Abstract
Background
Staphylococcus arlettae is a rarely reported coagulase-negative staphylococcus (CoNS) isolated from infected humans and livestock. Observing phage-bacteria interaction could improve the understanding of bacterial pathogenetic mechanisms, providing foundational evidence for phage therapy or phage detection. Herein, we aimed to characterise and annotate a novel bacteriophage, vB_SarS_BM31 (BM31), specific to S. arlettae. This bacteriophage was isolated from a milk sample associated with bovine mastitis and collected in the Sichuan Province, China.
Results
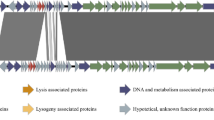
The BM31 genome comprised a linear double-stranded DNA of 42,271 base pair in length with a G + C content of 34.59%. A total of 65 open reading frames (ORFs) were assembled from phage DNA, of which 29 were functionally annotated. These functional genes were divided into four modules: the structural, DNA packing and replication, lysis, and lysogeny modules. Holin (ORF25), lysin (ORF26), and integrase (ORF28) were located closely in the entire BM31 genome and were important for lyse or lysogeny cycle of BM31. The phage was identified as a temperate phage according to whole genome analysis and life cycle assay, with basic biological characteristics such as small burst size, short latency period, and narrow host range, consistent with the characteristics of the family Siphoviridae, subcluster B14 of the Staphylococcus bacteriophage.
Conclusions
The present isolation and characterisation of BM31 contributes to the Staphylococcus bacteriophage database and provides a theoretical foundation for its potential applications. To the best of our knowledge, BM31 is the only shared and completely reported phage against S. arlettae in the entire public database.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Staphylococci can directly or indirectly cause arthritis and comb necrosis in chickens, mastitis in dairy cattle, extraocular infections in horses, ovine staphylococcal dermatitis in sheep, and exudative epidermitis in pigs (Oduor et al. 2020). In clinical diagnosis, staphylococci are divided into two groups based on their capacity for coagulase production: coagulase-positive (CoPS) and coagulase-negative (CoNS) staphylococci (Hatoum-Aslan 2021). Staphylococcus aureus and Staphylococcus epidermidis have been studied extensively as typical CoPS and CoNS strains, respectively, while other types of CoNS strains remain poorly understood. S. arlettae is commonly regarded as a commensal species; however, it is associated with different types of infection in contexts of large-scale antibiotic use (Lavecchia et al. 2019). S. arlettae strains have been previously isolated from patients afflicted with human chronic prostatitis, otologic infection, rheumatic mitral stenosis, bovine mastitis, dairy goat intramammary infection, and pig exudative epidermidis (Bernier Gosselin et al. 2019; Dinakaran et al. 2012; Hou et al. 2000; Park et al. 2013; ** giant. Emerg Top Life Sci 1(1):93–103. https://doi.org/10.1042/etls20170002 " href="/article/10.1007/s10123-022-00292-3#ref-CR72" id="ref-link-section-d93575127e722">2017). Consistent study of temperate phages may be beneficial to assess the efficiency and safety of introducing phage-derived proteins, e.g. endolysin, or introducing genetic mutations that convert the phage to a virulent type, increasing phage lytic ability, and making it suitable for use in phage cocktails or in combination with antibiotics (Al-Anany et al. 2021; Dedrick et al. 2019; Hargreaves and Clokie 2014). Phages are also used as diagnostic tools, in food production, and in biotechnology (Dunne et al. 2018). For example, they can regulate the intestinal flora of livestock, prevent contamination of ready-meals by foodborne bacteria (Salmonella, Listeria), and may be used as bio-preservatives (Guenther et al. 2012; Guenther et al. 2009; Kazi and Annapure 2016; Kim et al. 2017). Phages and their derived proteins can be engineered to detect S. aureus, Salmonella, Campylobacter jejuni, and Shigella flexneri (Bhardwaj et al. 2017; Kittler et al. 2013; Lakshmanan et al. 2007; Singh et al. 2013).
Based on these potential applications, novel phages specific to different host bacteria are needed. In response to an appeal of the United States Food and Drug Administration, phages are not used in clinical treatment if any of their genes encode toxins or other proteins that may enhance bacterial virulence (Abedon et al. 2011). Therefore, all isolated phages must be fully characterised through complete genome sequencing and functional analysis before application. Detailed phage information is required when bacteriophage-derived proteins are used to detect or remove pathogenic bacteria in diagnostic or therapeutic products, or food (Schmelcher and Loessner 2014). This study aimed to characterise a temperate phage vB_SarS_BM31, which is first reported to against S. arlettae.
Materials and methods
Bacterial strains and growth conditions
The host strain S. arlettae was isolated from a milk sample obtained at a dairy farm in Sichuan, China. Other Staphylococcus spp. used in this study were also isolated from milk samples obtained from animals with clinical or subclinical mastitis and stored at − 80 °C with 50% glycerol in the laboratory. Partial bacterial 16S rRNA polymerase chain reaction (PCR) analysis was used to verify the strains (Table 1). All strains were grown in Luria–Bertani (LB) broth (pH 7.0 ± 0.1) at 37 °C.
Phage isolation, purification, and amplification
Phage BM31 was isolated from a milk sample (obtained from the Sichuan Province) contaminated with S. arlettae using the double-layer agar method described previously, with slight modifications. Briefly, 1 mL of milk sample was added to 5 mL of LB broth suspended with 500 μL of exponentially growing culture of S. arlettae, and incubated at 37 °C for 4 h and shaken at 225 rpm. Flocculent precipitates in milk were removed by centrifugation at 4000 rpm for 10 min at room temperature and filtered through a 0.45-μm syringe filter. Subsequently, 200 μL of exponentially growing culture of S. arlettae and 2 mL of filtrate were mixed with 2 mL of LB broth, incubated at 37 °C for 24 h and shaken. Cell debris was removed by centrifugation at 8000 rpm for 5 min at 4 °C and filtered through a 0.22-μm syringe filter. The suspected phage sample was obtained and stored at 4 °C in a dark room.
A mixture of 200 μL of S. arlettae and 200 μL of suspected phage sample, which were adsorbed over 25 min at room temperature, was mixed into 3 mL of warm LB soft agar (0.6% w/v, pH 7.0 ± 0.1) and poured onto LB agar plates (1.2% w/v), maintained stable for 30 min, and then incubated at 37 °C overnight. The presence of phages was confirmed by visualising the presence of clear or turbid plaques. The clear plaque was picked out using a sterile Pasteur pipette and dissolved into a chloride-magnesium sulphate (SM) buffer (5.8 g/L of NaCl, 2.0 g/L of MgSO4, 50 mL/L of 1 M Tris, pH 7.5, and 5 mL/L of pre-sterilised 2% gelatine) in a 1.5-mL centrifugation tube, shaking for 2 h at 37 °C or stably standing overnight at 4 °C (Anand et al. 2015). The phage was tenfold serially diluted, and the double-layer agar method was performed to obtain new phage plaques. The process was repeated four or more times until the plaques were uniform.
When the plaque morphology was uniform, 5 mL of SM buffer was added to the plate with gentle shaking for 4 h at 37 °C. The liquid on the top of the plate was centrifuged at 5500 rcf for 5 min and filtered through a 0.22-μm syringe filter to obtain a high-titre phage.
Transmission electron microscopy
One microlitre of purified phage BM31, obtained using the method described above, was placed on top of a carbon-coated copper grid and stained with 2% phosphotungstic acid (pH = 7.0). The stained phage was observed using a transmission electron microscope (Lanzhou Veterinary Research Institute, China) at 80 kV 30,000 × magnification (Kyoung Min et al. 2018).
Chloroform and ether sensitivity
Phage sensitivity to chloroform and ether was used to determine whether a lipid substance was present in the capsid or tail fibre of phage BM31 (Espejo and Canelo 1968; Lu et al. 2017; Wei et al. 2020). For this process, 100 μL of phage (~ 108 plaque forming units [PFU]/mL) in SM buffer was mixed with 900 μL of chloroform and ether, shaken vigorously for 1 min, and incubated at room temperature for 1 h. The treated samples were immediately diluted in SM buffer and plated for phage titration using double-layer agar plates inoculated with S. arlettae; the original sample was used as a control group. The assay was performed in triplicates.
Thermal and pH stability
To evaluate the thermal stability of the phage, 1 mL of phage with SM buffer was incubated at 40, 50, 60, 70, and 80 °C. Samples were removed every 20 min for 1 h and phage titres were immediately determined at each time interval. To investigate the pH stability of the phage, 100 μL of phage was mixed with 900 μL of SM buffer, which was adjusted to pH 3, 5, 9, and 11 using HCl or NaOH, respectively. Phages in SM buffer at pH 7 served as controls. The phages in solutions of various pH values were incubated for 1 h at room temperature, following which, the phage titres were determined. The assays were performed in triplicate.
Multiplicity of infection assay
To achieve the highest efficacy of the phage at the lowest cost and set a standard for further study, a multiplicity of infection (MOI) assay was established to determine the optimal ratio of PFU/colony forming units (CFU) (Feng et al. 2021). Briefly, the phage was diluted to 1 × 105–12 PFU/mL in an SM buffer and incubated with S. arlettae at a concentration of 1 × 109. The double-layer agar method was used to determine phage titration, and the assay was performed in triplicate.
One-step growth curve assay
The host strain S. arlettae (1 × 107 CFU/mL) and phage BM31 (1 × 104 PFU/mL) were first adsorbed at an MOI of 0.001 for 15 min at 37 °C, and subsequently centrifuged at 12,000 rcf for 1 min. The supernatant, which contained free phage, was discarded and washed twice in sterilised LB broth. The pellets that consisted of a bacteria-phage complex were resuspended using warm LB broth and immediately incubated at 37 °C and shaken for 150 min. The samples were removed and immediately stored at 4 °C for simultaneous phage titration at 0 min, at 5-min intervals within 30 min, 10-min intervals within the next 1 h, and 30-min intervals within the next 1 h. This assay was repeated three times. The latency period and burst size were calculated, representing the time required from phage invasion to progeny maturation and cell lysis, and the number of phage particles released from the bacterial cell, respectively (Madurantakam Royam and Nachimuthu 2020). The burst size was calculated using the formula: median number of plaques at tend of the replication cycle/tstart of the replication cycle (Eckstein et al. 2021).
Host range
In this study, 1 × 109 PFU/mL of phage BM31 was prepared for the host range assay for all 13 Staphylococcus spp. strains (all isolated from bovine milk, Sichuan Province of China) involved in the host strain S. arlettae. Briefly, 100 μL of each strain was coated on LB agar plates and dried at room temperature for a few minutes until there were no water drops. Subsequently, 10 μL of BM31 and 10 μL of sterilised saline, which served as a control, were dropped on the coating plates and incubated overnight to observe whether there were any lytic plaques on each plate.
Identification and extraction of nucleic acid
Following the manufacturer’s instructions, DNAse I, RNAse A, and mung bean nuclease (Takara Biomedical Technology Co., Ltd., Bei**g, China) were used to confirm the type of nucleic acid in the phage (Kumar et al. 2021).
Phage DNA was obtained from the purified phage suspension using a viral RNA/DNA extraction kit (Takara Biomedical Technology Co., Ltd., Bei**g, China) according to the manufacturer’s instructions.
Genome assembly and annotation
Phage genomic DNA was sequenced by Personalbio (Personalbio Technology Co., Ltd., Shanghai) using next-generation sequencing on an Illumina NovaSeq platform. Quality control analysis was performed using FastQC v0.11.7. A5-MiSeq v20160825 and SPAdes v3.12.0 were used for genome sequence assembly.
ORFs were predicted using the GeneMarkS v4.32 (Besemer et al. 2001). Homology searching was performed with BLASTp against the NCBI nonredundant database, and sequences with an E-value of < 10−5 were considered homologues. HHpred against the Protein Data Bank and Pfam databases were used to predict more distant homologues (Söding et al. 2005). The genome was analysed for tRNA genes using the tRNAscan-SE 2.0 (Lowe and Chan 2016). The Comprehensive Antibiotic Resistance Database (https://card.mcmaster.ca/) and virulence factors of pathogenic bacteria (http://www.mgc.ac.cn/VFs/main.htm) were used to detect the antibiotic resistance and virulence genes, respectively. PHASTER (http://phaster.ca/) was used to identify the most closely related phages and the presence of prophages (Dakheel et al. 2019). The replication method and life cycle of BM31 were predicted using the phage classification prediction program (http://www.phantome.org/PHACTS/index.php) (Lu et al. 2017). A genomic map was generated using CGView (CGView server) (Grant and Stothard 2008).
Life cycle
The lifecycle of BM31 was determined based on whether the specific structural proteins harboured in phage BM31 were present in S. arlettae (host strain) before and after lysis with BM31 (Lu et al. 2017). Briefly, 5 mL of LB broth was added to the double agar plate lawned with S. arlettae, which was lysed by BM31 and rocked gently at room temperature for 2 h. The broth at the top of the agar was collected in microcentrifuge tubes and centrifuged at 5500 rcf for 5 min. The supernatant was discarded, and 1 mL of fresh LB broth was added to resuspend the pellets, followed by centrifugation. The contaminating phage was removed by repeating the washing step thrice. After the final centrifugation step and removal of the supernatant, the bacteria pellets and the original isolated S. arlettae, which was stored at − 80 °C, were both used to streak for isolation on LB agar plates and incubated overnight at 37 °C. Single colonies were randomly collected for PCR using primers specific to BM31 (Table 1) designed according to phage genome annotation, targeting ORF4 (portal protein), ORF6 (capsid protein), and ORF15 (tape measure protein). All these primers were generated using SnapGene (version 4.3.6). Visualisation of specific bands under transilluminator suggests that BM31 can integrate its own genome into the chromosome of host bacteria, as this ability belongs to lysogeny phages. In contrast, it suggests that the phage possesses a lytic life cycle.
Taxonomic analysis
The whole genome and major capsid protein sequences in phage BM31 were compared phylogenetically with those in other similar phages (compared using BLASTN) extracted from the GenBank database (http://www.ncbi.nih.gov/) using MEGA 7 software (version 7.0.26). ClustalW was used to align the inferred amino acid sequences using default parameters. Based on multiple sequence alignment, the Tamura-Nei model was selected, and a maximum likelihood tree was constructed with 1000 bootstrap replicates.
ANI values between BM31 and other Staphylococcus phages were generated using the ANI Calculator (ANI Calculator | Ezbiocloud.net). A heat map was created using TBtools v1. 0,986,853. Genome comparison between BM31 and its closest similar phage was performed using Geneious Prime (Mauve Plugin version 1.1.3).
Statistical analysis
Data were collected and analysed using GraphPad Prism version 7 software. The t-test was used to analyse the differences between two groups, and ANOVA was used for more than two groups. Error bars in the figures represent the standard deviation of the mean of three replicate experiments. Differences were considered statistically significant at p values of < 0.05.
Results and discussion
Phage morphology
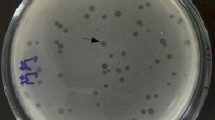
The isolated phage formed large and clear plaques of 1.98 ± 0.02 mm diameter with turbid halos after overnight incubation on its indicator host (Fig. 1a). Such halos may indicate the presence of a phage-encoded depolymerase that could eliminate exopolysaccharides in the bacterial cell wall (Chmielewska-Jeznach et al. 2020).
Transmission electron microscopy (TEM) classified BM31 as a Siphoviridae phage with a B1 morphotype based on an isometric head averaging 63 ± 7 nm in diameter and a long, non-contractile tail with an average tail length and width of 168 ± 7 nm and 14 ± 7, respectively (Ackermann and Eisenstark 1974). A baseplate was observed at the end of the tail (Fig. 1b).
Basic characteristics
No phage particles survived in chloroform or ether, whereas the phage titre in the control group (SM buffer) remained approximately 2 × 108 PFU/mL (Fig. 2a). This result suggests that phage BM31 is susceptible to both chloroform and ether, implying the presence of lipids in the capsid or a surrounding lipid layer in the phage (Wang and Li 2018). Therefore, chloroform or ether should not be used in further studies on BM31.
In the pH stability assay, BM31 showed the highest titre at an average of 2 × 107 PFU/mL at pH 7, surviving incubation at pH 9 for 1 h with an average titre of 1.4 × 107 PFU/mL (70% survival rate); no phage particle survived in other pH groups (Fig. 2b). This finding indicates that BM31 could stand a weakly alkaline environment, but the optimal survival environment may be neutral.
The thermal stability assay indicated that the tolerable temperature for BM31 was 40–60 °C (Fig. 2c), and this phage demonstrated good thermal stability. The results showed that after 60 min of incubation at 40 °C, the phage titre was 1.07 × 1012 (14.50% survivability) when the original titre was 7.40 × 1012. After 60 min of incubation at 50 °C and 60 °C, the phage titres were determined to be 1.70 × 1011 (2.29% survivability) and 2.32 × 1010 (0.31% survivability), respectively. BM31 could not survive at a temperature of 70 °C for more than 20 min, and at 80 °C, the survival duration was less than 20 min.
The results of the MOI assay showed that BM31 exhibited the best capacity for host lysis and self-replication when the MOI value reached 0.001. The optimal MOI value was associated with relatively low costs and yielded a phage titre significantly higher than that of other groups (Fig. 2d). Specifically, the viability of BM31 is stable, which is related to its structure since the Siphoviridae family is reportedly the most stable phage (Wintachai et al. 2020).
One-step growth curve
For phage BM31, the latency period was 20 min with a burst size of 49 phage particles per infected cell. Compared to other phages, BM31 had a shorter latency period and smaller burst size. The burst size is correlated with the rates of synthesis and assembly of phage components, latency period, metabolic activity, living environment, and protein synthesis machinery of host bacteria, rather than the cell size or DNA composition of the phage (Pan et al. 2021). The one-step curve for BM31 is represented in Fig. 3.
Host range
In all 13 staphylococcal strains tested for host range (Table 2), BM31 could only lyse the host strain of S. arlettae, suggesting that it is specific to its host with an extremely narrow host range.
Genomic overview
Treatment of the nucleic acid in BM31 with DNAse I resulted in digestion of its genome, whereas treatment with RNAse A and mung bean nuclease showed no sensitivity, thereby revealing that BM31 is a dsDNA virus (Fig. 4). The complete dsDNA genome of phage BM31 was 42,271 bp in size, with 34.59% of G + C (GenBank accession no. MZ488273). Functional prediction of the 65 ORFs using BLASTp analysis produced significant matches for 53 proteins. Of the 53 proteins with hits, only 26 were assigned to putative functions (Table S1). The remaining 12 proteins could not be matched with any of the proteins in the National Centre for Biotechnology Information (NCBI) database and were annotated as hypothetical. ORFs with distant homology to BM31, identified via HHpred analysis, are listed in Table 3. Although these hypothetical proteins were distributed throughout the BM31 genome, a clear modular organisation was evident, consisting of genes involved in virion morphogenesis (blue), DNA packing and replication (green), and lysis (purple) (Fig. 5). The most known functional genes were on the positive strand, except the integrase (ORF28) and transcriptional regulator (ORF32), which were in the middle of the gene cycle. No tRNA genes, antibiotic-resistance genes, or prophages were detected. However, a virulence factor named pemK and mazF-like toxin (ORF30, pfam02452), which is an endoribonuclease toxin of the type II toxin-antitoxin system, were observed (Oliveira et al. 2019).
Nucleic acid identification of BM31. M, 1-kb DNA ladder (COOLABER SCIENCE & TECHNOLOGY Co., LTD., Bei**g, China); 1, BM31 treated with mung bean nuclease; 2, BM31 treated with DNAse I; 3, BM31 treated with RNAse A; N, original BM31 genome as control. Original gel is presented in Supplementary Fig. 1
Structural module
According to the function of annotated genes, the whole genome of BM31 was divided into four modules: structural, lysis, lysogeny, and DNA packing and replication genes. Proteins involved in capsid assembly and packing included the terminase small subunit (TerS) (ORF1), terminase large subunit (TerL) (ORF2), portal protein (ORF4), prohead protease (ORF5), and capsid protein (ORF6). The small and large subunits of phage terminase are key enzymes involved in DNA translocation and head filling (Mitchell and Rao 2006). The prohead protease is observed in some bacteria, likely as a result of horizontal transfer. The portal protein is a crucial element of the packing motor, which pumps the phage genome into the capsid with the assistance of a large subunit terminase (ATPase activity) (Kyrkou et al. 2020). Seven proteins were identified as structural proteins involved in tail morphogenesis and phage assembly: a head–tail adaptor (ORF9), head–tail joining protein (ORF10), major tail protein (ORF12), tail length tape measure protein (ORF15), tail fibre protein (ORF16, ORF17), and minor structural protein (ORF19). ORF15 was the longest sequence in the complete genome of BM31 and was annotated as a tail length tape measure protein (TMP), a tail-associated protein. TMP is not only present in tailed phages, but also in tail-less phages, determining the tail length and DNA transition into the host cell during infection (Gao et al. 2012; Mahony et al. 2016; Zhang et al. 2020). For most phages, tail fibres are the first proteins to recognise receptors on the bacterial membrane and initiate an infection (Geng et al. 2020). The location of capsid assembly genes upstream of the tail assembly genes and the presence of the longest TMP revealed that this structural module corresponds to the typical Siphoviridae morphogenesis module (Hatfull 2008).
Lysis module
The lytic cassette is composed of holin (ORF25) and lysin (ORF26). The lysin (ORF26) component consisted of an N-terminal CHAP endopeptidase domain (pfam 05,257, amino acids 27–115) and an N-acetylmuramoyl-L-alanine-amidase domain (MurNAc-LAA, cd02696, amino acids 173–355), which was the same as the domain components of the lysin in phage Ph28 (Muharram et al. 2020). In addition, the lysin component of BM31 contained a SH3 peptidoglycan-binding domain (amino acids 377–438) in the C-terminus, which confirms that modularity is prevalent in staphylococcal lysins (Oliveira et al. 2013; Pertics et al. 2020). The holin (ORF25) component belongs to the SPP1 family (PF04688), which corresponds to 90.59% similarity to an uncultured Caudovirales phage from South Africa (van Zyl et al. 2018). Consecutive use of lysin and holin could lead most tailed phages to lyse and control the length of the infection cycle (Lu et al. 2017).
Lysogeny module
The lysogeny cassette contains integrase (ORF28) but lacks a CI-type repressor, which is normally present in other staphylococcal phages (García et al. 2009). However, the lambda repressor-like, Cro/C1-type helix-turn-helix (HTH) domain (ORF33), which is a DNA-binding domain and transcriptional regulator, was predicted downstream of the XRE family transcriptional regulator (ORF32) (Hargreaves et al. 2014). The HTH domain may indicate the presence of a repressor that cannot be annotated beyond unknown function proteins or hypothetical proteins in the BM31 complete genome. The phage_integrase domain (PF00589.24) of BM31 belongs to pfam00589, indicating that the integrase is a tyrosine recombinase (Y-Int), which possesses tyrosine as a catalytic residue (Oliveira et al. 2019). Preliminary evidence suggests that BM31 possesses two lyse systems and tends to obey the lysogenic cycle. Exception may be conditions that involve cellular stress, wherein anti-repressor proteins in the phage interfere with the function of the repressor protein and force the phage to adopt the lytic cycle in a process called a lytic-lysogenic switch (Das et al. 2020). The transcriptional activator RinB (ORF62) is associated with integrative recombination of phages. For the staphylococcal phage phi11, rinA and rinB are required for active expression of the int gene, which is the sole viral gene responsible for integrative recombination.
DNA packing and replication module
The DNA packing and replication module consists of a single-stranded DNA-binding protein (ORF42), a replication initiation protein (ORF44), helicase DnaB (ORF46, ORF48), and others. All known temperate staphylococcal phages belong to the family Siphoviridae, suggesting that phage BM31 may be classified as a lysogenic phage (Ingmer et al. 2019); specifically, when integrase (ORF28), pathogenicity island protein (ORF34), and dUTPase (ORF60) are annotated in BM31. The pathogenicity island may be a type of phage-related chromosomal island, such as S. aureus pathogenicity islands (SaPIs), which are highly mobile and superantigen-encoding genetic elements closely associated with temperate phages (Novick et al. 2010). SaPIs carry genes encoding the toxic shock syndrome toxin, staphylococcal enterotoxin B, and other important virulence factors (Lindsay et al. 1998; Moller et al. 2019; Novick et al. 2010). SaPIs reside stably in prophages and can only achieve transduction by hijacking capsids of helper phages, rather than by themselves. The stl gene inside SaPI is a master repressor and can be depressed by dUTPase, which is expressed by the helper phage, so that the SaPI lytic cycle is activated. SaPI DNA was packed into these phage capsids after TerS recognised and cleaved the SaPI pac sites, resulting in the release of mature progeny phages (decreased in number), and SaPI particles (with smaller capsids than progeny phages) were released after bacterial lysis (Moller et al. 2019; Novick et al. 2010). HNH homing endonuclease (ORF40, ORF47, ORF65) genes are highly specialised selfish genetic components commonly present in phage genomes (Kyrkou et al. 2019, 2020). The HNH endonuclease may facilitate its own and relevant genes’ mobility, even excluding other competing phages by cleaving their DNA (Goodrich-Blair and Shub 1996; Mohamed et al. 2012). It may also interfere with the expression of genes surrounding its insertion position and disrupt the active ORFs (Pan et al. 2021).
Life cycle
All phage proteins used in this study (capsid protein, portal protein, and tape measure protein) were detected in S. arlettae colonies which were randomly selected, irrespective of whether the strain was invaded by BM31 (Fig. 6). The results suggest that BM31 has a lysogeny-like life cycle.
Electrophoresis map showing specific structural protein of phage BM31 in S. arlettae. Lane M, DL 2000 DNA marker; lane N, negative control: PCR product without bacterial template. a Lanes 1–16, random clone of S. arlettae using primer CP, and original gel is presented in Supplementary Fig. 2. b Lanes 1–16, random clone of S. arlettae revealed using primer CP1, and original gel is presented in Supplementary Fig. 3. c Lanes 1–6, 7–12, and 13–18, random clone of S. arlettae revealed using primers CP675, TMP81, and PP488, respectively; and original gel is presented in Supplementary Fig. 4
All randomly selected colonies had several phage genes encoding phage structural proteins, both before and after lysis by phage BM31. Phage genes detected in the host strain genes may be the result of their ability to integrate its own genome into the host chromosome and achieve self-replication by bacterial replication, suggesting that the examined phage is a temperate phage. The phage genes were also observed in the colony that was never treated with BM31, likely because both S. arlettae and BM31 were isolated from one milk sample. As we first isolated the host strain, it is likely that the phage had already undergone the lysogenic cycle and integrated into the host strain chromosome. Findings from both genome annotation and the PHACTS online software analyses suggested that BM31 may be a temperate phage with a pac site.
Taxonomic analysis
BM31 showed the closest sequence identity to phage phiRS7 (KF589919), with 36% query coverage and 83.82% nucleotide identity. To investigate the relationship between phage BM31 and other related phages stored in GenBank, two maximum likelihood phylogenetic trees were constructed based on the amino acid sequences of the phage capsid protein (ORF6) and complete genome sequences. Both trees revealed a close relationship between BM31 and Staphylococcus phage phiRS7, which belongs to the family Siphoviridae (Fig. 7). This result is consistent with that of the PHASTER online analysis software. Combined with the morphology characteristics determined by TEM, phage BM31 was defined as a member of family Siphoviridae.
Phylogenetic tree of BM31 and other related phages obtained from GenBank. a Phylogenetic tree based on capsid protein; b phylogenetic tree based on complete genome. Black inverted triangle, phage BM31 in this study. The number on the branch point represents credibility (values closer to 100 represent stronger credibility). The length of the branches represents the genetic distance; the shorter the ruler, the closer the relationship
All 205 Staphylococcus phages can be divided into four clusters (A, B, C, and D) (Oliveira et al. 2019). Following this standard, we aimed to classify phage BM31 into one of them. Three conserved pfams of BM31 were annotated; pfam1520 (ORF26) was the same as subcluster C3; pfam692 (ORF60) and pfam1844 (ORF65) were the same as subcluster D1, also presenting with temperate behaviour and long tail morphology, which were specific to cluster B. Therefore, 132 phages of cluster B, 5 phages of subcluster C3, and 2 phages of subcluster D1 were compared to BM31 through average nucleotide identity (ANI) analysis (Supplementary Dataset File). The results showed that BM31 contained the highest ANI of 79.83% with phiRS7 (KF589919); therefore, we classified BM31 into the same subcluster as phiRS7 in B14, which is a subcluster under cluster B (Fig. 8).
To obtain a more nuanced understanding of the genomic structural differences between BM31 and other phages, BM31 was comparatively analysed with the genome of the most homologous phiRS7 phage (Fig. 9). The results showed high degree of similarity and conservation between the genomes of the two phages, especially in the structural genes. Insertion and deletion variations in parts of the genes are mainly reflected in integrase, helicase, and HNH endonuclease. The results also prove the possible existence of a RinB gene in the BM31 genome that was originally annotated as a hypothetical protein (ORF62).
While we aim to use BM31 as a therapy/detection application, one problem is the virulence of the phage itself. The presence of the lysogenic cycle and pemK and mazF-like toxin may determine the rare possibility of phage BM31 being an antibiotic agent. Under these circumstances, whether the endolysin protein derived from BM31 may be a safe sole agent or used in coordination with antibiotics requires further study. Research has shown that the phage particle, the endolysin derived from the phage, and the cell wall binding domain derived from endolysin are all valid detection agents. The extremely narrow host range for BM31 indicates the high specificity of the phage acting as a detector. To achieve simultaneous multiple bacterial detection, the phage requires a wide host spectrum. Whether the expression of endolysin protein will broaden the host range also needs to be verified in subsequent experiments.
Conclusion
BM31 is a novel and rare phage that is highly specific to S. arlettae. The present findings supplement the phage database of CoNS, elucidating the mechanisms underlying bacteria-phage interactions, which may translate into clinical applications. BM31 survived in both normal and stable environments and exhibited a small burst size and short latency period. The small burst size may be associated with its lysogenic life cycle, and shorter latencies may suggest a rapid and high efficacy when the phage is applied. Although S. arlettae is not a major pathogenic bacterium, it was isolated from a bovine mastitis milk sample in this study, and it may contaminate dairy products. BM31 could not be used as a therapeutic agent due to lysogenic replication; however, phage-derived proteins such as lysin may be used to detect and control S. arlettae in further studies.
Data availability
All data generated or analysed during this study are included in this article and supplementary information files. The complete genome sequence of the novel phage BM31 has been deposited in GenBank under accession number MZ488273.
References
Abedon ST, Kuhl SJ, Blasdel BG, Kutter EM (2011) Phage treatment of human infections. Bacteriophage 1(2):66–85. https://doi.org/10.4161/bact.1.2.15845
Ackermann HW, Eisenstark A (1974) The present state of phage taxonomy. Intervirology 3(4):201–219. https://doi.org/10.1159/000149758
Al-Anany AM, Fatima R, Hynes AP (2021) Temperate phage-antibiotic synergy eradicates bacteria through depletion of lysogens. Cell Rep 35(8). https://doi.org/10.1016/j.celrep.2021.109172
Anand T, Vaid RK, Bera BC, Barua S, Riyesh T, Virmani N, Yadav N, Malik P (2015) Isolation and characterization of a bacteriophage with broad host range, displaying potential in preventing bovine diarrhoea. Virus Genes 51(2):315–321. https://doi.org/10.1007/s11262-015-1222-9
Beeton ML, Alves DR, Enright MC, Jenkins ATA (2015) Assessing phage therapy against Pseudomonas aeruginosa using a Galleria mellonella infection model. Int J Antimicrob Agents 46(2):196–200. https://doi.org/10.1016/j.ijantimicag.2015.04.005
Bernier Gosselin V, Dufour S, Adkins PRF, Middleton JR (2019) Persistence of coagulase negative staphylococcal intramammary infections in dairy goats. J Dairy Res 86(2):211–216. https://doi.org/10.1017/s0022029919000311
Besemer J, Lomsadze A, Borodovsky M (2001) GeneMarkS: a self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions. Nucleic Acids Res 29(12):2607–2618. https://doi.org/10.1093/nar/29.12.2607
Bhardwaj N, Bhardwaj SK, Mehta J, Kim KH, Deep A (2017) MOF-bacteriophage biosensor for highly sensitive and specific detection of Staphylococcus aureus. ACS Appl Mater Interfaces 9(39):33589–33598. https://doi.org/10.1021/acsami.7b07818
Borysowski J, Międzybrodzki R, Wierzbicki P, Kłosowska D, Korczak-Kowalska G, Weber-Dąbrowska B, Górski A (2017) A3R phage and Staphylococcus aureus lysate do not induce neutrophil degranulation. Viruses 9(2):36. https://doi.org/10.3390/v9020036
Brüssow H, Hendrix RW (2002) Phage genomics: small is beautiful. Cell 108(1):13–16. https://doi.org/10.1016/S0092-8674(01)00637-7
Cepko LCS, Garling EE, Dinsdale MJ, Scott WP, Bandy L, Nice T, Faber-Hammond J, Mellies JL (2020) Myoviridae phage PDX kills enteroaggregative Escherichia coli without human microbiome dysbiosis. J Med Microbiol 69(2):309–323. https://doi.org/10.1099/jmm.0.001162
Chmielewska-Jeznach M, Bardowski JK, Szczepankowska AK (2020) Lactococcus ceduovirus phages isolated from industrial dairy plants-from physiological to genomic analyses. Viruses 12(3):280. https://doi.org/10.3390/v12030280
Dakheel KH, Rahim RA, Neela VK, Al-Obaidi JR, Hun TG, Isa MNM, Yusoff K (2019) Genomic analyses of two novel biofilm-degrading methicillin-resistant Staphylococcus aureus phages. BMC Microbiol 19(1):114–114. https://doi.org/10.1186/s12866-019-1484-9
Das A, Mandal S, Hemmadi V, Ratre V, Biswas M (2020) Studies on the gene regulation involved in the lytic–lysogenic switch in Staphylococcus aureus temperate bacteriophage Phi11. J Biochem 168(6):659–668. https://doi.org/10.1093/jb/mvaa080
Dedrick RM, Guerrero-Bustamante CA, Garlena RA, Russell DA, Ford K, Harris K, Gilmour KC, Soothill J, Jacobs-Sera D, Schooley RT, Hatfull GF, Spencer H (2019) Engineered bacteriophages for treatment of a patient with a disseminated drug-resistant Mycobacterium abscessus. Nat Med 25(5):730–733. https://doi.org/10.1038/s41591-019-0437-z
Dinakaran V, Shankar M, Jayashree S, Rathinavel A, Gunasekaran P, Rajendhran J (2012) Genome sequence of Staphylococcus arlettae strain CVD059, isolated from the blood of a cardiovascular disease patient. J Bacteriol 194(23):6615–6616. https://doi.org/10.1128/JB.01732-12
Dunne M, Hupfeld M, Klumpp J, Loessner MJ (2018) Molecular basis of bacterial host interactions by Gram-positive targeting bacteriophages. Viruses 10(8):397. https://doi.org/10.3390/v10080397
Eckstein S, Stender J, Mzoughi S, Vogele K, Kühn J, Friese D, Bugert C, Handrick S, Ferjani M, Wölfel R, Millard A, Ben Moussa M, Bugert JJ (2021) Isolation and characterization of lytic phage TUN1 specific for Klebsiella pneumoniae K64 clinical isolates from Tunisia. BMC Microbiol 21(1):186. https://doi.org/10.1186/s12866-021-02251-w
Espejo RT, Canelo ES (1968) Properties of bacteriophage PM2: A lipid-containing bacterial virus. Virology 34(4):738–747. https://doi.org/10.1016/0042-6822(68)90094-9
Feng J, Gao L, Li L, Zhang Z, Wu C, Li F, Tong Y (2021) Characterization and genome analysis of novel Klebsiella phage BUCT556A with lytic activity against carbapenemase-producing Klebsiella pneumoniae. Virus Res 303:198506. https://doi.org/10.1016/j.virusres.2021.198506
Gao EB, Gui J-F, Zhang Q-Y (2012) A novel cyanophage with a cyanobacterial nonbleaching protein A gene in the genome. J Virol 86(1):236–245. https://doi.org/10.1128/JVI.06282-11
García P, Martínez B, Obeso JM, Lavigne R, Lurz R, Rodríguez A (2009) Functional genomic analysis of two Staphylococcus aureus phages isolated from the dairy environment. Appl Environ Microbiol 75(24):7663–7673. https://doi.org/10.1128/AEM.01864-09
Geng H, Zhang M, Li X, Wang L, Cong C, Cui H, Wang L, Xu Y (2020) Complete genome analysis of a Staphylococcus aureus phage (vBSM-A1). Arch Microbiol 202(7):1617–1626. https://doi.org/10.1007/s00203-020-01867-2
Goodrich-Blair H, Shub DA (1996) Beyond homing: competition between intron endonucleases confers a selective advantage on flanking genetic markers. Cell 84(2):211–221. https://doi.org/10.1016/S0092-8674(00)80976-9
Gordillo Altamirano FL, Barr JJ (2019) Phage therapy in the postantibiotic era. Clin Microbiol Rev 32(2):e00066-e18. https://doi.org/10.1128/CMR.00066-18
Grant JR, Stothard P (2008) The CGView Server: a comparative genomics tool for circular genomes. Nucleic Acids Res 36(Web Server issue):W181-W184. https://doi.org/10.1093/nar/gkn179
Guenther S, Huwyler D, Richard S, Loessner MJ (2009) Virulent bacteriophage for efficient biocontrol of Listeria monocytogenes in ready-to-eat foods. Appl Environ Microbiol 75(1):93–100. https://doi.org/10.1128/AEM.01711-08
Guenther S, Herzig O, Fieseler L, Klumpp J, Loessner MJ (2012) Biocontrol of Salmonella typhimurium in RTE foods with the virulent bacteriophage FO1-E2. Int J Food Microbiol 154(1–2):66–72. https://doi.org/10.1016/j.ijfoodmicro.2011.12.023
Hargreaves KR, Kropinski AM, Clokie MRJ (2014) What does the talking?: quorum sensing signalling genes discovered in a bacteriophage genome. PLoS One 9(1):e85131–e85131. https://doi.org/10.1371/journal.pone.0085131
Hargreaves KR, Clokie MRJ (2014) Clostridium difficile phages: still difficult? [Review]. Front Microbiol 5(184). https://doi.org/10.3389/fmicb.2014.00184
Hatfull GF (2008) Bacteriophage genomics. Curr Opin Microbiol 11(5):447–453. https://doi.org/10.1016/j.mib.2008.09.004
Hatoum-Aslan A (2021) The phages of staphylococci: critical catalysts in health and disease. Trends Microbiol. https://doi.org/10.1016/j.tim.2021.04.008
Holguín AV, Rangel G, Clavijo V, Prada C, Mantilla M, Gomez MC, Kutter E, Taylor C, Fineran PC, Barrios AFG, Vives MJ (2015) Phage ΦPan70, a putative temperate phage, controls Pseudomonas aeruginosa in planktonic, biofilm and burn mouse model assays. Viruses 7(8):4602–4623. https://doi.org/10.3390/v7082835
Hou D, Mai C, Hou C (2000) A case report: Staphylococcus arlette causes chronic prostatitis (Chinese). Chin J Derm Venereol 14(3):181. https://doi.org/10.3969/j.issn.1001-7089.2000.03.026
Ingmer H, Gerlach D, Wolz C (2019) Temperate phages of Staphylococcus aureus. Microbiol Spectr 7(5). https://doi.org/10.1128/microbiolspec.GPP3-0058-2018
Kazi M, Annapure US (2016) Bacteriophage biocontrol of foodborne pathogens. J Food Sci Technol 53(3):1355–1362. https://doi.org/10.1007/s13197-015-1996-8
Kim JS, Hosseindoust A, Lee SH, Choi YH, Kim MJ, Lee JH, Kwon IK, Chae BJ (2017) Bacteriophage cocktail and multi-strain probiotics in the feed for weanling pigs: effects on intestine morphology and targeted intestinal coliforms and Clostridium. Animal 11(1):45–53. https://doi.org/10.1017/S1751731116001166
Kittler S, Fischer S, Abdulmawjood A, Glünder G, Klein G (2013) Effect of bacteriophage application on Campylobacter jejuni loads in commercial broiler flocks. Appl Environ Microbiol 79(23):7525–7533. https://doi.org/10.1128/AEM.02703-13
Kumar P, Meghvansi MK, Kamboj DV (2021) Isolation, phenotypic characterization and comparative genomic analysis of 2019SD1, a polyvalent enterobacteria phage. Sci Rep 11(1):22197. https://doi.org/10.1038/s41598-021-01419-8
Kyoung Min G, In Young C, **young L, Jun-Hyun O, Mi-Kyung P (2018) Isolation and characterization of a lytic and highly specific phage against Yersinia enterocolitica as a novel biocontrol agent. J Microbiol Biotechnol 28(11):1946–1954. https://doi.org/10.4014/jmb.1808.08001
Kyrkou I, Byth Carstens A, Ellegaard-Jensen L, Kot W, Zervas A, Djurhuus AM, Neve H, Hansen M, Hestbjerg Hansen L (2019) Expanding the diversity of Myoviridae phages infecting Lactobacillus plantarum-a novel lineage of Lactobacillus phages comprising five new members. Viruses 11(7):611. https://doi.org/10.3390/v11070611
Kyrkou I, Carstens AB, Ellegaard-Jensen L, Kot W, Zervas A, Djurhuus AM, Neve H, Franz CMAP, Hansen M, Hansen LH (2020) Isolation and characterisation of novel phages infecting Lactobacillus plantarum and proposal of a new genus, “Silenusvirus.” Sci Rep 10(1):8763–8763. https://doi.org/10.1038/s41598-020-65366-6
Lakshmanan RS, Guntupalli R, Hu J, Kim DJ, Petrenko VA, Barbaree JM, Chin BA (2007) Phage immobilized magnetoelastic sensor for the detection of Salmonella typhimurium. J Microbiol Methods 71(1):55–60. https://doi.org/10.1016/j.mimet.2007.07.012
Langdon A, Crook N, Dantas G (2016) The effects of antibiotics on the microbiome throughout development and alternative approaches for therapeutic modulation. Genome Med 8(1):39–39. https://doi.org/10.1186/s13073-016-0294-z
Lavecchia A, Chiara M, De Virgilio C, Manzari C, Monno R, De Carlo A, Pazzani C, Horner D, Pesole G, Placido A (2019) Staphylococcus arlettae genomics: novel insights on candidate antibiotic resistance and virulence genes in an emerging opportunistic pathogen. Microorganisms 7(11):580. https://doi.org/10.3390/microorganisms7110580
Lawrence D, Baldridge MT, Handley SA (2019) Phages and human health: more than idle hitchhikers. Viruses 11(7):587. https://doi.org/10.3390/v11070587
Lindsay JA, Ruzin A, Ross HF, Kurepina N, Novick RP (1998) The gene for toxic shock toxin is carried by a family of mobile pathogenicity islands in Staphylococcus aureus. Mol Microbiol 29(2):527–543. https://doi.org/10.1046/j.1365-2958.1998.00947.x
Lowe TM, Chan PP (2016) tRNAscan-SE On-line: integrating search and context for analysis of transfer RNA genes. Nucleic Acids Res 44(W1):W54–W57. https://doi.org/10.1093/nar/gkw413
Lu L, Cai L, Jiao N, Zhang R (2017) Isolation and characterization of the first phage infecting ecologically important marine bacteria Erythrobacter. Virology J 14(1):104–104. https://doi.org/10.1186/s12985-017-0773-x
Łusiak-Szelachowska M, Żaczek M, Weber-Dąbrowska B, Międzybrodzki R, Letkiewicz S, Fortuna W, Rogóż P, Szufnarowski K, Jończyk-Matysiak E, Olchawa E, Walaszek KM, Górski A (2017) Antiphage activity of sera during phage therapy in relation to its outcome. Future Microbiol 12:109–117. https://doi.org/10.2217/fmb-2016-0156
Ly M, Abeles SR, Boehm TK, Robles-Sikisaka R, Naidu M, Santiago-Rodriguez T, Pride DT (2014) Altered oral viral ecology in association with periodontal disease. mBio 5(3):e01133. https://doi.org/10.1128/mBio.01133-14
Ma Y, You X, Mai G, Tokuyasu T, Liu C (2018) A human gut phage catalog correlates the gut phageome with type 2 diabetes. Microbiome 6(1):24–24. https://doi.org/10.1186/s40168-018-0410-y
Madurantakam Royam M, Nachimuthu R (2020) Isolation, characterization, and efficacy of bacteriophages isolated against Citrobacter spp. an in vivo approach in a zebrafish model (Danio rerio). Res Microbiol 171(8):341–350. https://doi.org/10.1016/j.resmic.2020.08.003
Mahony J, Alqarni M, Stockdale S, Spinelli S, Feyereisen M, Cambillau C, Sinderen DV (2016) Functional and structural dissection of the tape measure protein of lactococcal phage TP901-1. Sci Rep 6:36667–36667. https://doi.org/10.1038/srep36667
Merabishvili M, Pirnay J-P, Verbeken G, Chanishvili N, Tediashvili M, Lashkhi N, Glonti T, Krylov V, Mast J, Van Parys L, Lavigne R, Volckaert G, Mattheus W, Verween G, De Corte P, Rose T, Jennes S, Zizi M, De Vos D, Vaneechoutte M (2009) Quality-controlled small-scale production of a well-defined bacteriophage cocktail for use in human clinical trials. PLoS ONE 4(3):e4944–e4944. https://doi.org/10.1371/journal.pone.0004944
Międzybrodzki R, Borysowski J, Kłak M, Jończyk-Matysiak E, Obmińska-Mrukowicz B, Suszko-Pawłowska A, Bubak B, Weber-Dąbrowska B, Górski A (2017) In vivo studies on the influence of bacteriophage preparations on the autoimmune inflammatory process. Biomed Res Int 2017:3612015–3612015. https://doi.org/10.1155/2017/3612015
Miller-Ensminger T, Mormando R, Maskeri L, Shapiro JW, Wolfe AJ, Putonti C (2020) Introducing Lu-1, a novel Lactobacillus jensenii phage abundant in the urogenital tract. PLoS One 15(6):e0234159–e0234159. https://doi.org/10.1371/journal.pone.0234159
Mitchell MS, Rao VB (2006) Functional analysis of the bacteriophage T4 DNA-packaging ATPase motor*. J Biol Chem 281(1):518–527. https://doi.org/10.1074/jbc.M507719200
Mohamed H, Hausner G, Bonen L (2012) Homing endonucleases: DNA scissors on a mission. Genome 55(8):553–569. https://doi.org/10.1139/g2012-049%M22891613
Moller AG, Lindsay JA, Read TD (2019) Determinants of phage host range in Staphylococcus species. Appl Environ Microbiol 85(11):e00209-00219. https://doi.org/10.1128/AEM.00209-19
Muharram, M. M., Abulhamd, A. T., Aldawsari, M. F., Alqarni, M. H., & Labrou, N. E. (2020). Development of Staphylococcus enzybiotics: the Ph28 gene of Staphylococcus epidermidis phage PH15 is a two-domain endolysin. Antibiotics (Basel), 9(4). https://doi.org/10.3390/antibiotics9040148
Nasioudis D, Linhares IM, Ledger WJ, Witkin SS (2017) Bacterial vaginosis: a critical analysis of current knowledge. BJOG 124(1):61–69. https://doi.org/10.1111/1471-0528.14209
Novick RP, Christie GE, Penadés JR (2010) The phage-related chromosomal islands of Gram-positive bacteria. Nat Rev Microbiol 8(8):541–551. https://doi.org/10.1038/nrmicro2393
Oduor JMO, Kadija E, Nyachieo A, Mureithi MW, Skurnik M (2020) Bioprospecting Staphylococcus phages with therapeutic and bio-control potential. Viruses 12(2):133. https://doi.org/10.3390/v12020133
Oliveira H, Melo LDR, Santos SB, Nóbrega FL, Ferreira EC, Cerca N, Azeredo J, Kluskens LD (2013) Molecular aspects and comparative genomics of bacteriophage endolysins. J Virol 87(8):4558–4570. https://doi.org/10.1128/JVI.03277-12
Oliveira H, Sampaio M, Melo LDR, Dias O, Pope WH, Hatfull GF, Azeredo J (2019) Staphylococci phages display vast genomic diversity and evolutionary relationships. BMC Genomics 20(1):357. https://doi.org/10.1186/s12864-019-5647-8
Pan L, Li D, Sun Z, Lin W, Hong B, Qin W, Xu L, Liu W, Zhou Q, Wang F, Cai R, Qian M, Tong Y (2021) First characterization of a hafnia phage reveals extraordinarily large burst size and unusual plaque polymorphism. Front Microbiol 12:754331. https://doi.org/10.3389/fmicb.2021.754331
Park J, Friendship RM, Weese JS, Poljak Z, Dewey CE (2013) An investigation of resistance to β-lactam antimicrobials among staphylococci isolated from pigs with exudative epidermitis. BMC Vet Res 9:211–211. https://doi.org/10.1186/1746-6148-9-211
Pertics BZ, Szénásy D, Dunai D, Born Y, Fieseler L, Kovács T, Schneider G (2020) Isolation of a novel lytic bacteriophage against a nosocomial methicillin-resistant Staphylococcus aureus belonging to ST45. Biomed Res Int 2020:5463801. https://doi.org/10.1155/2020/5463801
Pincus NB, Reckhow JD, Saleem D, Jammeh ML, Datta SK, Myles IA (2015) Strain specific phage treatment for Staphylococcus aureus infection is influenced by host immunity and site of infection. PLoS One 10(4):e0124280–e0124280. https://doi.org/10.1371/journal.pone.0124280
Rhoads DD, Wolcott RD, Kuskowski MA, Wolcott BM, Ward LS, Sulakvelidze A (2009) Bacteriophage therapy of venous leg ulcers in humans: results of a phase I safety trial. J Wound Care 18(6):237–243. https://doi.org/10.12968/jowc.2009.18.6.42801
Roach DR, Debarbieux L (2017) Phage therapy: awakening a slee** giant. Emerg Top Life Sci 1(1):93–103. https://doi.org/10.1042/etls20170002
Rohde C, Wittmann J, Kutter E (2018) Bacteriophages: a therapy concept against multi-drug-resistant bacteria. Surg Infect 19(8):737–744. https://doi.org/10.1089/sur.2018.184
Rose T, Verbeken G, Vos DD, Merabishvili M, Vaneechoutte M, Lavigne R, Jennes S, Zizi M, Pirnay J-P (2014) Experimental phage therapy of burn wound infection: difficult first steps. Int J Burns Trauma 4(2):66–73. https://pubmed.ncbi.nlm.nih.gov/25356373
Schmelcher M, Loessner MJ (2014) Application of bacteriophages for detection of foodborne pathogens. Bacteriophage 4(1):e28137. https://doi.org/10.4161/bact.28137
Seed KD, Dennis JJ (2009) Experimental bacteriophage therapy increases survival of Galleria mellonella larvae infected with clinically relevant strains of the Burkholderia cepacia complex. Antimicrob Agents Chemother 53(5):2205–2208. https://doi.org/10.1128/AAC.01166-08
Semler DD, Goudie AD, Finlay WH, Dennis JJ (2014) Aerosol phage therapy efficacy in Burkholderia cepacia complex respiratory infections. Antimicrob Agents Chemother 58(7):4005–4013. https://doi.org/10.1128/AAC.02388-13
Shareefdeen H, Hynes AP (2021) Does over a century of aerobic phage work provide a solid framework for the study of phages in the gut? Anaerobe 68:102319. https://doi.org/10.1016/j.anaerobe.2021.102319
Singh A, Poshtiban S, Evoy S (2013) Recent advances in bacteriophage based biosensors for food-borne pathogen detection. Sensors (basel, Switzerland) 13(2):1763–1786. https://doi.org/10.3390/s130201763
Söding J, Biegert A, Lupas AN (2005) The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res 33(Web Server issue):W244-W248. https://doi.org/10.1093/nar/gki408
van Zyl LJ, Sunda F, Taylor MP, Cowan DA, Trindade MI (2015) Identification and characterization of a novel Geobacillus thermoglucosidasius bacteriophage, GVE3. Arch Virol 160(9):2269–2282. https://doi.org/10.1007/s00705-015-2497-9
van Zyl LJ, Abrahams Y, Stander EA, Kirby-McCollough B, Jourdain R, Clavaud C, Breton L, Trindade M (2018) Novel phages of healthy skin metaviromes from South Africa. Sci Rep 8(1):12265. https://doi.org/10.1038/s41598-018-30705-1
Wagner J, Maksimovic J, Farries G, Sim WH, Bishop RF, Cameron DJ, Catto-Smith AG, Kirkwood CD (2013) Bacteriophages in gut samples from pediatric Crohn’s disease patients: metagenomic analysis using 454 pyrosequencing. Inflamm Bowel Dis 19(8):1598–1608. https://doi.org/10.1097/MIB.0b013e318292477c
Wang CX, Li X (2018) JMT-1: a novel, spherical lytic halotolerant phage isolated from Yuncheng saline lake. Braz J Microbiol 49(Suppl 1):262–268. https://doi.org/10.1016/j.bjm.2018.03.004
Wang J, Gao Y, Zhao F (2016) Phage–bacteria interaction network in human oral microbiome. Environ Microbiol 18(7):2143–2158. https://doi.org/10.1111/1462-2920.12923
Wei B, Cong C, Yu W, Xu Y, Li J, Li S (2020) Isolation, identification, biological characteristics and whole genomic analysis of a multi-drug resistant E coli phage. J Jilin Agric Univ. https://doi.org/10.13327/j.jjlau.2020.5773
Wintachai P, Naknaen A, Thammaphet J, Pomwised R, Phaonakrop N, Roytrakul S, Smith DR (2020) Characterization of extended-spectrum-β-lactamase producing Klebsiella pneumoniae phage KP1801 and evaluation of therapeutic efficacy in vitro and in vivo. Sci Rep 10(1):11803. https://doi.org/10.1038/s41598-020-68702-y
Wright A, Hawkins CH, Änggård EE, Harper DR (2009) A controlled clinical trial of a therapeutic bacteriophage preparation in chronic otitis due to antibiotic-resistant Pseudomonas aeruginosa; a preliminary report of efficacy. Clin Otolaryngol 34(4):349–357. https://doi.org/10.1111/j.1749-4486.2009.01973.x
**ao H, Yuan C, Liang S, Chen Z, Jiang X (2021) A case report and review of the literature: otologic infection due to Staphylococcus arlettae. Int J Lab Med 42(6):764–766. https://doi.org/10.3969/j.issn.1673-4130.2021.06.031
Zhang D, You F, He Y, Te SH, Gin KY-H (2020) Isolation and characterization of the first freshwater cyanophage infecting Pseudanabaena. J Virol 94(17):e00682-e620. https://doi.org/10.1128/JVI.00682-20
Zurabov F, Zhilenkov E (2021) Characterization of four virulent Klebsiella pneumoniae bacteriophages, and evaluation of their potential use in complex phage preparation. Virol J 18(1):9–9. https://doi.org/10.1186/s12985-020-01485-w
Acknowledgements
We would like to thank Editage (www.editage.cn) for English language editing.
Funding
This study was supported by the Key R&D Program of the Department of Science and Technology of Gansu Province (No. 20YF8NA029), Project of the Department of Science and Technology of Sichuan Province (No. 2019YJ0671), and Innovation Research Group on Quality and Safety of Featured Agricultural Products in Three Gorges Reservoir Area (No. CXQTP19037).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Conceptualisation: Guangli Han and Xue** Yao; methodology: Zidan Luo; investigation: Biao Lu and Jieru Zhang; resources: Pengfei Zhang; data curation: Guangli Han; writing, original draft preparation: Guangli Han; review and supervision: Kang Yong, Yin Wang, Yan Luo, Zexiao Yang, Meishen Ren, Xue** Yao, and Suizhong Cao. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All experimental protocols in this study were approved by experimental ethics committee of Sichuan Agricultural University. All methods were carried out in accordance with the 1964 Helsinki declaration and its later amendments ethical standards. The collection of milk samples had been permitted by dairy farm.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Guangli Han and Jieru Zhang contributed equally to this work and share first authorship.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Han, G., Zhang, J., Luo, Z. et al. Characteristics of a novel temperate bacteriophage against Staphylococcus arlettae (vB_SarS_BM31). Int Microbiol 26, 327–341 (2023). https://doi.org/10.1007/s10123-022-00292-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10123-022-00292-3