Abstract
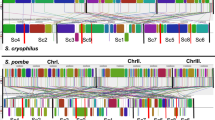
The genome of the lower eukaryote Dictyostelium discoideum comprises six chromosomes. Here we report the sequence of the largest, chromosome 2, which at 8 megabases (Mb) represents about 25% of the genome. Despite an A + T content of nearly 80%, the chromosome codes for 2,799 predicted protein coding genes and 73 transfer RNA genes. This gene density, about 1 gene per 2.6 kilobases (kb), is surpassed only by Saccharomyces cerevisiae (one per 2 kb) and is similar to that of Schizosaccharomyces pombe (one per 2.5 kb)1,2. If we assume that the other chromosomes have a similar gene density, we can expect around 11,000 genes in the D. discoideum genome. A significant number of the genes show higher similarities to genes of vertebrates than to those of other fully sequenced eukaryotes1,2,3,4,5,6. This analysis strengthens the view that the evolutionary position of D. discoideum is located before the branching of metazoa and fungi but after the divergence of the plant kingdom7, placing it close to the base of metazoan evolution.



Similar content being viewed by others
References
Goffeau, A. et al. Life with 6000 genes. Science 274, 546–567 (1996)
Wood, V. et al. The genome sequence of Schizosaccharomyces pombe. Nature 415, 871–880 (2002)
The C. elegans Sequencing Consortium Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282, 2012–2018 (1998)
Adams, M. D. et al. The genome sequence of Drosophila melanogaster. Science 287, 2185–2195 (2000)
The Arabidopsis Genome Initiative. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408, 796–815 (2000)
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001)
Baldauf, S. L., Roger, A. J., Wenk-Siefert, I. & Doolittle, W. F. A kingdom-level phylogeny of eukaryotes based on combined protein data. Science 290, 972–977 (2000)
Loomis, W. F. Genetic networks that regulate development in Dictyostelium cells. Microbiol. Rev. 60, 135–150 (1996)
Eichinger, L., Lee, S. S. & Schleicher, M. Dictyostelium as model system for studies of the actin cytoskeleton by molecular genetics. Microsc. Res. Technol. 47, 124–134 (1999)
Parent, C. A. & Devreotes, P. N. A cell's sense of direction. Science 284, 765–770 (1999)
Firtel, R. A. & Meili, R. Dictyostelium: a model for regulated cell movement during morphogenesis. Curr. Opin. Genet. Dev. 10, 421–427 (2000)
Noegel, A. A. & Schleicher, M. The actin cytoskeleton of Dictyostelium: a story told by mutants. J. Cell Sci. 113, 759–766 (2000)
Kay, R. R. & Williams, J. G. The Dictyostelium genome project: an invitation to species hop**. Trends Genet. 15, 294–297 (1999)
Cox, E. C., Vocke, C. D., Walter, S., Gregg, K. Y. & Bain, E. S. Electrophoretic karyotype for Dictyostelium discoideum. Proc. Natl Acad. Sci. USA 87, 8247–8251 (1990)
Loomis, W. F. & Kuspa, A. Dictyostelium—A Model System for Cell and Developmental Biology (eds Maeda, Y., Inouye, K. & Takeuchi, I.) 15–30 (Universal Academic, Tokyo, 1997)
Gardner, M. J. et al. Chromosome 2 sequence of the human malaria parasite Plasmodium falciparum. Science 282, 1126–1132 (1998)
Bowman, S. et al. The complete nucleotide sequence of chromosome 3 of Plasmodium falciparum. Nature 400, 532–538 (1999)
Glöckner, G. Large scale sequencing and analysis of AT rich eukaryote genomes. Curr. Genom. 1, 289–299 (2000)
Glöckner, G. et al. The complex repeats of Dictyostelium discoideum. Genome Res. 11, 585–594 (2001)
Dear, P. H. in Genome Map**—A Practical Approach (ed. Dear, P. H.) 95–124 (IRL Press, Oxford, 1997)
Pan, W. C. & Blackburn, E. H. Single extrachromosomal ribosomal RNA gene copies are synthesized during amplification of the rDNA in Tetrahymena. Cell 23, 459–466 (1981)
Lim, L. P. & Burge, C. B. A computational analysis of sequence features involved in recognition of short introns. Proc. Natl Acad. Sci. USA 98, 11193–11198 (2001)
Morio, T. et al. The Dictyostelium developmental cDNA project: generation and analysis of expressed sequence tags from the first-finger stage of development. DNA Res. 5, 335–340 (1998)
Apweiler, R. et al. InterPro—an integrated documentation resource for protein families, domains and functional sites. Bioinformatics 16, 1145–1150 (2000)
Goebl, M. G. & Petes, T. D. Most of the yeast genomic sequences are not essential for cell growth and division. Cell 46, 983–992 (1986)
Konfortov, B. A., Cohen, H. M., Bankier, A. T. & Dear, P. H. A high-resolution HAPPY map of Dictyostelium discoideum chromosome 6. Genome Res. 10, 1737–1742 (2000)
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997)
Parra, G., Blanco, E. & Guigo, R. GeneID in Drosophila. Genome Res. 10, 511–515 (2000)
The Gene Ontology Consortium. Gene ontology: tool for the unification of biology. Nature Genet. 25, 25–29 (2000)
Fortini, M. E., Skupski, M. P., Boguski, M. S. & Hariharan, I. K. A survey of human disease gene counterparts in the Drosophila genome. J. Cell Biol. 150, F23–F30 (2000)
Acknowledgements
We thank S. Förste, N. Zeisse, S. Rothe, S. Landmann, R. Schultz, S. Müller and R. Müller for expert technical assistance. We also thank the working team of the Japanese cDNA project (http://www.csm.biol.tsukuba.ac.jp/cDNAproject.html) for sharing data. The sequencing of chromosome 2 was supported by the Deutsche Forschungsgemeinschaft, with partial support by Köln Fortune. Additional support was obtained from the NIH, the Medical Research Council and the EU. Members of the Sanger Institute Dictyostelium discoideum sequencing team are listed at http://www.sanger.ac.uk/Projects/D_discoideum/team.shtml.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Glöckner, G., Eichinger, L., Szafranski, K. et al. Sequence and analysis of chromosome 2 of Dictyostelium discoideum. Nature 418, 79–85 (2002). https://doi.org/10.1038/nature00847
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature00847
- Springer Nature Limited
This article is cited by
-
Adenylyl cyclase mRNA localizes to the posterior of polarized DICTYOSTELIUM cells during chemotaxis
BMC Cell Biology (2017)
-
A moso bamboo WRKY gene PeWRKY83 confers salinity tolerance in transgenic Arabidopsis plants
Scientific Reports (2017)
-
Genome-wide investigation and transcriptome analysis of the WRKY gene family in Gossypium
Molecular Genetics and Genomics (2015)
-
Insights into the genome structure and copy-number variation of Eimeria tenella
BMC Genomics (2012)
-
A matricellular protein and EGF-like repeat signalling in the social amoebozoan Dictyostelium discoideum
Cellular and Molecular Life Sciences (2012)