Abstract
The ubiquitous RNA-processing molecule TDP-43 is involved in neuromuscular diseases such as inclusion body myositis, a late-onset acquired inflammatory myopathy. TDP-43 solubility and function are disrupted in certain viral infections. Certain viruses, high viremia, co-infections, reactivation of latent viruses, and post-acute expansion of cytotoxic T cells may all contribute to inclusion body myositis, mainly in an age-shaped immune landscape. The virally induced senescent, interferon gamma-producing cytotoxic CD8+ T cells with increased inflammatory, and cytotoxic features are involved in the occurrence of inclusion body myositis in most such cases, in a genetically predisposed host. We discuss the putative mechanisms linking inclusion body myositis, TDP-43, and viral infections untangling the links between viruses, interferon, and neuromuscular degeneration could shed a light on the pathogenesis of the inclusion body myositis and other TDP-43-related neuromuscular diseases, with possible therapeutic implications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Inclusion body myositis (IBM) is an inflammatory myopathy occurring after middle age, with autoimmune and degenerative mechanisms [1, 2]. Other idiopathic inflammatory myopathies (IIMs) are dermatomyositis (DM), polymyositis (PM), overlap syndromes including anti-synthetase syndrome and necrotizing pauci-immune myositis [3]. The distinction between IBM, PM, and PM with mitochondrial pathology is not neat, raising the question whether IBM is a variant of PM occurring in the older age, related to immunosenescence [4]. IBM pathogenesis centrally involves cytotoxic, senescent CD8+ T cells, defects of autophagy and ubiquitin–proteasome system (UPS) resulting in proteostasis impairment and abnormal sarcoplasmic protein aggregation, along with endoplasmic reticulum and mitochondrial alterations, and antibodies to the cytosolic 5′-nucleotidase 1A (anti-cN1A) [1, 5]. The driving mechanisms of this pathology, however, are still evasive.
IBM belongs to a group of neurological disorders, the TDP-43 proteinopathies, which pathogenically involve TDP-43 [TAR-DNA-binding protein 43 (transactive response DNA-binding protein of 43 kDa)] [6]. TDP-43, encoded by the TARDBP gene, an RNA- and DNA-binding nuclear regulatory protein, member of the heterogeneous nuclear ribonucleoprotein (hnRNP) family [7, 8]. In skeletal muscles, TDP-43 is involved in transcription regulation, RNA splicing, mRNA stability, RNA transport, and quality control and undergoes post-translational modifications with functional consequences [9]. TDP-43 functions in muscles are complex, including myoregeneration (Table 1). In neurodegeneration, the mechanisms of TDP-43 involvement include cytotoxic aggregations, nuclear loss, alteration of cellular functions, and others [6, 10].
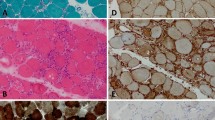
IBM muscle biopsies reveal cytoplasmic aggregation of TDP-43 and TDP-43 nuclear loss [10]. Even an 1% amount of myofibers staining for TDP-43 in a muscle biopsy was highly sensitive and specific for IBM [11].
TDP-43 may have an emerging intriguing role in viral infections [12]. TDP-43 is involved in controlling IFN responses triggered by endogenous RNA, but the TDP-43 role as an RNA-binding protein in viral infections is rarely investigated [13, 14]. Loss of TDP-43 results in dsRNA intracellular accumulation and interferon (IFN) triggering [13]. The TDP-43 ortholog of Caenorhabditis elegans called TDP1 limits dsRNA accumulation [21]. Also, knockdown of TARDBP increases viral replication in macrophages [14] and TDP-43 knockdown amplifies enterovirus infections, suggesting an antiviral effect of TDP-43 [22]. Moreover, TDP-43 binding is protective against HIV-1 by sterically hindering a HIV-1 promoter [23]. Also, after TDP-43 knockdown in mouse brain, the type I IFN-inducible genes, including the mouse orthologs of the intracellular sensor molecules RIG-I and MDA-5 which detect viral RNA, are the most overexpressed [21, 57]. IFN-ƴ induces ER stress and aggregation of TDP-43 and other proteins [1, 5, 54]. IFN-I may also be induced by anti-Ro52, present in some IBM patients [50, 58]. Ro52 or TRIM21 (tripartite motif proteins), is an IFN–inducible E3 ligase involved in IFN type I downregulation [59]. Other infection-related factors may intervene in IBM, such as activation of NLRP3 inflammasome, heat shock proteins (HSP), ribosomal proteins, or molecular mimicry with a mycobacterial protein guanylate-binding protein 2 (GBP2) with antiviral and anti-tuberculous functions [5, 60]. GBP2 is involved in the control of mRNA splicing [5] with possible relevance in TDP-43 dysfunction when mRNA splicing is altered.
Also, the glycogen synthase kinase 3 (GSK3), a serine/threonine kinase with 2 isoforms (α and β), is activated in IBM [33]. GSK3, involved in many cellular processes, is an immunomodulator in IBM [33]. GSK3 delays and decreases IFN-1 production, enhances IFNγ signaling, but also increases and delays pro-inflammatory cytokines production [33]. Moreover, GSK3β is one of the protein kinases involved in the TDP-43 phosphorylation [34]. TDP-43 expression activates GSK3, and GSK inhibition decreases TDP-43 aggregation [35].
Also, activation of autophagy is part of the innate immune response, and autophagy receptors may become viral targets [15]. Amongst these autophagy receptors, NBR1 (neighbor of BRCA1), a ubiquitin-binding scaffold protein, increases in viral infections [61], and NBR1 accumulates and is abnormal in IBM muscle [62].
In IBM, the dysregulation of a deubiquitinase called cylindromatosis (CYLD) reduces the autophagic clearance of protein aggregates [63]. CYLD is expressed with phosphorylated TDP-43 in the sIBM myofibers [63]. CYLD, required for antiviral host defense, is involved in the STING cleavage [64] and negatively regulates NF-kB [63].
IFN-ƴ and low RNA amounts in cytoplasm also stimulate aggregation of TDP-43 and other RBPs with “prion-like” low-complexity (LC) domains, favored by proteins misfolding in aging [1, 65].
Potential links between IBM and viral infections
The IBM occurrence may reflect various pathogenic associations, including viral infections [4]. In general, chronic IIM may be triggered by viruses such as Coxsackie B, enterovirus, parvovirus, HTLV-1, or HIV [66]. Mechanisms of viral-induced myositis hypothetically include direct invasion of myocytes by the virus, molecular mimicry, exposure of cryptic epitopes after conformational alterations, myotoxic cytokines such as IFNs and autoimmune reactions [66,67,68]. Latent viral infection, viral-induced denaturation of self-structures or homologies with various viral proteins could result in a prolonged immune response [66]. For instance, enterovirus 71 (EV71) may upregulate TRIM21 (Ro52), which degrades SAMHD1, a host antiviral molecule [59]. Also, during a viral infection, many ribonucleoproteins including TDP-43, are hijacked [12]. Coxsackie virus B3 protease 3C causes TDP-43 cytoplasmic redistribution and aggregation [12, 22].
Also, the aging cellular environment may make the myofiber susceptible to a newly invading virus, or may allow cytopathic manifestation of a virus, or a vertically transmitted genomic endogenous virus such as a retrovirus dormant for years, such as HTLV1, may start to be transcribed due to the age-modified milieu [16]. Endogenous retroviruses (ERVs, genomic remnants of ancient viral infections, most inactive and non-infectious) are mutually reinforcing with TDP-43 proteinopathies regarding neurodegeneration [17, 26]. Moreover, aging may favor both ERVs expression and TDP-43 proteinopathy [26].
However, no definite evidence for a viral etiology of IBM has been established [27]. Mumps virus was described as a potential IBM cause, later questioned in immunohistochemical studies [28]. IBM patients have an increased prevalence of hepatitis C virus (HCV) or human lymphotropic T virus-1 (HTLV1) [69,70,71]. The relationship between HCV and TDP-43 is yet to be clarified. TDP-43 binds YB (Y-box-binding protein-1), a host factor involved in HCV capsids assembling, and TDP-43 knockdown significantly decreased HCV replication [19]. The persistent HCV-related IFN upregulation and lymphocyte exhaustion may in fact contribute to the chronic myopathy in HCV [4]. TDP-43 facilitates HBV gene expression stimulating its transcription and assembly of protein complexes [12]. Furthermore, the clinical picture of IBM patients with HCV is different from the one of patients with IBM and HIV; therefore, no unique mechanism links a chronic viral infection to IBM [20].
Most of the HIV-positive patients with myositis had overlap** features of PM and IBM, which clinically progress to IBM, and most of them have anti-c1NA antibodies and rimmed vacuoles [20]. TDP-43 suppresses HIV-1 transcription by binding HIV-1 long terminal repeat [72]. Knocking down TDP-43 with siRNAs in cell cultures reactivates HIV-1 by reversing its latency [23]. Notwithstanding, HIV-1 can replicate in human immune cells independent of TDP-43 [73]. In viral-associated IBM in HIV and HTLV-1, the viral antigen is not present in myofibers but in the T cells and macrophages instead [1]. HIV infection can induce T cells immune senescence [74]. Thus, it is more conceivable that the virally induced senescent, IFN-ƴ producing cytotoxic CD8+ T cells lead to IBM.
IBM has been reported to be induced by Covid-19 in a 54-year female patient with diabetes mellitus and hyperlipidemia on statins [75]. Also, an axial paraspinal myopathy was reported in Covid-19 [76], and paraspinal myositis may be a feature of IBM [27]. However, long-term consequences of SARS-CoV2 infection, including muscular involvement, are starting to be recognized [77]. After COVID-19, the prevalence of myositis-specific antibodies and myositis-associated antibodies increases [78]. Possible mechanisms include type I IFN pathways, NLRP3 inflammasome activation, or a previous exposure to common coronaviruses [79]. SARS-CoV-2 impairs the stress granules (SGs) disassembly, and the SARS CoV-2 nucleocapsid N protein binds the SG-related amyloid proteins, favoring aggregation [89]. The HLA-DRB1*03 allele, as a component of the ancestral HLA 8.1 haplotype, is a susceptibility factor for IIMs and many other autoimmune diseases [90]. An arginine in position 74 of the DRβ1 chain confers the allelic risk for IBM [89]. HLA DRB1*01 is also associated with rheumatoid arthritis and hematologic malignancies, all overrepresented in IBM and associating age-related stochastic accumulation of CD8+ CD28- T cells [1, 86]. HLA-DRB1 alleles expression also impacts durable control of viral replication, HLA DR B1*03:01 being associated with high HIV viremia [91, 92], while HLA DRB1*01 was associated with spontaneous viral clearance of hepatitis C [92].
HLA DRB1*13 is common for IBM susceptibility and for protection against infection with several viruses, including HIV, HCV, HBV [87]. In IBM, the HLA DRB1*13:01 was associated with the highest age of onset and the lower strength [88]. Nevertheless, intriguingly, HLA DRB1*13 was protective against autoimmune diseases such as systemic lupus erythematosus, psoriasis, systemic sclerosis, and others [93]. However, HLA DRB1*13 is also associated with a slow progression of HIV [94]. HLA DRB1 *13 is associated with the clearance of hepatitis B as well [95]. Surprisingly, HLA DRB1*13 is neuroprotective, along with apoE, against age-related brain changes [96]. HLA-C*14:02:01 allele was higher in IBM patients with high LGL T cell expression [84]. HLA-C*14:02 allele was also associated with a T cell response in HIV-1 infection, which was nevertheless non-protective for the viral infection [97].
HLA-F, found in IBM and Sjogren’s syndrome, also elicits antiviral responses through activation of the KIR3DS1+ NK cells [20]. Therefore, HIV testing is advisable mostly in PM/IBM overlaps [20]. Also, pan-JAK inhibitors in aged mice alleviated the senescence—associated secretory phenotype but may also reactivate latent viruses [55]. Trials of immunosuppressive therapies in IBM have been recently nicely reviewed [105]. Immunosuppression is not routinely advised unless IBM is rapidly progressive or associated with other autoimmune diseases [105].
Followed both inflammatory and myodegenerative pathways presumed to be involved in IBM pathogenesis [105]. Most studies addressed inflammation or the involvement of T cells. Alemtuzumab (against CD52), natalizumab, anti-TNF alpha such as infliximab or etanercept, or IL-1 inhibitors as anakinra and canakinumab showed modest or no improvement [105], Rapamycin (sirolimus) targets mTOR important in IL-2 immune responses and protein metabolism (NCT04789070) [105]. Novel therapeutic avenues involve anti-KLRG1 antibodies, targeting a surface marker of the highly differentiated CD8T cells (NCT04659031) [84, 105]. Moreover, in HIV, the KLRG1 expression on NK cells correlates with HIV transcription, and targeting KLRG1 on NK cells potentially aids in elimination of HIV-infected cells [106]. Therapies against myodegeneration have recently become targets in clinical trials (arimoclomol, bimagrumab, follistatin, oxandrolone, rapamycin) [105].
Possible future directions may address other pathways. The attempts to reduce TDP-43 level led to muscle weakness and defective regeneration in myopathy models [6]. However, in neurological disorders such as ALS and other TDP-43-associated diseases, affecting skeletal and cardiac muscles besides neurons, there are several TDP-43 directed therapies [107, 108]. In ALS inhibition or deletion of cGAS and STING prevents TDP-43-induced upregulation of NF-kB and IFN type I [107]. Nevertheless, the neurological and muscular effects are not completely superposable [6].
Research including new therapies and repurposing for IBM some drugs used with other indications could serve as directions for the future [109]. Future therapeutic approaches could include inhibition of TDP-43 aggregation, the TDP-43-mitochondria association, proteasomal degradation of cytoplasmic TDP-43, or reducing TDP-43 aggregation-induced cell stress [37, 38, 110, 111]. Drugs stimulating the proteasome, such as chlorpromazine and other phenothiazines, methylene blue as a structural analogue of chlorpromazine and pyrazolones may target proteotoxic disorders [112]. The efficacy of zetomipzomib (KZR-616), a selective inhibitor of the immunoproteasome, is being studied in a phase 2 controlled multicenter study for active PM and DM [113, 114]. GSK3 inhibition decreases TDP-43 aggregation [34]. Lithium inhibits GSK-3 and induces autophagy, which may be relevant for IBM [115]. Also, lithium protected synapses from HIV-1 Tat-induced neuronal loss, in cultures and may be neuroprotective in HIV [116, 117]. Some other GSK3 inhibitors (including famotidine, naproxen, olanzapine, curcumin-all sterically hindering the enzyme binding pocket) may be tested for repurposing in IBM [33]. Also, regulating CYLD could be tested as a possible a therapeutic strategy in IBM [63].
The connection between a chronic viral infection and IBM deserves to be investigated further. There are questions waiting to be answered. Which factors are involved in transforming acute viral myositis into chronic inflammatory idiopathic myopathy? And moreover, why do some aged patients develop after a viral infection an IIM, for instance an anti-synthetase syndrome, and others an IBM? For instance, in HIV infection, what conditionate the switch from a PM phenotype to an IBM one? [4]. Serial studies in patients with chronic viral infections and signs of myopathy and/or sarcopenia would probably shed light on this progression, also regarding the progression to immunosenescence, mitochondrial dysfunction and proteinopathy, and the role of TDP-43 in this setting.
Conclusions
TDP-43 is important in preventing the dsRNA-induced IFN responses [13]. Viral infections may disrupt TDP-43 solubility and function, leading to its accumulation and lack of splicing regulation. The phenotypic differences between several IBM subtypes may be conditioned, besides genetic predisposing factors and age, also by environmental triggers such as certain viruses, and by epigenetic regulators [65]. Malat1 upregulation in certain viral infections may contribute to a protracted immune response [80].
Finding early disease markers and untangling mechanisms after a viral injury could inform whether there is a window of opportunity for the anti-inflammatory therapy, hopefully stop** or slowing the plethora of accompanying proteostasis, mitochondrial, and metabolic defects. Certain viruses, high viremia, coinfections, reactivation of latent viruses, and post-acute expansion of cytotoxic T cells may all contribute to IBM, mainly in an age-shaped immune landscape, with CD8+ T cells with IFN-ƴ production. In most such cases, the virally induced senescent, IFN-ƴ producing cytotoxic CD8+ T cells are the ones involved in IBM, in a genetically predisposed host. Immunophenoty** IBM patients to identify elevated CD8+ CD57+ populations may help stratify patients with prognostic and possibly therapeutic implications [84]. Identifying pathogenic mechanisms may lead to the identification of potential new treatments or to drug repurposing to improve the outcome in this debilitating disease.
Data availability
No datasets were generated or analysed during the current study.
Abbreviations
- aa:
-
Amino acid
- ALS:
-
Amyotrophic lateral sclerosis
- BRCA1:
-
Breast cancer susceptibility gene 1
- Anti-cN1A:
-
Antibodies against the cytosolic 5′-nucleotidase 1A
- CANDLE syndrome:
-
Chronic atypical neutrophilic dermatosis with lipodystrophy and elevated temperature
- cGAS:
-
Cyclic GMP–AMP synthase
- CHCHD10:
-
Coiled-coil-helix-coiled-coil-helix domain-containing protein 10
- CMV:
-
Cytomegalovirus
- CYLD:
-
Cylindromatosis, a deubiquitinating enzyme that negatively regulates signal transduction pathways, such as NF-kB signaling pathways
- DM:
-
Dermatomyositis
- dsRNA:
-
Double-stranded RNA
- EBV:
-
Epstein–Barr virus
- ERV:
-
Endogenous retroviruses
- GBP2:
-
Guanylate-binding protein 2
- GSK3:
-
Glycogen synthase kinase 3
- HCV:
-
Hepatitis C virus
- HIV:
-
Human immunodeficiency virus
- hnRNP:
-
Heterogeneous nuclear ribonucleoprotein
- HSPs:
-
Heat shock proteins
- HTLV1:
-
Human T-cell leukemia virus type 1
- IFN:
-
Interferon
- IBM:
-
Inclusion body myositis
- IIMs:
-
Idiopathic inflammatory myopathies
- iPS:
-
Immunoproteasomes
- IRF:
-
Interferon regulatory factor
- lncRNA:
-
Long non-coding RNA
- Malat1/MALAT1:
-
Metastasis-associated lung adenocarcinoma transcript-1
- MDA-5:
-
Melanoma differentiation-associated protein 5
- MHC:
-
Major histocompatibility complex
- miRNA:
-
MicroRNA
- mRNA:
-
Messenger RNA
- NBR1:
-
Neighbor of BRCA1
- NF-kB:
-
Nuclear factor kappa B
- NK:
-
Natural killer cells
- NLRP3:
-
NOD-, LRR-, and pyrin domain-containing protein 3
- PASC:
-
Post-acute sequelae SARS-CoV-2 infection
- PM:
-
Polymyositis
- PSMB8:
-
Proteasome subunit beta type-8
- Rbck1:
-
RanBP-type and C3HC4-type zinc finger-containing protein 1
- RBP:
-
RNA-binding proteins
- RIG-I:
-
Retinoic acid-inducible gene-I
- RNA:
-
Ribonucleic acid
- RRM:
-
RNA-recognition motif
- SARS-CoV-2:
-
Severe acute respiratory syndrome coronavirus 2
- SARS-CoV2 S1 RBD:
-
SARS-CoV-2 spike S1 protein receptor binding domain
- SAMHD1:
-
Sterile alpha motif domain and histidine-aspartate domain-containing protein 1
- SGs:
-
Stress granules
- ssRNA:
-
Single-stranded RNA
- STAT:
-
Signal transducer and activator of transcription
- STING:
-
Stimulator of interferon genes
- TARDBP:
-
TAR-DNA-binding-protein 43 (transactive response DNA-binding protein of 43 kDa)
- TDP-43:
-
TAR-DNA-binding protein 43
- TEMRA:
-
Effector memory T cells re-expressing CD45RA
- TRIM21:
-
Tripartite motif containing 21
- UPS:
-
Ubiquitin-proteasome system
- YB:
-
Y-box-binding protein-1
References
Greenberg SA. Inclusion body myositis: clinical features and pathogenesis. Nat Rev Rheumatol. 2019;15(5):257–72. https://doi.org/10.1038/s41584-019-0186-x.
McLeish E, Slater N, Sooda A, Wilson A, Coudert JD, Lloyd TE, Needham M. Inclusion body myositis: the interplay between ageing, muscle degeneration and autoimmunity. Best Pract Res Clin Rheumatol. 2022;36(2):101761. https://doi.org/10.1016/j.berh.2022.101761.
Lundberg IE, de Visser M, Werth VP. Classification of myositis. Nat Rev Rheumatol. 2018;14(5):269–78. https://doi.org/10.1038/nrrheum.2018.41.
Nelke C, Kleefeld F, Preusse C, Ruck T, Stenzel W. Inclusion body myositis and associated diseases: an argument for shared immune pathologies. Acta Neuropathol Commun. 2022;10(1):84. https://doi.org/10.1186/s40478-022-01389-6.
Snedden AM, Kellett KAB, Lilleker JB, Hooper NM, Chinoy H. The role of protein aggregation in the pathogenesis of inclusion body myositis. Clin Exp Rheumatol. 2022;40(2):414–24. https://doi.org/10.55563/clinexprheumatol/pp0oso.
Versluys L, Ervilha Pereira P, Schuermans N, De Paepe B, De Bleecker JL, Bogaert E, Dermaut B. Expanding the TDP-43 proteinopathy pathway from neurons to muscle: physiological and pathophysiological functions. Front Neurosci. 2022;16:815765. https://doi.org/10.3389/fnins.2022.815765.
Weihl CC, Temiz P, Miller SE, Watts G, Smith C, Forman M, Hanson PI, et al. TDP-43 accumulation in inclusion body myopathy muscle suggests a common pathogenic mechanism with frontotemporal dementia. J Neurol Neurosurg Psychiatry. 2008;79(10):1186–9. https://doi.org/10.1136/jnnp.2007.131334.
Šušnjar U, Škrabar N, Brown AL, Abbassi Y, Phatnani H, NYGC ALS Consortium, Cortese A, et al. Cell environment shapes TDP-43 function with implications in neuronal and muscle disease. Commun Biol. 2022;5(1):314. https://doi.org/10.1038/s42003-022-03253-8.
Buratti E. TDP-43 post-translational modifications in health and disease. Expert Opin Ther Targets. 2018;22(3):279–93. https://doi.org/10.1080/14728222.2018.1439923.
Britson KA, Ling JP, Braunstein KE, Montagne JM, Kastenschmidt JM, Wilson A, Ikenaga C, et al. Loss of TDP-43 function and rimmed vacuoles persist after T cell depletion in a xenograft model of sporadic inclusion body myositis. Sci Transl Med. 2022;14(628):eabi9196. https://doi.org/10.1126/scitranslmed.abi9196.
Salajegheh M, Pinkus JL, Taylor JP, Amato AA, Nazareno R, Baloh RH, Greenberg SA. Sarcoplasmic redistribution of nuclear TDP-43 in inclusion body myositis. Muscle Nerve. 2009;40(1):19–31. https://doi.org/10.1002/mus.21386.
Rahic Z, Buratti E, Cappelli S. Reviewing the potential links between viral infections and TDP-43 proteinopathies. Int J Mol Sci. 2023;24(2):1581. https://doi.org/10.3390/ijms24021581.
Dunker W, Ye X, Zhao Y, Liu L, Richardson A, Karijolich J. TDP-43 prevents endogenous RNAs from triggering a lethal RIG-I-dependent interferon response. Cell Rep. 2021;35(2):108976. https://doi.org/10.1016/j.celrep.2021.108976.
Liu W, Wang Z, Liu L, Yang Z, Liu S, Ma Z, Liu Y, et al. LncRNA Malat1 inhibition of TDP43 cleavage suppresses IRF3-initiated antiviral innate immunity. Proc Natl Acad Sci U S A. 2020;117(38):23695–706. https://doi.org/10.1073/pnas.2003932117.
Ylä-Anttila P. Autophagy receptors as viral targets. Cell Mol Biol Lett. 2021;26(1):29. https://doi.org/10.1186/s11658-021-00272-x.
Askanas V, Engel WK. Inclusion-body myositis: newest concepts of pathogenesis and relation to aging and Alzheimer disease. J Neuropathol Exp Neurol. 2001;60(1):1–14. https://doi.org/10.1093/jnen/60.1.1.
Dubowsky M, Theunissen F, Carr JM, Rogers ML. The molecular link between TDP-43, endogenous retroviruses and inflammatory neurodegeneration in amyotrophic lateral sclerosis: a potential target for Triumeq, an antiretroviral therapy. Mol Neurobiol. 2023;60(11):6330–45. https://doi.org/10.1007/s12035-023-03472-y.
Morosetti R, Broccolini A, Sancricca C, Gliubizzi C, Gidaro T, Tonali PA, Ricci E, Mirabella M. Increased aging in primary muscle cultures of sporadic inclusion-body myositis. Neurobiol Aging. 2010;31(7):1205–14. https://doi.org/10.1016/j.neurobiolaging.2008.08.011.
Germain MA, Chatel-Chaix L, Gagné B, Bonneil É, Thibault P, Pradezynski F, de Chassey B, et al. Elucidating novel hepatitis C virus-host interactions using combined mass spectrometry and functional genomics approaches. Mol Cell Proteomics. 2014;13(1):184–203. https://doi.org/10.1074/mcp.M113.030155.
Lloyd TE, Pinal-Fernandez I, Michelle EH, Christopher-Stine L, Pak K, Sacktor N, Mammen AL. Overlap** features of polymyositis and inclusion body myositis in HIV-infected patients. Neurology. 2017;88(15):1454–60. https://doi.org/10.1212/WNL.0000000000003821.
Saldi TK, Ash PE, Wilson G, Gonzales P, Garrido-Lecca A, Roberts CM, Dostal V, et al. TDP-1, the Caenorhabditis elegans ortholog of TDP-43, limits the accumulation of double-stranded RNA. EMBO J. 2014;33(24):2947–66. https://doi.org/10.15252/embj.201488740.
Fung G, Shi J, Deng H, Hou J, Wang C, Hong A, Zhang J, et al. Cytoplasmic translocation, aggregation, and cleavage of TDP-43 by enteroviral proteases modulate viral pathogenesis. Cell Death Differ. 2015;22(12):2087–97. https://doi.org/10.1038/cdd.2015.58.
Rathore A, Iketani S, Wang P, Jia M, Sahi V, Ho DD. CRISPR-based gene knockout screens reveal deubiquitinases involved in HIV-1 latency in two Jurkat cell models. Sci Rep. 2020;10(1):5350. https://doi.org/10.1038/s41598-020-62375-3.
Li Y, Lu S, Gu J, **a W, Zhang S, Zhang S, Wang Y, et al. SARS-CoV-2 impairs the disassembly of stress granules and promotes ALS-associated amyloid aggregation. Protein Cell. 2022;13(8):602–14. https://doi.org/10.1007/s13238-022-00905-7.
Xue Y, Zhang J, Ke J, Zeng L, Cheng K, Han X, Chen F, et al. LncGBP9 knockdown alleviates myocardial inflammation and apoptosis in mice with acute viral myocarditis via suppressing NF-kappaB signaling pathamianway. Inflamm Res. 2022;71(12):1559–76. https://doi.org/10.1007/s00011-022-01644-5.
Chang YH, Dubnau J. Endogenous retroviruses and TDP-43 proteinopathy form a sustaining feedback driving intercellular spread of Drosophila neurodegeneration. Nat Commun. 2023;14(1):966. https://doi.org/10.1038/s41467-023-36649-.
Keller CW, Schmidt J, Lunemann JD. Immune and myodegenerative pathomechanisms in inclusion body myositis. Ann Clin Transl Neurol. 2017;4(6):422–45. https://doi.org/10.1002/acn3.419.
Nishino H, Engel AG, Rima BK. Inclusion body myositis: the mumps virus hypothesis. Ann Neurol. 1989;25(3):260–4. https://doi.org/10.1002/ana.410250309.
Agergaard J, Leth S, Pedersen TH, Harbo T, Blicher JU, Karlsson P, Ostergaard L, et al. Myopathic changes in patients with long-term fatigue after COVID-19. Clin Neurophysiol. 2021;132(8):1974–81. https://doi.org/10.1016/j.clinph.2021.04.009.
Galeotti C, Bayry J. Autoimmune and inflammatory diseases following COVID-19. Nat Rev Rheumatol. 2020;16(8):413–4. https://doi.org/10.1038/s41584-020-0448-7.
Nguyen TM, Kabotyanski EB, Reineke LC, Shao J, **ong F, Lee JH, Dubrulle J, et al. The SINEB1 element in the long non-coding RNA Malat1 is necessary for TDP-43 proteostasis. Nucleic Acids Res. 2020;8(5):2621–42. https://doi.org/10.1093/nar/gkz1176.
Li XL, Ezelle HJ, Hsi TY, Hassel BA. A central role for RNA in the induction and biological activities of type 1 interferons. Wiley Interdiscip Rev RNA. 2011;2(1):58–78. https://doi.org/10.1002/wrna.32.
Piazzi M, Bavelloni A, Cenni V, Faenza I, Blalock WL. Revisiting the role of GSK3, a modulator of innate immunity, in idiopathic inclusion body myositis. Cells. 2021;10(11):3255. https://doi.org/10.3390/cells10113255.
Sreedharan J, Neukomm LJ, Brown Jr RH, Freeman MR. Age-dependent TDP-43-mediated motor neuron degeneration requires GSK3, hat-trick, and xmas-2. Curr Biol. 2015;25(16):2130–6. https://doi.org/10.1016/j.cub.2015.06.045.
Liu XL, Sun DD, Zheng MT, Li XT, Niu HH, Zhang L, Zhou ZW, et al. Maraviroc promotes recovery from traumatic brain injury in mice by suppression of neuroinflammation and activation of neurotoxic reactive astrocytes. Neural Regen Res. 2023;18(1):141–9. https://doi.org/10.4103/1673-5374.344829.
Medeiros GA, Silvério JC, Marino AP, Roffê E, Vieira V, Kroll-Palhares K, Carvalho CE, Silva AA, et al. Treatment of chronically Trypanosoma cruzi-infected mice with a CCR1/CCR5 antagonist (Met-RANTES) results in amelioration of cardiac tissue damage. Microbes Infect. 2009;11(2):264–73. https://doi.org/10.1016/j.micinf.2008.11.012.
Webber CJ, Murphy CN, Rondón-Ortiz AN, van der Spek SJF, Kelly EX, Lampl NM, Chiesa G, et al. Human herpesvirus 8 ORF57 protein is able to reduce TDP-43 pathology: network analysis identifies interacting pathways. Hum Mol Genet. 2023;32(20):2966–80. https://doi.org/10.1093/hmg/ddad122.
Huntley ML, Gao J, Termsarasab P, Wang L, Zeng S, Thammongkolchai T, Liu Y, et al. Association between TDP-43 and mitochondria in inclusion body myositis. Lab Invest. 2019;99(7):1041–8. https://doi.org/10.1038/s41374-019-0233-x.
Idrees D, Kumar V. SARS-CoV-2 spike protein interactions with amyloidogenic proteins: potential clues to neurodegeneration. Biochem Biophys Res Commun. 2021;554:94–8. https://doi.org/10.1016/j.bbrc.2021.03.100.
Huang K, Wang C, Vagts C, Raguveer V, Finn PW, Perkins DL. Long non-coding RNAs (lncRNAs) NEAT1 and MALAT1 are differentially expressed in severe COVID-19 patients: an integrated single-cell analysis. PLoS ONE. 2022;17(1):e0261242. https://doi.org/10.1371/journal.pone.0261242.
Menon MP, Hua KF. The long non-coding RNAs: paramount regulators of the NLRP3 inflammasome. Front Immunol. 2020;11:569524. https://doi.org/10.3389/fimmu.2020.569524.
Hamann PD, Roux BT, Heward JA, Love S, McHugh NJ, Jones SW, Lindsay MA. Transcriptional profiling identifies differential expression of long non-coding RNAs in Jo-1 associated and inclusion body myositis. Sci Rep. 2017;7(1):8024. https://doi.org/10.1038/s41598-017-08603-9.
Askanas V, Engel WK, Nogalska A. Inclusion body myositis: a degenerative muscle disease associated with intra-muscle fiber multi-protein aggregates, proteasome inhibition, endoplasmic reticulum stress and decreased lysosomal degradation. Brain Pathol. 2009;19(3):493–506. https://doi.org/10.1111/j.1750-3639.2009.00290.x.
Ghannam K, Martinez-Gamboa L, Spengler L, Krause S, Smiljanovic B, Bonin M, Bhattarai S, et al. Upregulation of immunoproteasome subunits in myositis indicates active inflammation with involvement of antigen presenting cells, CD8 T-cells and IFNGamma. PLoS ONE. 2014;9(8):e104048. https://doi.org/10.1371/journal.pone.0104048.
van den Eshof BL, Medfai L, Nolfi E, Wawrzyniuk M, Sijts AJAM. The function of immunoproteasomes—an immunologists’ perspective. Cells. 2021;10(12):3360. https://doi.org/10.3390/cells10123360.
Wellington D, Yin Z, Yu Z, Heilig R, Davis S, Fischer R, Felce SL, et al. SARS-CoV-2 mutations affect antigen processing by the proteasome to alter CD8+ T cell responses. Heliyon. 2023;9(10):e20076. https://doi.org/10.1016/j.heliyon.2023.e20076.
Ferrer I, Martin B, Castano JG, Lucas JJ, Moreno D, Olive M. Proteasomal expression, induction of immunoproteasome subunits, and local MHC class I presentation in myofibrillar myopathy and inclusion body myositis. J Neuropathol Exp Neurol. 2004;63(5):484–98. https://doi.org/10.1093/jnen/63.5.484.
Bolko L, Jiang W, Tawara N, Landon-Cardinal O, Anquetil C, Benveniste O, Allenbach Y. The role of interferons type I, II and III in myositis: a review. Brain Pathol. 2021;31(3):e12955. https://doi.org/10.1111/bpa.12955.
Ivanidze J, Hoffmann R, Lochmuller H, Engel AG, Hohlfeld R, Dornmair K. Inclusion body myositis: laser microdissection reveals differential up-regulation of IFN-gamma signaling cascade in attacked versus nonattacked myofibers. Am J Pathol. 2011;179(3):1347–59. https://doi.org/10.1016/j.ajpath.2011.05.055.
Ekholm L, Vosslamber S, Tjarnlund A, de Jong TD, Betteridge Z, McHugh N, Plestilova L, et al. Autoantibody specificities and type I interferon pathway activation in idiopathic inflammatory myopathies. Scand J Immunol. 2016;84(2):100–9. https://doi.org/10.1111/sji.12449.
Angeles A, Fung G, Luo H. Immune and non-immune functions of the immunoproteasome. Front Biosci (Landmark Ed). 2012;17(5):1904–16. https://doi.org/10.1093/jnen/60.1.1.
Ayaki T, Murata K, Kanazawa N, Uruha A, Ohmura K, Sugie K, Kasagi S, et al. Myositis with sarcoplasmic inclusions in Nakajo-Nishimura syndrome: a genetic inflammatory myopathy. Neuropathol Appl Neurobiol. 2020;46(6):579–87. https://doi.org/10.1111/nan.12614.
Pinal-Fernandez I, Casal-Dominguez M, Derfoul A, Pak K, Plotz P, Miller FW, Milisenda JC, et al. Identification of distinctive interferon gene signatures in different types of myositis. Neurology. 2019;93(12):e1193–204. https://doi.org/10.1212/WNL.0000000000008128.
Rigolet M, Hou C, Baba Amer Y, Aouizerate J, Periou B, Gherardi RK, et al. Distinct interferon signatures stratify inflammatory and dysimmune myopathies. RMD Open. 2019;5(1):e000811. https://doi.org/10.1136/rmdopen-2018-000811.
Goronzy JJ, Weyand CM. Mechanisms underlying T cell ageing. Nat Rev Immunol. 2019;19(9):573–83. https://doi.org/10.1038/s41577-019-0180-1.
Naddaf E, Shelly S, Mandrekar J, Chamberlain AM, Hoffman EM, Ernste FC, Liewluck T. Survival and associated comorbidities in inclusion body myositis. Rheumatology (Oxford). 2022;61(5):2016–24. https://doi.org/10.1093/rheumatology/keab716.
Goyal NA, Coulis G, Duarte J, Farahat PK, Mannaa AH, Cauchii J, Irani T, et al. Immunophenoty** of inclusion body myositis blood T and NK cells. Neurology. 2022;98(13):e1374–83. https://doi.org/10.1212/WNL.0000000000200013.
Eloranta ML, Barbasso Helmers S, Ulfgren AK, Ronnblom L, Alm GV, Lundberg IE. A possible mechanism for endogenous activation of the type I interferon system in myositis patients with anti-Jo-1 or anti-Ro 52/anti-Ro 60 autoantibodies. Arthritis Rheum. 2007;56(9):3112–24. https://doi.org/10.1002/art.22860.
Jones EL, Laidlaw SM, Dustin LB. TRIM21/Ro52—roles in innate immunity and autoimmune disease. Front Immunol. 2021;12:738473. https://doi.org/10.3389/fimmu.2021.738473.
Pinal-Fernandez I, Casal-Dominguez M, Derfoul A, Pak K, Miller FW, Milisenda JC, Grau-Junyent JM. Machine learning algorithms reveal unique gene expression profiles in muscle biopsies from patients with different types of myositis. Ann Rheum Dis. 2020;79(9):1234–42. https://doi.org/10.1136/annrheumdis-2019-216599.
Cai Y, Zhu Y, Zheng J, Zhang Y, Chen W. NBR1 mediates autophagic degradation of IRF3 to negatively regulate type I interferon production. Biochem Biophys Res Commun. 2022;623:140–7. https://doi.org/10.1016/j.bbrc.2022.07.043.
D’Agostino C, Nogalska A, Cacciottolo M, Engel WK, Askanas V. Abnormalities of NBR1, a novel autophagy-associated protein, in muscle fibers of sporadic inclusion-body myositis. Acta Neuropathol. 2011;122(5):627–36. https://doi.org/10.1007/s00401-011-0874-3.
Yamashita S, Matsuo Y, Tawara N, Hara K, Yamamoto M, Nishikami T, Kawakami K, et al. CYLD dysregulation in pathogenesis of sporadic inclusion body myositis. Sci Rep. 2019;9(1):11606. https://doi.org/10.1038/s41598-019-48115-2.
Zhang Q, Jia Q, Gao W, Zhang W. The role of deubiquitinases in virus replication and host innate immune response. Front Microbiol. 2022;13:839624. https://doi.org/10.3389/fmicb.2022.839624.
Gasset-Rosa F, Lu S, Yu H, Chen C, Melamed Z, Guo L, Shorter J, et al. Cytoplasmic TDP-43 de-mixing independent of stress granules drives inhibition of nuclear import, loss of nuclear TDP-43, and cell death. Neuron. 2019;102(2):339–57. https://doi.org/10.1016/j.neuron.2019.02.038.
Adler BL, Christopher-Stine L. Triggers of inflammatory myopathy: insights into pathogenesis. Discov Med. 2018;25(136):75–83.
Singh H, Talapatra P, Arya S, Gupta V. Viral myositis as a close mimicker of polymyositis. Ann Trop Med Public Health. 2013;6:324–6. https://doi.org/10.4103/1755-6783.120997.
Manzano GS, Woods JK, Amato AA. Covid-19-associated myopathy caused by type I interferonopathy. N Engl J Med. 2020;383(24):2389–90. https://doi.org/10.1056/NEJMc2031085.
Uruha A, Noguchi S, Hayashi YK, Tsuburaya RS, Yonekawa T, Nonaka I, Nishino I. Hepatitis C virus infection in inclusion body myositis: a case-control study. Neurology. 2016;86(3):211–7. https://doi.org/10.1212/WNL.0000000000002291.
Eura N. Anti-cytosolic 5’-nucleotidase 1A (cN1A) positivity in muscle is helpful in the diagnosis of sporadic inclusion body myositis: a study of 35 Japanese patients. J Neurol Neurosci. 2016. https://doi.org/10.3390/biomedicines11071963.
Matsuura E, Umehara F, Nose H, Higuchi I, Matsuoka E, Izumi K, Kubota R, et al. Inclusion body myositis associated with human T-lymphotropic virus-type I infection: eleven patients from an endemic area in Japan. J Neuropathol Exp Neurol. 2008;67(1):41–9. https://doi.org/10.1097/nen.0b013e31815f38b7.
Ou SH, Wu F, Harrich D, García-Martínez LF, Gaynor RB. Cloning and characterization of a novel cellular protein, TDP-43, that binds to human immunodeficiency virus type 1 TAR DNA sequence motifs. J Virol. 1995;69(6):3584–96. https://doi.org/10.1128/JVI.69.6.3584-3596.1995.
Nehls J, Koppensteiner H, Brack-Werner R, Floss T, Schindler M. HIV-1 replication in human immune cells is independent of TAR DNA binding protein 43 (TDP-43) expression. PLoS ONE. 2014;9(8):e105478. https://doi.org/10.1371/journal.pone.0105478.
Zhou L, Miranda-Saksena M, Saksena NK. Viruses and neurodegeneration. Virol J. 2013;10:172. https://doi.org/10.1186/1743-422X-10-172.
Dalal F, Dalal H, McNew G. COVID-19-induced sporadic inclusion body myositis. Cureus. 2022;14(10):e30808. https://doi.org/10.7759/cureus.30808.
Saud A, Naveen R, Aggarwal R, Gupta L. COVID-19 and myositis: what we know so far. Curr Rheumatol Rep. 2021;23(8):63. https://doi.org/10.1007/s11926-021-01023-9.
Jacob S, Kapadia R, Soule T, Luo H, Schellenberg KL, Douville RN, Pfeffer G. Neuromuscular complications of SARS-CoV-2 and other viral infections. Front Neurol. 2022;13:914411. https://doi.org/10.3389/fneur.2022.914411.
Swartzman I, Gu JJ, Toner Z, Grover R, Suresh L, Ullman LE. Prevalence of myositis-specific autoantibodies and myositis-associated autoantibodies in COVID-19 patients: a pilot study and literature review. Cureus. 2022;14(9):e29752. https://doi.org/10.7759/cureus.29752.
Holzer MT, Krusche M, Ruffer N, Haberstock H, Stephan M, Huber TB, Kotter I. New-onset dermatomyositis following SARS-CoV-2 infection and vaccination: a case-based review. Rheumatol Int. 2022;42(12):2267–76. https://doi.org/10.1007/s00296-022-05176-3.
Huang C, Huang L, Wang Y, Li X, Ren L, Gu X, Kang L, et al. 6-month consequences of COVID-19 in patients discharged from hospital: a cohort study. Lancet. 2021;397(10270):220–32. https://doi.org/10.1016/S0140-6736(20)32656-8.
Su Y, Yuan D, Chen DG, Ng RH, Wang K, Choi J, Li S, et al. Multiple early factors anticipate post-acute COVID-19 sequelae. Cell. 2022;185(5):881–95. https://doi.org/10.1016/j.cell.2022.01.014.
Ramakrishnan RK, Kashour T, Hamid Q, Halwani R, Tleyjeh IM. Unraveling the mystery surrounding post-acute sequelae of COVID-19. Front Immunol. 2021;12:686029. https://doi.org/10.3389/fimmu.2021.686029.
Jiang R, Roy B, Wu Q, Mohanty S, Nowak RJ, Shaw AC, Kleinstein SH, O’Connor KC. The plasma cell infiltrate populating the muscle tissue of patients with inclusion body myositis features distinct B cell receptor repertoire properties. Immunohorizons. 2023;7(5):310–22. https://doi.org/10.4049/immunohorizons.2200078.
McLeish E, Sooda A, Slater N, Kachigunda B, Beer K, Paramalingam S, Lamont PJ, Chopra A, Mastaglia FL, Needham M, Coudert JD. Uncovering the significance of expanded CD8+ large granular lymphocytes in inclusion body myositis: Insights into T cell phenotype and functional alterations, and disease severity. Front Immunol. 2023;14:1153789. https://doi.org/10.3389/fimmu.2023.1153789.
Rose A, Isenalumhe L, Van den Bergh M, Sokol L. Clonal T-cell large granular lymphocytic disorders manifesting in patients with HIV-1 infection: case series and review of the literature. Mediterr J Hematol Infect Dis. 2018;10(1):e2018036. https://doi.org/10.4084/MJHID.2018.036.
Goronzy JJ, Li G, Yang Z, Weyand CM. The Janus head of T cell aging—autoimmunity and immunodeficiency. Front Immunol. 2013;4:131. https://doi.org/10.3389/fimmu.2013.00131.
Rothwell S, Cooper RG, Lundberg IE, Gregersen PK, Hanna MG, Machado PM, Herbert MK, et al. Immune-array analysis in sporadic inclusion body myositis reveals HLA-DRB1 amino acid heterogeneity across the myositis spectrum. Arthritis Rheumatol. 2017;69(5):1090–9. https://doi.org/10.1002/art.40045.
Rojana-Udomsart A, Bundell C, James I, Castley A, Martinez P, Christiansen F, Hollingsworth P, Mastaglia F. Frequency of autoantibodies and correlation with HLA-DRB1 genotype in sporadic inclusion body myositis (s-IBM): a population control study. J Neuroimmunol. 2012;249(1–2):66–70. https://doi.org/10.1016/j.jneuroim.2012.04.007.
Slater N, Sooda A, McLeish E, Beer K, Brusch A, Shakya R, Bundell C, James I, Chopra A, Mastaglia FL, Needham M, Coudert JD. High-resolution HLA genoty** in inclusion body myositis refines 8.1 ancestral haplotype association to DRB1*03:01:01 and highlights pathogenic role of arginine-74 of DRβ1 chain. J Autoimmun. 2024;142:103150. https://doi.org/10.1016/j.jaut.2023.103150.
Miller FW, Lamb JA, Schmidt J, Nagaraju K. Risk factors and disease mechanisms in myositis. Nat Rev Rheumatol. 2018;14(5):255–68. https://doi.org/10.1038/nrrheum.2018.48.
Ranasinghe S, Cutler S, Davis I, Lu R, Soghoian DZ, Qi Y, Sidney J, Kranias G, Flanders MD, Lindqvist M, Kuhl B, Alter G, Deeks SG, Walker BD, Gao X, Sette A, Carrington M, Streeck H. Association of HLA-DRB1-restricted CD4+ T cell responses with HIV immune control. Nat Med. 2013;19(7):930–3. https://doi.org/10.1038/nm.3229.
Barrett S, Ryan E, Crowe J. Association of the HLA-DRB1*01 allele with spontaneous viral clearance in an Irish cohort infected with hepatitis C virus via contaminated anti-D immunoglobulin. J Hepatol. 1999;30(6):979–83. https://doi.org/10.1016/s0168-8278(99)80249-9.
Bettencourt A, Carvalho C, Leal B, Brás S, Lopes D, Martins da Silva A, Santos E, et al. The protective role of HLA-DRB1(∗)13 in autoimmune diseases. J Immunol Res. 2015. https://doi.org/10.1155/2015/948723.
Ferre AL, Hunt PW, McConnell DH, Morris MM, Garcia JC, Pollard RB, Yee HF, et al. HIV controllers with HLA-DRB1*13 and HLA-DQB1*06 alleles have strong, polyfunctional mucosal CD4+ T-cell responses. J Virol. 2010;84(21):11020–9. https://doi.org/10.1128/JVI.00980-10.
Kummee P, Tangkijvanich P, Poovorawan Y, Hirankarn N. Association of HLA-DRB1*13 and TNF-alpha gene polymorphisms with clearance of chronic hepatitis B infection and risk of hepatocellular carcinoma in Thai population. J Viral Hepat. 2007;14(12):841–8. https://doi.org/10.1111/j.1365-2893.2007.00880.x.
James LM, Christova P, Lewis SM, Engdahl BE, Georgopoulos A, Georgopoulos AP. Protective effect of human leukocyte antigen (HLA) allele DRB1*13:02 on age-related brain gray matter volume reduction in healthy women. EBioMedicine. 2018;29:31–7. https://doi.org/10.1016/j.ebiom.2018.02.005.
Chikata T, Paes W, Kuse N, Partridge T, Gatanaga H, Zhang Y, Kuroki K, Maenaka K, Ternette N, Oka S, Borrow P, Takiguchi M. Impact of micropolymorphism outside the peptide binding groove in the clinically relevant allele HLA-C*14 on T cell responses in HIV-1 infection. J Virol. 2022;96(10):e0043222. https://doi.org/10.1128/jvi.00432-22.
Zeng L, Chen K, **ao F, Zhu CY, Bai JY, Tan S, Long L, et al. Potential common molecular mechanisms between Sjögren syndrome and inclusion body myositis: a bioinformatic analysis and in vivo validation. Front Immunol. 2023;14:1161476. https://doi.org/10.3389/fimmu.2023.1161476.
Lin A, Yan WH. The emerging roles of human leukocyte antigen-F in immune modulation and viral infection. Front Immunol. 2019;10:964. https://doi.org/10.3389/fimmu.2019.00964.
Zhang J, Khasanova E, Zhang L. Bioinformatics analysis of gene expression profiles of Inclusion body myositis. Scand J Immunol. 2020;91(6):e12887. https://doi.org/10.1111/sji.12887.
Jasinska AJ, Pandrea I, Apetrei C. CCR5 as a coreceptor for human immunodeficiency virus and simian immunodeficiency viruses: a prototypic love-hate affair. Front Immunol. 2022;13:835994. https://doi.org/10.3389/fimmu.2022.835994.
Ellwanger JH, Kulmann-Leal B, Kaminski VL, Rodrigues AG, Bragatte MAS, Chies JAB. Beyond HIV infection: neglected and varied impacts of CCR5 and CCR5Δ32 on viral diseases. Virus Res. 2020;286:198040. https://doi.org/10.1016/j.virusres.2020.198040.
Roos A, Preusse C, Hathazi D, Goebel HH, Stenzel W. Proteomic profiling unravels a key role of specific macrophage subtypes in sporadic inclusion body myositis. Front Immunol. 2019;10:1040. https://doi.org/10.3389/fimmu.2019.01040.
Trifone C, Salido J, Ruiz MJ, Leng L, Quiroga MF, Salomón H, Bucala R, et al. Interaction between macrophage migration inhibitory factor and CD74 in human immunodeficiency virus type I infected primary monocyte-derived macrophages triggers the production of proinflammatory mediators and enhances infection of unactivated CD4+ T cells. Front Immunol. 2018;9:1494. https://doi.org/10.3389/fimmu.2018.01494.
Connolly CM, Plomp L, Paik JJ, Allenbach Y. Possible future avenues for myositis therapeutics: DM, IMNM and IBM. Best Pract Res Clin Rheumatol. 2022;36(2):101762. https://doi.org/10.1016/j.berh.2022.101762.
Astorga-Gamaza A, Perea D, Sanchez-Gaona N, Calvet-Mirabent M, Gallego-Cortés A, Grau-Expósito J, Sanchez-Cerrillo I, Rey J, Castellví J, Curran A, Burgos J, Navarro J, Suanzes P, Falcó V, Genescà M, Martín-Gayo E, Buzon MJ. KLRG1 expression on natural killer cells is associated with HIV persistence, and its targeting promotes the reduction of the viral reservoir. Cell Rep Med. 2023;4(10):101202. https://doi.org/10.1016/j.xcrm.2023.101202.
Yu CH, Davidson S, Harapas CR, Hilton JB, Mlodzianoski MJ, Laohamonthonkul P, Louis C, et al. TDP-43 triggers mitochondrial DNA release via mPTP to activate cGAS/STING in ALS. Cell. 2020;183(3):636–49. https://doi.org/10.1016/j.cell.2020.09.020.
Mori F, Tada M, Kon T, Miki Y, Tanji K, Kurotaki H, Tomiyama M, et al. Phosphorylated TDP-43 aggregates in skeletal and cardiac muscle are a marker of myogenic degeneration in amyotrophic lateral sclerosis and various conditions. Acta Neuropathol Commun. 2019;7(1):165. https://doi.org/10.1186/s40478-019-0824-1.
Damian L, Login CC, Solomon C, Belizna C, Encica S, Urian L, Jurcut C, et al. Inclusion body myositis and neoplasia: a narrative review. Int J Mol Sci. 2022;23(13):7358. https://doi.org/10.3390/ijms23137358.
Pozzi S, Codron P, Soucy G, Renaud L, Cordeau PJ, Dutta K, Bareil C, Julien JP. Monoclonal full-length antibody against TAR DNA binding protein 43 reduces related proteinopathy in neurons. JCI Insight. 2020;5(21):e140420. https://doi.org/10.1172/jci.insight.140420.
Wang P, Deng J, Dong J, Liu J, Bigio EH, Mesulam M, Wang T, et al. TDP-43 induces mitochondrial damage and activates the mitochondrial unfolded protein response. PLoS Genet. 2019;15(5):e1007947. https://doi.org/10.1371/journal.pgen.1007947.
Njomen E, Tepe JJ. Proteasome activation as a new therapeutic approach to target proteotoxic disorders. J Med Chem. 2019;62(14):6469–81. https://doi.org/10.1021/acs.jmedchem.9b00101.
Moghadam-Kia S, Oddis CV. Current and new targets for treating myositis. Curr Opin Pharmacol. 2022;65:102257. https://doi.org/10.1016/j.coph.2022.102257.
Guglielmi V, Cheli M, Tonin P, Vattemi G. Sporadic inclusion body myositis at the crossroads between muscle degeneration, inflammation, and aging. Int J Mol Sci. 2024;25(5):2742. https://doi.org/10.3390/ijms25052742.
Terracciano C, Nogalska A, Engel WK, Askanas V. In AbetaPP-overexpressing cultured human muscle fibers proteasome inhibition enhances phosphorylation of AbetaPP751 and GSK3beta activation: effects mitigated by lithium and apparently relevant to sporadic inclusion-body myositis. J Neurochem. 2010;112(2):389–96.
Hong N, Park JS, Kim HJ. Synapto-protective effect of lithium on HIV-1 Tat-induced synapse loss in rat hippocampal cultures. Anim Cells Syst (Seoul). 2021;26(1):1–9. https://doi.org/10.1080/19768354.2021.2018044.
Schifitto G, Zhong J, Gill D, Peterson DR, Gaugh MD, Zhu T, Tivarus M, Cruttenden K, Maggirwar SB, Gendelman HE, Dewhurst S, Gelbard HA. Lithium therapy for human immunodeficiency virus type 1-associated neurocognitive impairment. J Neurovirol. 2009;15(2):176–86. https://doi.org/10.1080/13550280902758973.
Acknowledgements
Not applicable.
Funding
Partial support for open access publication has been received from Foundation Vital Team.
Author information
Authors and Affiliations
Contributions
Conceptualization was presented by LD; documentation was collected by VV, AC, and RV; imaging was provided by RV and LD; writing—original draft preparation was revised by LD; writing—review and editing were prepared by RV, AC, and VV. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Văcăraş, V., Vulturar, R., Chiş, A. et al. Inclusion body myositis, viral infections, and TDP-43: a narrative review. Clin Exp Med 24, 91 (2024). https://doi.org/10.1007/s10238-024-01353-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10238-024-01353-9