Abstract
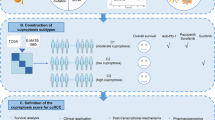
Cuproptosis is a type of programmed cell death triggered by accumulation of intracellular copper which was considered closely related to tumor progression. The study of cuproptosis in multiple myeloma (MM) is however limited. To determine the prognostic significance of cuproptosis-related gene signature in MM, we interrogated gene expression and overall survival with other available clinical variables from public datasets. Four cuproptosis-related genes were included to establish a prognostic survival model by least absolute shrinkage and selection operator (LASSO) Cox regression analysis, which showed a good performance on prognosis prediction in both training and validation cohorts. Patients with higher cuproptosis-related risk score (CRRS) exhibited worse prognosis compared with lower risk score. Survival prediction capacity and clinical benefit were elevated after integrating CRRS to existing prognostic stratification system (International Staging System, ISS or Revised International Staging System, RISS) both on 3-year and 5-year survival. Based on CRRS groups, functional enrichment analysis and immune infiltration in bone marrow microenvironment revealed correlation between CRRS and immunosuppression. In conclusion, our study found that cuproptosis-related gene signature is an independent poor prognostic factor and functions negatively on immune microenvironment, which provides another perspective on prognosis assessment and immunotherapy strategy in MM.









Similar content being viewed by others
Data Availability
Public datasets can be found here: [https://portal.gdc.cancer.gov/], [https://www.ncbi.nlm.nih.gov/geo/]. The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Brigle, K., & Rogers, B. (2017). Pathobiology and diagnosis of multiple myeloma. Seminars in Oncology Nursing, 33(3), 225–236. https://doi.org/10.1016/j.soncn.2017.05.012
Kazandjian, D. (2016). Multiple myeloma epidemiology and survival: A unique malignancy. Seminars in Oncology, 43(6), 676–681. https://doi.org/10.1053/j.seminoncol.2016.11.004
Rodriguez-Otero, P., Paiva, B., & San-Miguel, J. F. (2021). Roadmap to cure multiple myeloma. Cancer Treat Rev, 100, 102284. https://doi.org/10.1016/j.ctrv.2021.102284
Davies, F. E., Pawlyn, C., Usmani, S. Z., San-Miguel, J. F., Einsele, H., Boyle, E. M., Corre, J., Auclair, D., Cho, H. J., Lonial, S., et al. (2022). Perspectives on the risk-stratified treatment of multiple myeloma. Blood Cancer Discov, 3(4), 273–284. https://doi.org/10.1158/2643-3230.Bcd-21-0205
Ge, E. J., Bush, A. I., Casini, A., Cobine, P. A., Cross, J. R., DeNicola, G. M., Dou, Q. P., Franz, K. J., Gohil, V. M., Gupta, S., et al. (2022). Connecting copper and cancer: From transition metal signalling to metalloplasia. Nature Reviews Cancer, 22(2), 102–113. https://doi.org/10.1038/s41568-021-00417-2
Xue, Q., Kang, R., Klionsky, D. J., Tang, D., Liu, J., & Chen, X. (2023). Copper metabolism in cell death and autophagy. Autophagy. https://doi.org/10.1080/15548627.2023.2200554
Lopez, J., Ramchandani, D., & Vahdat, L. (2019). Copper depletion as a therapeutic strategy in cancer. Met Ions Life Sci. https://doi.org/10.1515/9783110527872-018
Adeoti, M. L., Oguntola, A. S., Akanni, E. O., Agodirin, O. S., & Oyeyemi, G. M. (2015). Trace elements; copper, zinc and selenium, in breast cancer afflicted female patients in LAUTECH Osogbo Nigeria. Indian Journal of Cancer, 52(1), 106–109. https://doi.org/10.4103/0019-509x.175573
Dìez, M., Cerdàn, F. J., Arroyo, M., & Balibrea, J. L. (1989). Use of the copper/zinc ratio in the diagnosis of lung cancer. Cancer, 63(4), 726–730. https://doi.org/10.1002/1097-0142(19890215)63:4%3c726::aid-cncr2820630421%3e3.0.co;2-p
Stepien, M., Jenab, M., Freisling, H., Becker, N. P., Czuban, M., Tjønneland, A., Olsen, A., Overvad, K., Boutron-Ruault, M. C., Mancini, F. R., et al. (2017). Pre-diagnostic copper and zinc biomarkers and colorectal cancer risk in the European Prospective Investigation into Cancer and Nutrition cohort. Carcinogenesis, 38(7), 699–707. https://doi.org/10.1093/carcin/bgx051
Saleh, S. A. K., Adly, H. M., Abdelkhaliq, A. A., & Nassir, A. M. (2020). Serum levels of selenium, zinc, copper, manganese, and iron in prostate cancer patients. Current Urology, 14(1), 44–49. https://doi.org/10.1159/000499261
Jiang, Y., Huo, Z., Qi, X., Zuo, T., & Wu, Z. (2022). Copper-induced tumor cell death mechanisms and antitumor theragnostic applications of copper complexes. Nanomedicine (London, England), 17(5), 303–324. https://doi.org/10.2217/nnm-2021-0374
Li, H., Wang, J., Wu, C., Wang, L., Chen, Z. S., & Cui, W. (2020). The combination of disulfiram and copper for cancer treatment. Drug Discovery Today, 25(6), 1099–1108. https://doi.org/10.1016/j.drudis.2020.04.003
Weyh, C., Krüger, K., Peeling, P., & Castell, L. (2022). The role of minerals in the optimal functioning of the immune system. Nutrients. https://doi.org/10.3390/nu14030644
Stabel, J. R., & Spears, J. W. (1989). Effect of copper on immune function and disease resistance. Advances in Experimental Medicine and Biology, 258, 243–252. https://doi.org/10.1007/978-1-4613-0537-8_22
Voli, F., Valli, E., Lerra, L., Kimpton, K., Saletta, F., Giorgi, F. M., Mercatelli, D., Rouaen, J. R. C., Shen, S., Murray, J. E., et al. (2020). Intratumoral copper modulates PD-L1 Expression and Influences Tumor Immune Evasion. Cancer Research, 80(19), 4129–4144. https://doi.org/10.1158/0008-5472.Can-20-0471
Tsvetkov, P., Coy, S., Petrova, B., Dreishpoon, M., Verma, A., Abdusamad, M., Rossen, J., Joesch-Cohen, L., Humeidi, R., Spangler, R. D., et al. (2022). Copper induces cell death by targeting lipoylated TCA cycle proteins. Science, 375(6586), 1254–1261. https://doi.org/10.1126/science.abf0529
Tang, D., Chen, X., & Kroemer, G. (2022). Cuproptosis: A copper-triggered modality of mitochondrial cell death. Cell Research, 32(5), 417–418. https://doi.org/10.1038/s41422-022-00653-7
Han, J., Hu, Y., Liu, S., Jiang, J., & Wang, H. (2022). A newly established cuproptosis-associated long non-coding RNA signature for predicting prognosis and indicating immune microenvironment features in soft tissue sarcoma. Journal of Oncology, 2022, 8489387. https://doi.org/10.1155/2022/8489387
Li, Z., Zhang, H., Wang, X., Wang, Q., Xue, J., Shi, Y., Wang, M., Wang, G., & Zhang, J. (2022). Identification of cuproptosis-related subtypes, characterization of tumor microenvironment infiltration, and development of a prognosis model in breast cancer. Frontiers in Immunology, 13, 996836. https://doi.org/10.3389/fimmu.2022.996836
Lv, H., Liu, X., Zeng, X., Liu, Y., Zhang, C., Zhang, Q., & Xu, J. (2022). Comprehensive analysis of cuproptosis-related genes in immune infiltration and prognosis in melanoma. Frontiers in Pharmacology, 13, 930041. https://doi.org/10.3389/fphar.2022.930041
Xu, Y., Zhou, Q., Feng, X., Dai, Y., Jiang, Y., Jiang, W., Liu, X., **ng, X., Wang, Y., Ni, Y., et al. (2020). Disulfiram/copper markedly induced myeloma cell apoptosis through activation of JNK and intrinsic and extrinsic apoptosis pathways. Biomedicine & Pharmacotherapy, 126, 110048. https://doi.org/10.1016/j.biopha.2020.110048
Agarwal, A., Khandelwal, A., Pal, K., Khare, N. K., Jadhav, V., Gurjar, M., Punatar, S., Gokarn, A., Bonda, A., Nayak, L., et al. (2022). A novel pro-oxidant combination of resveratrol and copper reduces transplant related toxicities in patients receiving high dose melphalan for multiple myeloma (RESCU 001). PLoS One, 17(2), e0262212. https://doi.org/10.1371/journal.pone.0262212
Liu, R., Shen, Y., Hu, J., Wang, X., Wu, D., Zhai, M., Bai, J., & He, A. (2021). Comprehensive analysis of m6A RNA methylation Regulators in the prognosis and immune microenvironment of multiple myeloma. Frontiers in Oncology, 11, 731957. https://doi.org/10.3389/fonc.2021.731957
Wu, T., Hu, E., Xu, S., Chen, M., Guo, P., Dai, Z., Feng, T., Zhou, L., Tang, W., Zhan, L., et al. (2021). clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. The Innovation (Camb), 2(3), 100141. https://doi.org/10.1016/j.xinn.2021.100141
Jia, Q., Wu, W., Wang, Y., Alexander, P. B., Sun, C., Gong, Z., Cheng, J. N., Sun, H., Guan, Y., **a, X., et al. (2018). Local mutational diversity drives intratumoral immune heterogeneity in non-small cell lung cancer. Nature Communications, 9(1), 5361. https://doi.org/10.1038/s41467-018-07767-w
Charoentong, P., Finotello, F., Angelova, M., Mayer, C., Efremova, M., Rieder, D., Hackl, H., & Trajanoski, Z. (2017). Pan-cancer immunogenomic analyses reveal genotype-immunophenotype relationships and predictors of response to checkpoint blockade. Cell Reports, 18(1), 248–262. https://doi.org/10.1016/j.celrep.2016.12.019
Wallington-Beddoe, C. T., & Mynott, R. L. (2021). Prognostic and predictive biomarker developments in multiple myeloma. Journal of Hematology & Oncology, 14(1), 151. https://doi.org/10.1186/s13045-021-01162-7
Szczepińska, T., Kalisiak, K., Tomecki, R., Labno, A., Borowski, L. S., Kulinski, T. M., Adamska, D., Kosinska, J., & Dziembowski, A. (2015). DIS3 shapes the RNA polymerase II transcriptome in humans by degrading a variety of unwanted transcripts. Genome Research, 25(11), 1622–1633. https://doi.org/10.1101/gr.189597.115
Boyle, E. M., Ashby, C., Tytarenko, R. G., Deshpande, S., Wang, H., Wang, Y., Rosenthal, A., Sawyer, J., Tian, E., Flynt, E., et al. (2020). BRAF and DIS3 mutations associate with adverse outcome in a long-term follow-up of patients with multiple myeloma. Clinical Cancer Research, 26(10), 2422–2432. https://doi.org/10.1158/1078-0432.Ccr-19-1507
Zhu, J., Kong, W., & **e, Z. (2021). Expression and prognostic characteristics of ferroptosis-related genes in colon cancer. International Journal of Molecular Sciences. https://doi.org/10.3390/ijms22115652
Tsvetkov, P., Detappe, A., Cai, K., Keys, H. R., Brune, Z., Ying, W., Thiru, P., Reidy, M., Kugener, G., Rossen, J., et al. (2019). Mitochondrial metabolism promotes adaptation to proteotoxic stress. Nature Chemical Biology, 15(7), 681–689. https://doi.org/10.1038/s41589-019-0291-9
Arroyo, J. D., Jourdain, A. A., Calvo, S. E., Ballarano, C. A., Doench, J. G., Root, D. E., & Mootha, V. K. (2016). A genome-wide CRISPR death screen identifies genes essential for oxidative phosphorylation. Cell Metabolism, 24(6), 875–885. https://doi.org/10.1016/j.cmet.2016.08.017
Yumnam, S., Kang, M. C., Oh, S. H., Kwon, H. C., Kim, J. C., Jung, E. S., Lee, C. H., Lee, A. Y., Hwang, J. I., & Kim, S. Y. (2021). Downregulation of dihydrolipoyl dehydrogenase by UVA suppresses melanoma progression via triggering oxidative stress and altering energy metabolism. Free Radical Biology & Medicine, 162, 77–87. https://doi.org/10.1016/j.freeradbiomed.2020.11.037
Prochownik, E. V., & Wang, H. (2021). The metabolic fates of pyruvate in normal and neoplastic cells. Cells. https://doi.org/10.3390/cells10040762
Deng, L., Jiang, A., Zeng, H., Peng, X., & Song, L. (2022). Comprehensive analyses of PDHA1 that serves as a predictive biomarker for immunotherapy response in cancer. Frontiers in Pharmacology, 13, 947372. https://doi.org/10.3389/fphar.2022.947372
Liggett, W. H., Jr., & Sidransky, D. (1998). Role of the p16 tumor suppressor gene in cancer. Journal of Clinical Oncology, 16(3), 1197–1206. https://doi.org/10.1200/jco.1998.16.3.1197
Bartsch, D. K., Sina-Frey, M., Lang, S., Wild, A., Gerdes, B., Barth, P., Kress, R., Grützmann, R., Colombo-Benkmann, M., Ziegler, A., et al. (2002). CDKN2A germline mutations in familial pancreatic cancer. Annals of Surgery, 236(6), 730–737. https://doi.org/10.1097/00000658-200212000-00005
Sievers, P., Hielscher, T., Schrimpf, D., Stichel, D., Reuss, D. E., Berghoff, A. S., Neidert, M. C., Wirsching, H. G., Mawrin, C., Ketter, R., et al. (2020). CDKN2A/B homozygous deletion is associated with early recurrence in meningiomas. Acta Neuropathologica, 140(3), 409–413. https://doi.org/10.1007/s00401-020-02188-w
Martínez-Baños, D., Sánchez-Hernández, B., Jiménez, G., Barrera-Lumbreras, G., & Barrales-Benítez, O. (2017). Global methylation and promoter-specific methylation of the P16, SOCS-1, E-cadherin, P73 and SHP-1 genes and their expression in patients with multiple myeloma during active disease and remission. Experimental and Therapeutic Medicine, 13(5), 2442–2450. https://doi.org/10.3892/etm.2017.4274
Mateos, M. V., García-Sanz, R., López-Pérez, R., Moro, M. J., Ocio, E., Hernández, J., Megido, M., Caballero, M. D., Fernández-Calvo, J., Bárez, A., et al. (2002). Methylation is an inactivating mechanism of the p16 gene in multiple myeloma associated with high plasma cell proliferation and short survival. British Journal of Haematology, 118(4), 1034–1040. https://doi.org/10.1046/j.1365-2141.2002.03749.x
Park, G., Kang, S. H., Lee, J. H., Suh, C., Kim, M., Park, S. M., Kim, T. Y., Oh, B., Min, H. J., Yoon, S. S., et al. (2011). Concurrent p16 methylation pattern as an adverse prognostic factor in multiple myeloma: A methylation-specific polymerase chain reaction study using two different primer sets. Annals of Hematology, 90(1), 73–79. https://doi.org/10.1007/s00277-010-1043-9
Dilworth, D., Liu, L., Stewart, A. K., Berenson, J. R., Lassam, N., & Hogg, D. (2000). Germline CDKN2A mutation implicated in predisposition to multiple myeloma. Blood, 95(5), 1869–1871.
Mitchell, J. S., Li, N., Weinhold, N., Försti, A., Ali, M., van Duin, M., Thorleifsson, G., Johnson, D. C., Chen, B., Halvarsson, B. M., et al. (2016). Genome-wide association study identifies multiple susceptibility loci for multiple myeloma. Nature Communications, 7, 12050. https://doi.org/10.1038/ncomms12050
Shah, V., Boyd, K. D., Houlston, R. S., & Kaiser, M. F. (2017). Constitutional mutation in CDKN2A is associated with long term survivorship in multiple myeloma: A case report. BMC Cancer, 17(1), 718. https://doi.org/10.1186/s12885-017-3715-5
Sautès-Fridman, C., Verneau, J., Sun, C. M., Moreira, M., Chen, T. W., Meylan, M., Petitprez, F., & Fridman, W. H. (2020). Tertiary Lymphoid Structures and B cells: Clinical impact and therapeutic modulation in cancer. Seminars in Immunology, 48, 101406. https://doi.org/10.1016/j.smim.2020.101406
Lopez-Yrigoyen, M., Cassetta, L., & Pollard, J. W. (2021). Macrophage targeting in cancer. Annals of the New York Academy of Sciences, 1499(1), 18–41. https://doi.org/10.1111/nyas.14377
Cui, C., Wang, J., Fagerberg, E., Chen, P. M., Connolly, K. A., Damo, M., Cheung, J. F., Mao, T., Askari, A. S., Chen, S., et al. (2021). Neoantigen-driven B cell and CD4 T follicular helper cell collaboration promotes anti-tumor CD8 T cell responses. Cell, 184(25), 6101-6118.e6113. https://doi.org/10.1016/j.cell.2021.11.007
Neefjes, J., Jongsma, M. L., Paul, P., & Bakke, O. (2011). Towards a systems understanding of MHC class I and MHC class II antigen presentation. Nature Reviews Immunology, 11(12), 823–836. https://doi.org/10.1038/nri3084
Li, B., Chan, H. L., & Chen, P. (2019). Immune checkpoint inhibitors: Basics and challenges. Current Medicinal Chemistry, 26(17), 3009–3025. https://doi.org/10.2174/0929867324666170804143706
Wei, G., Zhang, H., Zhao, H., Wang, J., Wu, N., Li, L., Wu, J., & Zhang, D. (2021). Emerging immune checkpoints in the tumor microenvironment: Implications for cancer immunotherapy. Cancer Letters, 511, 68–76. https://doi.org/10.1016/j.canlet.2021.04.021
Caserta, S., Innao, V., Musolino, C., & Allegra, A. (2020). Immune checkpoint inhibitors in multiple myeloma: A review of the literature. Pathology Research and Practice, 216(10), 153114. https://doi.org/10.1016/j.prp.2020.153114
Salik, B., Smyth, M. J., & Nakamura, K. (2020). Targeting immune checkpoints in hematological malignancies. Journal of Hematology & Oncology, 13(1), 111. https://doi.org/10.1186/s13045-020-00947-6
Acknowledgements
We appreciate all the participants and researchers for their contributions and commitment to this study.
Author information
Authors and Affiliations
Contributions
Conceptualization: HL; Methodology: HL and SC; Formal analysis and investigation: HL; Writing and original draft preparation: HL and SC; Writing, reviewing, and editing of the manuscript: HL and ML; Supervision: SC. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical Approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, H., Chan, S., Li, M. et al. Cuproptosis-Related Gene Signature Contributes to Prognostic Prediction and Immunosuppression in Multiple Myeloma. Mol Biotechnol 66, 475–488 (2024). https://doi.org/10.1007/s12033-023-00770-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-023-00770-7