would like to thank the FACS-core facility of the Nikolaus-Fiebiger-Zentrum of the Friedrich-Alexander University Erlangen-Nuremberg.
Funding
This work was supported by the Interdisciplinary Center for Clinical Research of the University of Erlangen-Nuremberg (PC) and the German Research Foundation grant DFG-CE-281/5-1 (PC). PC was also supported by IASLC Young Investigator Award. CP and HY were supported by the EU (MSCA grant no.: 861196). Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
Conception and design: PNG, PC; Development of methodology: PNG, BP, ASi, PC; Acquisition of data (provided animals, acquired and managed patients, provided facilities, etc.): PNG., BP, ASi, HY, VR, FN, ASc, RN, DM, CP, TB; Analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis): PNG, HY, VR, DM, PC; Writing, review, and/or revision of the manuscript: PNG, IA, TB, CP, PC; Administrative, technical, or material support (i.e., reporting or organizing data, constructing databases): IA, CP, PC; Study supervision: PC. The author(s) read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gollavilli, P.N., Parma, B., Siddiqui, A. et al. The role of miR-200b/c in balancing EMT and proliferation revealed by an activity reporter. Oncogene 40, 2309–2322 (2021). https://doi.org/10.1038/s41388-021-01708-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-021-01708-6
- Springer Nature Limited
This article is cited by
-
A Narrative Review of Diabetic Macroangiopathy: From Molecular Mechanism to Therapeutic Approaches
Diabetes Therapy (2024)
-
MiR-200b categorizes patients into pancreas cystic lesion subgroups with different malignant potential
Scientific Reports (2023)
-
Clinical significance of MATN1-AS1 as ceRNA of Mir-200b in tissues and serum of patients with cervical cancer
Scientific Reports (2023)
-
Metabolic impairment of non-small cell lung cancers by mitochondrial HSPD1 targeting
Journal of Experimental & Clinical Cancer Research (2021)