Abstract
Background:
MicroRNAs (miRNAs) regulate the biological properties of colorectal cancer (CRC) cells and might serve as potential prognostic factors and therapeutic targets. In this study, we therefore globally profiled miRNAs associated with E-cadherin expression in CRC cells in an attempt to identify miRNAs that are associated with aggressive clinical course in CRC patients.
Methods:
Two CRC cell lines (Caco-2 and HRT-18) with different E-cadherin expression pattern were profiled for differences in abundance for more than 1000 human miRNAs using microarray technology. One of the most differentially expressed miRNAs, miR-200a was evaluated for its prognostic role in a cohort of 111 patients and independently validated in 217 patients of the Cancer Genome Atlas data set. To further characterise the biological role of miR-200a expression in CRC, in vitro miR-200a inhibition and overexpression were performed and the effects on cellular growth, apoptosis and epithelial–mesenchymal transition (EMT)-related gene expression were explored.
Results:
In situ hybridisation specifically localised miR-200a in CRC cells. In both cohorts, a low miR-200a expression was associated with poor survival (P<0.05). Multivariate Cox regression analysis identified low levels of miR-200a expression as an independent prognostic factor with respect to cancer-specific survival (HR=2.04, CI=1.28–3.25, P<0.002). Gain and loss of function assays for miR-200a in vitro led to a significantly differential and converse expression of EMT-related genes (P<0.001.) A low expression of miR-200a was also observed in cancer stem cell-enriched spheroid growth conditions (P<0.05).
Conclusions:
In conclusion, our data suggest that low miR-200a expression is associated with poor prognosis in CRC patients. MiR-200a has a regulatory effect on EMT and is associated with cancer stem cell properties in CRC.
Similar content being viewed by others
Main
Colorectal cancer (CRC) represents the third most commonly diagnosed cancer in males and ranks second in females, with 143 460 new cancer cases and 51 690 deaths estimated to occur in 2012 in the United States (Siegel et al, 2012). Despite the introduction of novel treatment options, including anti-angiogenic agents and targeted agents against the epidermal growth factor receptor, metastatic CRC still remains an incurable disease in most cases (Chua et al, 2012). Given the high number of CRC-related deaths, the toxic side effects of current treatment modalities and their high costs, there remains a clear need for further deciphering the pathogenesis of CRC and establish novel prognostic and potential therapeutic factors (Heitzer et al, 2013a, 2013b). In the last 10 years, the classical concept of tumour driving protein-coding genes was expanded by the identification of a novel class of small non-protein-coding RNA molecules known as microRNAs (miRNAs) (Calin and Croce, 2006). Due to their chemical stability, they provide great potential as novel diagnostic and prognostic biomarkers (Cortez et al, 2011; Pichler et al, 2012; Schwarzenbacher et al, 2013). Currently, several miRNAs have been established as differentially expressed and relevant for CRC pathogenesis including miR-150 (Ma et al, 2011), miR-28-5p/-3p (Schmittgen and Livak, 2008).
Profiling EMT-related genes on a RT 2 profiler PCR array
We used the Epithelial to Mesenchymal Transition RT2 Profiler PCR Arrays (Qiagen) to screen the mRNA expression of multiple genes related to EMT in HCT116 cells, which were transfected with the specific miR-200a inhibitor and mimetic in comparison with the respective control (miScript negative Control, Qiagen). RNA was isolated using the RNeasy Mini Kit (Qiagen) and 500 ng of total RNA was reverse transcribed into cDNA using the RT2 First Strand Kit (Qiagen) according to the manufacturer’s instructions. Quantitative RT–PCR was carried out in triplicates for each sample on a LightCycler 480 Real-Time PCR System (Roche Diagnostics) using RT2 Real-Time SYBR Green qPCR Master Mix (SABiosciences, Qiagen). All analyses were performed using the web-based PCR Array Data Analysis Software tool provided on the SABiosciences website. Geometric mean of the housekee** genes ACTB, GAPDH and RPLP0 was used for normalisation and fold change was used to analyse differences in gene expression.
WST-1 proliferation assay
To test whether miR-200a expression influences cellular growth rates of CRC cells, we performed modulation experiments with the miR-200a mimetic and inhibitor and measured the cellular growth rate by applying the WST-1 proliferation assay. In more detail, after standard trypsinisation 2 × 104 HCT116 cells per well were seeded in a 96-well culture plate. After transfection with a miR-200a mimetic, cells were incubated in regular McCoy’s 5A modified Medium growth medium (Biochrom) for 48 h and the WST-1 proliferation reagent (Roche Applied Science) was applied according to the manufacturer’s recommendations. After 4 h, colorimetric changes were measured using a SpectraMax Plus (Molecular Devices) at a wavelength of 450 nm with a reference wavelength at 620 nm.
Apoptosis assay
To clarify the role of miR-200a in apoptosis, we carried out a commercially available apoptosis assay after modulation of miR-200a. In more detail, 2 × 104 HCT116 cells were seeded in each well of a 96-well plate and were transiently transfected with Syn-Hsa-mir-200a (Qiagen) and Anti-Hsa-mir-200a (Qiagen). Caspase activity was measured after 24 and 48 h in HCT116 cells using the Caspase-Glo 3/7 Assay Systems (Promega, Madison, WI, USA) according to manufacturer’s instructions. Luminescence was measured in a POLARstar OPTIMA microplate reader (BMG Labtech, Ortenberg, Germany).
Tumour sphere formation
To measure differences in normal adherent HCT116 cells and tumour spheres, which are considered to be enriched of stem/progenitor cancer cells, we established a spheroid growth model as previously described (Fang et al, 2010). In detail, the adherent growing HCT116 cell line was dissociated into single cells using trypsin/EDTA and seeded in ultra-low attachment flasks (Corning) using in serum-free McCoy5A medium. Serum-free McCoy5A medium was supplemented with 1 × B27 supplement (GIBCO), 20 ng ml−1 human epidermal growth factor (Peprotech, Rocky Hill, NJ, USA), 20 ng ml−1 human basic fibroblast growth factor (Peprotech), 20 IU ml−1 Heparin (Baxter, Vienna, Austria), 4 IU l−1 insulin Actrapid (Novo Nordisk, Vienna, Austria) and 1% antibiotic/antimycotic solution (Sigma-Aldrich, Vienna, Austria). Self-renewing capacity of HCT116 spheres was shown by dissociating the spheres into single cells that were reseeded to yield the next generation of colon spheres. Spheres were maintained for several passages. Splitting and dissociation of colon spheres were done using TrypLE Select (GIBCO). RNA was extracted by miRNeasy Kit (Qiagen) and miR-200a levels were compared between adherent and spheroid growing cells.
Statistical analyses
All statistical analyses were performed using SPSS version 19.0 software (SPSS, Chicago, IL, USA). As reported previously, the continuous variable miR-200a was analysed as dichotomous variable according to approximate cut off point (Atzpodien et al, 2003). The value that best discriminated between good and poor survival, that is with the most significant P-value according to the log-rank test, was determined by testing all possible cut offs. This cut off was then rounded to clinically convenient value. For miR-200a expression, this means that a cut off near the normal expression level (i.e. neoplastic miR-200a expression level is approximately equal to normal colon mucosa) was identified as the optimal cut off, and therefore we defined low miRNA-200a expression as a lower expression level than that of adjacent non-cancerous tissue (n=60), whereas high miR-200a expression was defined as expression levels being higher than in adjacent non-tumour tissue (n=51). Fisher’s exact test, λ2test and Mann–Whitney were used to analyse the association between miR-200a expression and clinicopathological parameters. Cancer-specific survival (CSS) was defined as the time from date of diagnosis to the date of cancer-related death. Cancer-specific survival was assessed using the Kaplan–Meier method, and the log-rank test was performed to compare the survival curves of individual groups. Univariate and multivariate Cox proportional hazard models including age, gender, tumour stage (according to the AJCC/UICC 2010 TNM classification), tumour grade, number of therapies and miR-200a expression were used to determine the clinicopathological parameters that were statistically significant for CSS. The reported results included hazard ratios (HR) and 95% confidence intervals (CI). All experiments were at least repeated for three independent times.
Results
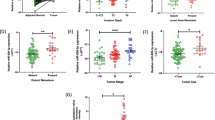
To identify miRNAs that determine the course of aggressive biological behaviour of CRC, we globally profiled the miRNA expression levels of two different CRC cell lines according to their E-cadherin expression pattern. The Caco-2 CRC cells exhibited a lower E-cadherin expression level with pronounced cytoplasmic staining pattern, in comparison with the HRT-18 CRC cell line, which showed a strong membranous E-Cadherin staining (Supplementary Figure 1). For detection of miRNA expression differences between both cell lines, we used a comprehensive miRNA detection system that detects more than 1000 human miRNAs. The rationale for using cell lines instead of CRC tissue was to avoid contamination of miRNAs from tumour microenvironment. A comparison of miRNA expression levels between the two different cell lines is shown in Figure 1. Overall, we identified 32 small RNAs including small nucleolar RNAs as significantly differentially expressed (P<0.05), and 19 of these RNAs remained significant after performing a Benjamini–Hochberg correction for multiple testing. The differentially expressed miRNAs included miR-200a that has been previously involved in the pathogenesis of many different human cancer types (Gregory et al, 2008). This prompted us to focus our further analysis on miR-200a and at first, we confirmed the array-based findings by qRT–PCR comparing the two cell lines (P<0.05, data not shown). To substantiate our findings from cell lines and to explore whether miR-200a deregulation might serve as a tissue-based biomarker that is not confounded by surrounding non-epithelial cells, we attempt to visualise and localise miR-200a expression within the CRC tissue by ISH. As demonstrated by ISH, miR-200a predominantly localises in CRC cells, whereas the surrounding non-epithelial cells showed only weak or no staining (Figure 2). The difference in expression in different cell lines and the specific localisation in CRC tissue, prompted us to test whether miR-200a expression might serve as a prognostic biomarker for CRC patients. Thus, we measured miR-200a expression on a collection of 111 CRC patients. Clinicopathological characteristics of the study cohort are summarised in Supplementary Table 1. Median follow-up was 38 months (interquartile range: 24–64). The median expression level of miR-200a was 0.88-fold (ranging from 0.11–10.3, Supplementary Table 2) lower in neoplastic tissue compared with adjacent non-neoplastic colon mucosa. For evaluating whether the levels of miR-200a expression influence survival of CRC patients, we divided the cohort into two groups according to a calculated cut off value, which was optimised as described above in the Material and Methods section (Atzpodien et al, 2003). The level of miR-200a expression was categorised into low (n=60) and high (n=51), respectively, compared with the expression in the corresponding non-neoplastic colon tissue. Subsequently, we used this cut off for clinicopathological parameters and survival analysis. Low miR-200a expression was not significantly associated with gender, tumour location, T stage, presence of metastasis at diagnosis, tumour grade and clinical stage (all P-values >0.05, Supplementary Table 3). Univariate analysis identified colonic location (colon versus rectum), high tumour grade (G3 versus G1+G2), advanced tumour stage (stage IV versus stage II–III), no metastasectomy and low expression of miR-200a as poor prognosticators for CSS (all P-values <0.05), whereas age, gender and number of therapy lines were not significantly associated with CSS (Table 1). The median survival for patients with high miR-200a expression was 61 months, whereas the median survival for patients with low miR-200a expression was 33 months (P<0.001). Figure 3 shows the Kaplan–Meier curve for CSS and reveals that low miR-200a expression is a significant factor for poor prognosis in CRC (P<0.001, log-rank test). To test whether the prognostic value of low miR-200a expression was independent of other risk factors for poor survival, a multivariate analysis was performed using a Cox proportional hazard model. Multivariate analyses including age, gender, tumour location, tumour grade, tumour stage, miR-200a expression, lines of therapy and metastasectomy demonstrated that low miR-200a expression was an independent predictor for poor CSS in CRC patients (HR=2.04, CI=1.28–3.25, P=0.002). Statistically significant results were also obtained for high tumour grade (P=0.02), advanced tumour stage (P<0.001), whereas all other parameters were not significant for CSS (Table 2).
Microarray-based miRNA expression profiling in the CRC cell lines Caco-2 versus HRT-18 revealed 19 miRNAs as significantly differentially expressed after Benjamini–Hochberg correction for multiple testing (red line) including miR-200a (red arrow). Red colour code represents an upregulation in HRT-18 cells, green colour code represents a downregulation compared with Caco-2 cells. The full colour version of this figure is available at British Journal of Cancer online.
In situ hybridisation to specifically localise the expression of miR-200a in CRC tissue. (A) Hematoxylin-eosin stained cancer tissue (CRC) and adjacent non-neoplastic tissue (N). (B) Positive control RNU6B localises all epithelial and non-epithelial cells (blue staining). (C and E) The specific probe localizes miR-200a mainly to invasive CRC formations. (D and F) Corresponding tissue probed with negative control for scrambled RNA is shown. Arrow denotes gland shown in high power view of inset (C and D). The full colour version of this figure is available at British Journal of Cancer online.
For further validation of the prognostic significance of miR-200a expression in an independent data set, we also analysed RNA seq data from the TCGA data set. In 217 colon cancer patients, we confirmed that a low expression of miR-200a is associated with poor survival (P=0.0166, Figure 4).
In an attempt to identify possible tumour characteristics and mechanisms that might be associated with worse clinical outcome in CRC patients with low miR-200a expression, we conducted in vitro overexpression/inhibition experiments with miR-200a mimetic/inhibitor. After transfection of miR-200a mimetic/inhibitor, we measured a dose-dependent increase of 500–4000-fold overexpression with the mimetic and a decrease of miR-200a expression between 30–50% by using the inhibitor, respectively (data not shown). First, we explored the role of miR-200a in EMT-related gene expression. Using a PCR array that contains a dozen of EMT-related genes, we screened whether the enhanced expression of miR-200a induces changes in gene expression of this gene set. Several genes including the well-known EMT-related E-cadherin (upregulation), vimentin or ZEB1 (downregulation) were differentially expressed (Supplementary Figure 2). We confirmed these findings by a RT–PCR assay for E-cadherin and vimentin, two well-known EMT-marker genes, which demonstrated converse effects on E-cadherin (miR-200a overexpression: 1.48±0.23-fold change versus miR-200a inhibition 0.87±0.09-fold change, P<0.001) and vimentin (miR-200a overexpression: 0.57±0.1-fold change versus miR-200a inhibition 1.1±0.05-fold change, P<0.001, Supplementary Figure 3). Regarding cell growth, proliferation-related gene expression or apoptotic activity, we could not observe any differences in miR-200a overexpressing HCT116 cells compared with miR-200a-inhibited cells (data not shown). Finally, we compared the expression of miR-200a in adherent versus spheroid growing HCT116 CRC cells, and detected a significant lower expression in the spheroid cancer stem cell model (0.48±0.07-fold change in spheroid cells versus adherent growing, P<0.05, Supplementary Figure 4).
Discussion
Novel biomarkers that are able to predict the clinical outcome of CRC patients will provide a valuable tool for risk stratification and patient surveillance and ideally provide a basis for decision making to avoid a potentially harmful, expensive and ineffective therapy. Different strategies have been performed to identify differentially expressed miRNAs in CRC, which often provided a comparison of normal versus cancerous tissue. However, contamination by cells of the tumour microenvironment, different regions of the cancerous tissue (tumour center or invasion front) and different sampling strategies (including different times of hypoxia and stress response) can impact miRNA levels in previously reported profiling studies (Iorio and Croce, 2012). Therefore, we first profiled miRNA expression based on differences in E-cadherin expression pattern in two different CRC cell lines for differences for more than 1000 miRNAs and identified several miRNAs as differentially expressed. One of these, miR-200a has been previously shown to be involved in regulation of E-cadherin expression as well as having a role in the pathogenesis of several types of cancer (Saydam et al, 2009). Therefore, we focused on miR-200a. Using in situ hybridisation, we confirmed specific expression in CRC cells, which provided the basis for further evaluation of the clinical value of miR-200a in a cohort of CRC patients. We identified low miR-200a expression as an independent prognostic parameter for CSS in CRC patients and independently confirmed our findings in a data set of an independent cohort. Encouraged by previously reported data about the role of miR-200 family as a regulator of EMT, we demonstrated by using two different assays an important role of miR-200a in EMT-related gene expression levels. In addition to EMT-related gene expression, we could also observe lower levels of miR-200a in spheroid growing cancer cells, a cell culture model that is assumed to contain more cancer stem cells (Fang et al, 2010). Thus, the shorter survival within the group of CRC patients with miR-200a low expression might be explained in part by differences in the EMT and stem cell formation capacity of their tumours. In line with our data, is the role of miR-200a that has been reported for the first time by Gregory et al (2008) who demonstrated that miR-200a was markedly downregulated in breast cancer cells that had undergone EMT, a process that is strongly associated with cancer cell invasion and metastases. On the molecular level, miR-200a suppresses the expression of the E-cadherin transcriptional repressors ZEB1 and ZEB2, transcriptions factors that induce the transition from an epithelial phenotype to a mesenchymal phenotype (Gregory et al, 2008). The transcriptional activity of miR-200a promoter region is repressed when ZEB1 and ZEB2 are upregulated through a conserved pair of ZEB-type E-box elements located proximal to the transcription start site (Bracken et al, 2008). Another self-reinforcing regulatory loop represents the interaction of miR-200a and SLUG, a protein that is not only a target of miR-200a but also directly represses miR-200a transcription in prostate adenocarcinoma (Liu et al, 2012). Recently, one study reported about the association of SIX1 and EMT in CRC. This study showed that SIX1 promotes EMT by repression of miR-200a in CRC cells (Ono et al, 2012). Wellner et al (2009) also linked the ZEB1 expression and members of the miR-200 family to tumour-initiating stem cells. Davalos et al (2012) demonstrated that miR-200a expression is commonly regulated by DNA methylation in human cancer. Recently, a study by Paterson (2013) showed that members of the miR-200 family are downregulated in the initial stages of stromal invasion. Besides the biological functions of members of the miR-200 family, they carry also great potential as non-invasive prognostic biomarkers. A very recently published study indicates that high miR-200c serum levels might represent a novel non-invasive prognostic marker in CRC patients (Toiyama et al, 2013).
In conclusion, this study indicates that high miR-200a expression is associated with membranous E-cadherin expression in CRC and a loss of miR-200a expression identifies patients with poor survival. MiR-200a regulates EMT-related gene expression and seems to be differentially expressed in CRC stem cells, which indicates this miRNA as a potential drug target in CRC.
Accession codes
Change history
18 March 2014
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Almeida MI, Nicoloso MS, Zeng L, Ivan C, Spizzo R, Gafa R, **ao L, Zhang X, Vannini I, Fanini F, Fabbri M, Lanza G, Reis RM, Zweidler-McKay PA, Calin GA (2012) Strand-specific miR-28-5p and miR-28-3p have distinct effects in colorectal cancer cells. Gastroenterology 142 (4): 886–896, e9.
Atzpodien J, Royston P, Wandert T, Reitz M (2003) Metastatic renal carcinoma comprehensive prognostic system. Br J Cancer 88 (3): 348–353.
Bartley AN, Yao H, Barkoh BA, Ivan C, Mishra BM, Rashid A, Calin GA, Luthra R, Hamilton SR (2011) Complex patterns of altered MicroRNA expression during the adenoma-adenocarcinoma sequence for microsatellite-stable colorectal cancer. Clin Cancer Res 17 (23): 7283–7293.
Bracken CP, Gregory PA, Kolesnikoff N, Bert AG, Wang J, Shannon MF, Goodall GJ (2008) A double-negative feedback loop between ZEB1-SIP1 and the microRNA-200 family regulates epithelial-mesenchymal transition. Cancer Res 68 (19): 7846–7854.
Buda A, Pignatelli M (2011) E-cadherin and the cytoskeletal network in colorectal cancer development and metastasis. Cell Commun Adhes 18 (6): 133–143.
Calin GA, Croce CM (2006) MicroRNA signatures in human cancers. Nat Rev Cancer 6 (11): 857–866.
Chen X, Wang Y, **a H, Wang Q, Jiang X, Lin Z, Ma Y, Yang Y, Hu M (2012) Loss of E-cadherin promotes the growth, invasion and drug resistance of colorectal cancer cells and is associated with liver metastasis. Mol Biol Rep 39 (6): 6707–6714.
Chua TC, Liauw W, Chu F, Morris DL (2012) Viewing metastatic colorectal cancer as a curable chronic disease. Am J Clin Oncol 35 (1): 77–80.
Cortez MA, Bueso-Ramos C, Ferdin J, Lopez-Berestein G, Sood AK, Calin GA (2011) MicroRNAs in body fluids—the mix of hormones and biomarkers. Nat Rev Clin Oncol 8 (8): 467–477.
Davalos V, Moutinho C, Villanueva A, Boque R, Silva P, Carneiro F, Esteller M (2012) Dynamic epigenetic regulation of the microRNA-200 family mediates epithelial and mesenchymal transitions in human tumorigenesis. Oncogene 31 (16): 2062–2074.
Fang DD, Kim YJ, Lee CN, Aggarwal S, McKinnon K, Mesmer D, Norton J, Birse CE, He T, Ruben SM, Moore PA (2010) Expansion of CD133(+) colon cancer cultures retaining stem cell properties to enable cancer stem cell target discovery. Br J Cancer 102 (8): 1265–1275.
Gregory PA, Bert AG, Paterson EL, Barry SC, Tsykin A, Farshid G, Vadas MA, Khew-Goodall Y, Goodall GJ (2008) The miR-200 family and miR-205 regulate epithelial to mesenchymal transition by targeting ZEB1 and SIP1. Nat Cell Biol 10 (5): 593–601.
Heitzer E, Auer M, Gasch C, Pichler M, Ulz P, Hoffmann EM, Lax S, Waldispuehl-Geigl J, Mauermann O, Lackner C, Hofler G, Eisner F, Sill H, Samonigg H, Pantel K, Riethdorf S, Bauernhofer T, Geigl JB, Speicher MR (2013a) Complex tumor genomes inferred from single circulating tumor cells by array-CGH and next-generation sequencing. Cancer Res 73 (10): 2965–2975.
Heitzer E, Auer M, Hoffmann EM, Pichler M, Gasch C, Ulz P, Lax S, Waldispuehl-Geigl J, Mauermann O, Mohan S, Pristauz G, Lackner C, Hofler G, Eisner F, Petru E, Sill H, Samonigg H, Pantel K, Riethdorf S, Bauernhofer T, Geigl JB, Speicher MR (2013b) Establishment of tumor-specific copy number alterations from plasma DNA of patients with cancer. Int J Cancer 133 (2): 346–356.
Iorio MV, Croce CM (2012) MicroRNA dysregulation in cancer: diagnostics, monitoring and therapeutics. A comprehensive review. EMBO Mol Med 4 (3): 143–159.
Liu YN, Yin JJ, Abou-Kheir W, Hynes PG, Casey OM, Fang L, Yi M, Stephens RM, Seng V, Sheppard-Tillman H, Martin P, Kelly K (2012) MiR-1 and miR-200 inhibit EMT via Slug-dependent and tumorigenesis via Slug-independent mechanisms. Oncogene 32 (3): 296–306.
Ma Y, Zhang P, Wang F, Zhang H, Yang J, Peng J, Liu W, Qin H (2011) miR-150 as a potential biomarker associated with prognosis and therapeutic outcome in colorectal cancer. Gut 61 (10): 1447–1453.
Ono H, Imoto I, Kozaki K, Tsuda H, Matsui T, Kurasawa Y, Muramatsu T, Sugihara K, Inazawa J (2012) SIX1 promotes epithelial-mesenchymal transition in colorectal cancer through ZEB1 activation. Oncogene 31 (47): 4923–4934.
Paterson EL, Kazenwadel J, Bert AG, Khew-Goodall Y, Ruszkiewicz A, Goodall GJ (2013) Down-regulation of the miRNA-200 family at the invasive front of colorectal cancers with degraded basement membrane indicates EMT is involved in cancer progression. Neoplasia 15 (2): 180–191.
Pichler M, Winter E, Stotz M, Eberhard K, Samonigg H, Lax S, Hoefler G (2012) Down-regulation of KRAS-interacting miRNA-143 predicts poor prognosis but not response to EGFR-targeted agents in colorectal cancer. Br J Cancer 106 (11): 1826–1832.
Pieler R, Sanchez-Cabo F, Hackl H, Thallinger GG, Trajanoski Z (2004) ArrayNorm: comprehensive normalization and analysis of microarray data. Bioinformatics 20 (12): 1971–1973.
Saydam O, Shen Y, Wurdinger T, Senol O, Boke E, James MF, Tannous BA, Stemmer-Rachamimov AO, Yi M, Stephens RM, Fraefel C, Gusella JF, Krichevsky AM, Breakefield XO (2009) Downregulated microRNA-200a in meningiomas promotes tumor growth by reducing E-cadherin and activating the Wnt/beta-catenin signaling pathway. Mol Cell Biol 29 (21): 5923–5940.
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 3 (6): 1101–1108.
Schwarzenbacher D, Balic M, Pichler M (2013) The role of microRNAs in breast cancer stem cells. Int J Mol Sci 14 (7): 14712–14723.
Siegel R, Naishadham D, Jemal A (2012) Cancer statistics, 2012. CA Cancer J Clin 62 (1): 10–29.
Sturn A, Quackenbush J, Trajanoski Z (2002) Genesis: cluster analysis of microarray data. Bioinformatics 18 (1): 207–208.
Toiyama Y, Hur K, Tanaka K, Inoue Y, Kusunoki M, Boland CR, Goel A (2013) Serum miR-200c is a novel prognostic and metastasis-predictive biomarker in patients with colorectal cancer. Ann Surg; e-pub ahead of print 26 August 2013; doi:10.1097/SLA.0b013e3182a6909d.
Van Cutsem E, Nordlinger B, Cervantes A (2010) Advanced colorectal cancer: ESMO Clinical Practice Guidelines for treatment. Ann Oncol 21 (Suppl 5): v93–v97.
Wellner U, Schubert J, Burk UC, Schmalhofer O, Zhu F, Sonntag A, Waldvogel B, Vannier C, Darling D, zur Hausen A, Brunton VG, Morton J, Sansom O, Schuler J, Stemmler MP, Herzberger C, Hopt U, Keck T, Brabletz S, Brabletz T (2009) The EMT-activator ZEB1 promotes tumorigenicity by repressing stemness-inhibiting microRNAs. Nat Cell Biol 11 (12): 1487–1495.
Acknowledgements
Martin Pichler was supported by the funding of the START Funding programme of the Medical University of Graz, Austria, the Styrian government, by an Erwin Schroedinger Scholarship of the Austrian Science Funds (No. J3389-B23) (all to MP) and by Oesterreichische Nationalbank (Anniversary Fund, project number: 14869). Armin Gerger is supported by research grants from the Oesterreichische Nationalbank (Anniversary Fund, project number: 15400) and the Austrian Science Fund (FWF): (KLI 191).
Author information
Authors and Affiliations
Corresponding author
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Pichler, M., Ress, A., Winter, E. et al. MiR-200a regulates epithelial to mesenchymal transition-related gene expression and determines prognosis in colorectal cancer patients. Br J Cancer 110, 1614–1621 (2014). https://doi.org/10.1038/bjc.2014.51
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2014.51
- Springer Nature Limited
Keywords
This article is cited by
-
LncRNA EBLN3P attributes methotrexate resistance in osteosarcoma cells through miR-200a-3p/O-GlcNAc transferase pathway
Journal of Orthopaedic Surgery and Research (2022)
-
A novel miRNA-based classification model of risks and stages for clear cell renal cell carcinoma patients
BMC Bioinformatics (2021)
-
CBFβ promotes colorectal cancer progression through transcriptionally activating OPN, FAM129A, and UPP1 in a RUNX2-dependent manner
Cell Death & Differentiation (2021)
-
An improved unsupervised learning approach for potential human microRNA–disease association inference using cluster knowledge
Network Modeling Analysis in Health Informatics and Bioinformatics (2021)
-
Integrated characterization and validation of the prognostic significance of microRNA-200s in colorectal cancer
Cancer Cell International (2020)