Abstract
Populus alba ‘Berolinensis’ is a fast-growing, high-yielding species with strong biotic and abiotic stress resistance, and widely planted for timber, shelter belts and aesthetic purposes. In this study, molecular development is explored and the important genes regulating xylem formation in P. alba ‘Berolinensis’ under artificial bending treatments was identified. Anatomical investigation indicated that tension wood (TW) was characterized by eccentric growth of xylem and was enriched in cellulose; the degree of lignification was lower than for normal wood (NW) and opposite wood (OW). RNA-Seq-based transcriptome analysis was performed using develo** xylem from three wood types (TW, OW and NW). A large number of differentially expressed genes (DEGs) were screened and 4889 counted. In GO and KEGG enrichment results, genes involved in plant hormone signal transduction, phenylpropanoid biosynthesis, and cell wall and secondary cell wall biogenesis play major roles in xylem development under artificial bending. Eight expansin (PalEXP) genes were identified from the RNA-seq data; four were differentially expressed during tension wood formation. Phylogenetic analysis indicated that PalEXLB1 belongs to the EXPB subfamily and that the other PalEXPs are members of the EXPA subfamily. A transcriptional regulatory network construction showed 10 transcription factors located in the first and second layers upstream of EXP, including WRKY, ERF and bHLH. RT‒qPCR analysis in leaves, stems and roots combined with transcriptome analysis suggests that PalEXPA2, PalEXPA4 and PalEXPA15 play significant regulatory roles in cell wall formation during tension wood development. The candidate genes involved in xylem cell wall development during tension wood formation marks an important step toward identifying the molecular regulatory mechanism of xylem development and wood property improvement in P. alba ‘Berolinensis’.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As an important renewable resource, wood provides a large number of ecological services and biomass products. Wood formation depends on the growth of secondary xylem, which involves several processes, including cell division and expansion, secondary wall deposition, lignification, and programmed cell death (Mellerowicz et al 2001; Zhang et al 2020). Cellulose, lignin, hemicellulose and several proteins are the main components of cell walls (Rennie and Scheller 2014).
With regards to the bent or curved portions of woody dicotyledonous plant stems or branches, xylem tissue formed on the outside due to tensile force is called tension wood (TW), making the upper side of the stems or branches of trees thicker and eccentric to a certain extent (Timell 1969). Opposite wood (OW) refers to wood on the inner side of the bent wood (Pilate et al 2004a). Wood without any irregularities is termed normal wood (NW). Due to external forces, many plants produce a gelatinous fiber layer (G layer) inside tension wood cell walls, and its main component is crystalline cellulose (Pilate et al 2004b). Compared with normal wood, its appearance replaces the S3 layer of secondary cell walls (Andersson-Gunnerås et al 2006). Its most remarkable feature is that the cellulose content increases, making the cell wall thicker, and the lignin and xylan contents less. Anatomically, it has been found that the amount and size of vessels in tension wood are fewer, and fibers and vessels are longer. Certain chemical attributes have the potential to enhance the effectiveness of wood conversion for biofuel and material manufacturing (Yu et al 2021), but their physical properties are affected due to the presence of gelatinous fibers, making them unfavorable for fiber board (Vilkovská et al 2018). Improving the understanding of the molecular and genetic regulatory mechanisms involved in tension wood formation can facilitate strategic improvements in wood fiber and energy utilization and play a key role in promoting wood engineering to produce materials and energy more effectively.
Populus is a genus of fast-growing species and identifying the molecular mechanism of wood formation is of considerable significance for the improvement of timber resources. RNA sequencing (RNA-seq) has been used to explore the differentially expressed genes during tension wood development in various poplar species such as P. trichocarpa Torr. & A. Gray ex Hook. (Yu et al 2021), P. × euramericana ‘Zhonglin46’ (Liu et al 2023), P. davidiana × bolleana ‘Shanxin’ (Zhao et al 1999). It has rapidly become the most widely used variety of Populus in China because of its tolerance to cold and drought. Therefore, the identification of high-quality genes related to wood formation in P. alba ‘Berolinensis’ is of considerable significance for poplar wood improvement in semi-arid northeast China. In this study, the gene expression pathways regulating the development of tension wood were analyzed using transcriptome sequencing technology; the genes involved in the development of cell walls were screened. Eight EXP genes (PalEXPs) were screened in RNA-seq data, four were differentially expressed between TW, OW and NW. The bHLH, WRKY transcription factors were identified as upstream factors of PalEXP genes by transcriptional regulatory network construction, which revealed the molecular regulation mechanism in the formation of tension wood and provided information and gene resources for the improvement of wood properties using molecular biological methods.
Materials and methods
Plant materials and experimental treatment
Four-year-old clonal P. alba ‘Berolinensis’ saplings were obtained from the breeding base of Improved Variety Bases in Fulaerji District, Qiqihar, Heilongjiang province. The materials were cultivated outdoors (average temperature range 17–27 °C). The stems of the treatment group were bent in a lean angle of 45° in June (the period of active cambium cells and the develo** xylem). Straight stems were used as the control (Fig. S1). Five saplings were pooled into one sample with three replicates. A total 15 saplings were used for bending and another 15 were the control. After two weeks, stems of similar heights above the ground were removed from both groups. The bark was peeled and the xylem gently scraped away with a scalpel and collected. The upper side of the bent stems was removed and labelled as TW (tension wood), the lower side as OW (opposite wood), and the develo** xylem from the straight stems was collected as NW (normal wood). All samples were pre-cooled in liquid nitrogen and stored at −80 °C for subsequent transcriptome sequencing and quantitative reverse transcription polymerase chain reaction (RT-qPCR) experiments. Stem samples of 5 cm were used for anatomical analysis.
Anatomical analysis
A microscope slide was prepared of 3 cm of stem and fixed in FAA fixative (10% formalin, 5% acetic acid, 50% ethanol). To observe the develo** xylem, the fixed samples were sectioned with a sliding microtome (Leica 1007, Wetzlar, Germany). Paraffin sections were prepared with sequential steps of dehydration, transparency, wax dip**, embedding, block trimming, slicing, sticking and dewaxing. The samples were stained with safranin O-fast green or by the Wiesner (phloroglucinol-HCl) reaction (Pradhan Mitra et al. 2014; Chi et al 2023) and photographed under a stereo light microscope (Olympus SZX7, Tokyo, Japan) and an optical microscope (Olympus BX43, Tokyo, Japan).
RNA sequencing, de novo transcriptome assembly and assessment
The total RNA of nine samples (NW1, NW2, NW3, TW1, TW2, TW3, OW1, OW2, and OW3) was extracted by the CTAB method (Zhao et al 2012) and assessed using 1.0% agarose gel electrophoresis. The A260/280 ratio was measured by a NanoDrop 2000 UV–visible spectrophotometer (Thermo Scientific, Waltham, MA, USA). A library of standard-compliant RNA was constructed using the NEBNext® Ultra™ RNA Library Prep Kit, and the constructed library was sequenced on an Illumina® platform (New England Biolabs, Ipswich, MA, USA). Based on the filtered clean data, a full-length transcript sequence was assembled with Trinity (Trinity Release v2.4.0) (Grabherr et al 2011). The longest transcript sequence in each gene was taken as the unigene. All transcripts assembled by Trinity and unigene were evaluated by BUSCO to determine their integrity (Simão et al 2015). Based on the unigene sequence assembled by Trinity and the ORF sequence predicted by TransDecoder, the annotation information of the assembly result in the corresponding database was obtained by Tools including Blast (http://ww.ncbi.nlm.nih.gov/BLAST/), HmmScan (Finn et al 2011), SignalP (Petersen et al 2011), and TmHMMP (Krogh et al 2001), and the comprehensive functional annotation results were then obtained through Trinotate (Trinotate Release V3.0.2). The RNA-seq data were upload to the NCBI Short Read Archive (SRA) with the accession number PRJNA954564.
Expression level analysis and DEGs identification
FPKM was used for the quantitative estimation of unigene expression values (Trapnell 2010). Differential genes with |log2-fold change|≥ 1 and q < 0.05 were selected by DEG-seq (Wang 2010) as differentially expressed genes (DEGs). The data were normalized using the negative binomial distribution statistical method, and the resulting P value was subjected to multiple test correction to control false positives according to the Benjamini‒Hochberg procedure.
Gene function annotation, GO, KEGG analysis and identification of DEGs related to cell wall formation
The genes were compared in NR (NCBI nonredundant protein sequences), NT (nucleotide sequence database), and UniProt (universal protein resource) databases, and the function and characteristics of known nucleotide sequences were recorded. All DEGs were mapped to specific terms within the gene ontology (GO) database (http://www.geneontology.org/). Subsequently, the number of genes associated with each was determined. GO enrichment analysis was performed by the topGO R package to identify significantly enriched terms. KOBAS 3.0 software (http://kobas.cbi.pku.edu.cn/index.php) tested the statistical enrichment of DEGs in the Kyoto Encyclopedia of Genes and Genomes (KEGG, http://www.kegg.jp/) pathways (** or mature parts of the xylem; the black arrow points to G-layer; the red arrow points to annual ring of the previous year
The staining results show that the tension wood cell walls were rich in green (Fig. 1A). The develo** xylem cells near the cambium on the outer side of the TW, OW and NW were also green (Figs. 1A, D, B, F, C and H), indicating cellulose accumulation but lower lignin deposition. The mature xylem cells near the previous annual ring were stained red in the OW and NW (Figs. 1G and I) but were still green in TW (Fig. 1E). This indicates that the tension wood was rich in cellulose and higher than in OW and NW. The degree of lignification in OW and NW was higher than in TW. The existence of G-fibers, characteristic of TW markers, was found in mature tension wood stained green (Fig. 1E). The results of Weisner staining showed lighter red in TW and darker in OW and NW, indicating that the G-layer lacked lignin deposition and that the lignin in OW and NW was higher than in TW. (Figs. 1A', B', C', E', G' and I').
Transcriptome assembly and functional annotation of the unigenes in TW, OW and NW
Some 142,819,912 (TW), 146,012,770 (OW), and 152,134,284 (NW) raw reads were generated by RNA-seq. After filtering, 133,843,072 (TW), 133,709,274 (OW), and 141,165,714 (NW) clean reads with high quality were obtained. The Q30 values were above 90% in all samples and the data amount and base content were used for further analysis (Table S2). The proportion of clean reads for each sequence sample reflects the quality of sequencing data, and the analysis results show that the proportion of clean reads was close to 90%, indicating good quality data (Fig. S2). Altogether, 60,712 unigenes were obtained from the assembly results, of which the GC content was 39.9%. The length distribution of the assembled unigenes was as follows: the average length of the unigenes was 1017.09 bp, and the length of N50 was 1688 bp. Among these unigenes, 41,043 (67.6%) were > 400 bp in length, 21,706 (35.7%) were > 1000 bp, and 8666 (14.3%) were > 2000 bp (Table S3). BUSCO evaluation was conducted on all transcripts and unigenes assembled by Trinity. The number of missing BUSCOs was low, with a core protein alignment of 93.6%, annotation rate of 75.0%, and reads utilization rate of 75.5%. Accuracy, integrity, and core protein comparison evaluations indicated good transcriptome assembly quality (Table S3).
To characterize the gene functions involved in cell wall development in P. alba ‘Berolinensis’, the unigenes were annotated against public databases, including UniProt, NR, and NT. Annotated unigenes were counted, and the annotation of each database is shown using an UpSetR plot. The results showed that 18,382 genes were common to all databases (Fig. S3).
Expression level analysis of unigenes and identification of DEGs
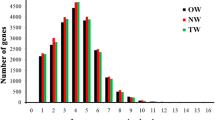
The fragments per kilobase per million (FPKM) method estimates gene expression levels using RNA-Seq (Trapnell 2010). In this study, the expression levels of unigenes in each sample were analyzed, and boxplots used to visualize the distribution of expression levels. The overall expression distribution was found to be similar across all samples, with only a small percentage of genes showing differential expression (Fig. S4). To investigate key genes involved in the regulation of xylem development during the formation of tension wood, differentially expressed genes (DEGs) were identified based on q < 0.05 and |log2_ratio|≥ 1 in the transcriptome data. A total of 4489 DEGs were identified in three comparisons. In the comparison between OW and NW, there were 1563 upregulated and 373 downregulated transcripts. In the comparison between TW and NW, there were 1407 upregulated and 507 downregulated transcripts and between TW and OW, there were 1326 upregulated and 1840 downregulated transcripts (Fig. S5A).
An upset plot was constructed to intuitively reflect the common and unique DEG numbers of each comparison group of the TW, OW and NW transcriptomes. There were 423 common DEGs between the OW vs. NW and TW vs. NW groups, 1123 between the TW vs. NW and TW vs. OW groups, and 139 DEGs common to all three groups. These results suggest that these 423 DEGs shared by OW vs. NW and TW vs. NW might be responsive to bending treatment stimulation, and the 1123 DEGs common to TW versus NW and TW versus OW might be involved in the development of tension wood (Fig. S5B).
GO enrichment analyses of DEGs
To analyze the biological function of DEGs involved in tension wood development, GO enrichment analyses were carried out. The figure shows that DEGs in GO terms, including cell wall organization or biogenesis and secondary cell wall biogenesis (biological processes), xylan O-acetyltransferase activity (molecular functions), and cell wall (cellular components), were enriched among the three groups. It is noteworthy that in the TW versus NW group, the DEGs were significantly highly enriched in cell wall organization or biogenesis, plant-type secondary cell wall biogenesis and hemicellulose metabolic process, among others (Fig. 2). In conclusion, the DEGs from TW versus NW and TW versus OW were enriched more in GO terms related to cell wall and secondary wall metabolic and biosynthesis processes, which suggests that transcripts involved in plant cell wall development play significant roles during the formation of tension wood in P. alba ‘Berolinensis’ and that these DEGs are key regulatory factors.
GO functional enrichment analysis of DEGs in P. alba ‘Berolinensis’. GO analysis of DEGs identified in OW vs. NW (A), TW versus NW (B), and TW versus OW (C). The x-axis represents the transcript ratios, and the y-axis the term enriched in BP (biological process), MF (molecular function) and CC (cellular component). The size of the point represents the transcript count; the color of the point represents the p value
KEGG enrichment analyses of DEGs
KEGG enrichment analysis further revealed the gene regulatory pathways in develo** xylem under bending stress. The DEGs in all three comparison groups were significantly enriched in the plant hormone signal transduction and phenylpropanoid biosynthesis pathways (Fig. 3), suggesting that these pathways played important roles in P. alba ‘Berolinensis’ xylem development in response to artificial bending.
Expression analysis and identification of DEGs related to cell wall formation
According to previous studies on genes associated with cell walls and through conducting GO and KEGG enrichment analyses, we identified DEGs associated with the formation of the cell wall in the transcriptomes of TW, OW, and NW. Most DEGs involved in lignin biosynthesis, like POD (peroxidase), CCR (cinnamoyl-CoA reductase), PAL (phenylalanine ammonia-lyase), CAD and CYP450 (cytochrome P450), were downregulated in TW vs. OW (Fig. 4A). Transcript levels of most DEGs encoding CesA (cellulose synthase) and SuSy (sucrose synthase), which are involved in cellulose synthesis, increased in TW vs. OW (Fig. 4B). Among the CesA transcripts, homologue of AtCesA8 (DN19943_c0_g3 and DN23646_c3_g1) were upregulated in TW vs. OW. Most DEGs involved in MAPK signaling pathway were upregulated in OW versus NW, downregulated in TW vs. OW. PR (pathogenesis-related protein) was significantly regulated among them (Fig. 4C). Based on the information provided, the expression levels of PEs (pectinesterases) were downregulated in the development of tension wood cell walls compared to opposite wood. FLAs (fasciclin-like arabinogalactan proteins) were upregulated in TW compared to NW and OW. EXPs were downregulated in OW compared to NW (Fig. 4D). In the process of plant hormone signal transduction, transcripts encoding AUX (auxin influx carriers), LAX and CCND3 (cyclin D3 protein) were upregulated in TW vs. OW and TW vs. NW. Transcripts encoding NPR3-like, AXR1 and BRU1-like were downregulated in TW compared with OW and NW (Fig. 4E). In total, 21 TFs shared among the three comparative groups were identified, including bHLH, ERF, WRKY, MYB, and NFY. These TF genes are mainly up-regulated in OW or TW compared to NW, indicating their involvement in the transcriptional regulation of xylem development in response to bending stimuli (Fig. 4F). These results indicate that the identified transcripts and TFs play significant roles in develo** xylem cell walls in response to artificial bending.
Candidate transcripts in response to artificial bending in P. alba ‘Berolinensis’. Heatmaps (A − F) show the expression of the DEGs identified in lignin biosynthesis (A), cellulose biosynthesis (B), MAPK signaling pathway (C), development of cell wall (D) plant hormone signal transduction (E) and transcription regulation (F) in response to artificial bending. Rows represent different transcripts and columns three kinds of groups (OW vs. NW, TW vs. NW, and TW vs. OW). The expression levels of transcripts are represented by squares, with red indicating up-regulation and blue indicating down-regulation. Differentially expressed genes (DEGs) were identified based on q < 0.05 and |log2_ratio|≥ 1, and then aligned with databases using the BLASTx algorithm at an E-value cutoff of 10−5
To validate the transcriptome analysis results, RT-qPCR analysis was carried out to assess the expression levels of genes associated with cell wall synthesis, including PAL (TRINITY_DN21468), PalCesA (TRINITY_DN23646), FLA (TRINITY_DN20133), EXPA2 (TRINITY_DN7305_c0_g1), EXPA4 (TRINITY_DN16617_c0_g3), EXPA15 (TRINITY_DN15609_c1_g2), and EXLB1 (TRINITY_DN8270_c0_g1). These genes exhibited differential expression between TW, OW, and NW transcriptomes. The results demonstrate a consistent expression pattern for PAL, CesA, FLA, and EXP genes in TW, OW, and NW when analyzed by both RT-qPCR and RNA-seq methods (Fig. 5).
Expression pattern validation of the RNA-seq DEG results by RT‒qPCR of PalPAL, PalCesA, PalFLA, PalEXPA2, PalEXPA4, PalEXLB1 and PalEXPA15. The expression was normalized to the tubulin gene. Relative expression levels of candidate transcripts were calculated by the 2–ΔΔCt method. Data are mean values ± standard error. Asterisks indicate statistically significant differences compared to the ratio obtained by RT‒qPCR alone (t-test; p < 0.05)
Identification and characterization of PalEXP family members using transcriptome data
Eight EXP family genes of P. alba ‘Berolinensis’ were screened from the transcriptome data. The protein lengths of PalEXPs ranged from 249 to 274 aa, with molecular weights from 26.59 to 30.12 kDa. The isoelectric points varied between 6.71 and 9.62, indicating that most of these proteins were hydrophilic (Table S4). By comparing and analyzing the amino acid sequences of the eight PalEXPs, it was observed that there were obvious conserved structural features between the sequences and that the C-terminal domains are relatively conserved (Fig. S6).
Phylogenetic analysis of EXP proteins
To explore the phylogenetic relationship of PalEXPs genes, a phylogenetic tree was constructed using EXP proteins from three species, P. alba ‘Berolinensis’, P. trichocarpa, and A. thaliana (Fig. 6). The results show that all the EXP proteins were clustered into four subfamilies (EXPA, EXPB, EXLA and EXLB) and named as PalEXPA2, PalEXPA3, PalEXPA4, PalEXPA6, PalEXPA13, PalEXPA15, PalEXPA16 and PalEXLB1 with Arabidopsis EXP proteins as reference according to the phylogenetic tree. Only PalEXLB1 was clustered into the EXLB subfamily, and the remaining PalEXPs proteins belonged to the EXPA subfamily.
Construction of transcriptional regulatory network of EXP upstream transcription factors
Eight PalEXP genes were identified, and four had different expressions between TW, OW and NW according to the comparative transcriptome analysis (Fig. 5). By identifying DEGs encoding TFs in each group, the transcriptional regulatory factors that control the expression of the PalEXPs can be identified (Fig. 4F). In total, 21 TFs are shared among the three comparative groups and a transcriptional regulatory network was constructed with four deferentially expressed PalEXPs using partial correlation coefficient algorithm. In this network, there were 10 TFs involved in the regulation of PalEXPs, and most belonged to WRKY, ERF, bHLH families (Fig. 7). Five of these are located in the first layer of the regulatory network, and they indirectly regulate EXPs by regulating five TFs in the second layer. These results show that these TFs are involved in the transcription regulation of PalEXPs when the xylem received external bending signals. The transcriptional regulation relationship needs further verification.
Tissue-specific expression analysis of PalEXP genes
The results of the RT‒qPCR analysis revealed distinct expression patterns of the PalEXP genes in different tissues of P. alba ‘Berolinensis’. PalEXPA2, PalEXPA4, and PalEXPA15 exhibited significantly higher expression levels in leaves compared to roots and stems. The expression levels were the lowest in roots. PalEXPA6 and PalEXPA3 showed high expression levels in stems and leaves, but relatively lower levels in roots. PalEXPA16 and PalEXPA13 exhibited slightly higher expression levels in stems compared to leaves. The highest expression levels of all four genes were shown in stems. PalEXLB1 was mainly expressed in roots (Fig. 8).