Abstract
It is a classical result of Stein and Waterman that the asymptotic number of RNA secondary structures is \(1.104366 \cdot n^{-3/2} \cdot 2.618034^n\). Motivated by the kinetics of RNA secondary structure formation, we are interested in determining the asymptotic number of secondary structures that are locally optimal, with respect to a particular energy model. In the Nussinov energy model, where each base pair contributes \(-1\) towards the energy of the structure, locally optimal structures are exactly the saturated structures, for which we have previously shown that asymptotically, there are \(1.07427\cdot n^{-3/2} \cdot 2.35467^n\) many saturated structures for a sequence of length \(n\). In this paper, we consider the base stacking energy model, a mild variant of the Nussinov model, where each stacked base pair contributes \(-1\) toward the energy of the structure. Locally optimal structures with respect to the base stacking energy model are exactly those secondary structures, whose stems cannot be extended. Such structures were first considered by Evers and Giegerich, who described a dynamic programming algorithm to enumerate all locally optimal structures. In this paper, we apply methods from enumerative combinatorics to compute the asymptotic number of such structures. Additionally, we consider analogous combinatorial problems for secondary structures with annotated single-stranded, stacking nucleotides (dangles).







Similar content being viewed by others
Notes
If the energy \(E(S)=0\) or if the temperature \(T=+\infty \), then the partition function is exactly equal to the number of secondary structures.
Jean Gaston Darboux (1842–1917).
The shape of a secondary structure was defined by Voss et al. (2006) to represent its branching topology; for instance, the shape of the well-known clover-leaf structure of tRNA is \(\,\mathtt [ \,\,\mathtt [ \,\,\mathtt ] \,\,\mathtt [ \,\,\mathtt ] \,\,\mathtt [ \,\,\mathtt ] \,\,\mathtt ] \,\). The asymptotic number of shapes for a length \(n\) sequence yields the run time for the Giegerich Lab software RNAshapes on length \(n\) sequences, since Steffen et al. (2006) report that RNAshapes runs in time \(O(n^3 k s)\) for \(s\) sequences, each of length at most \(n\) and \(k\) shapes.
To the best of our knowledge, UNAFOLD is currently the only software that computes the partition function over all secondary structures in a mathematically rigorous manner.
This is done, for instance, in grammar \(G_4\) by replacing the rule \(\bullet _{\ge \theta } \rightarrow \bullet \) by \(\bullet _{\ge \theta } \rightarrow \bullet ^{\theta }\), where \(\bullet ^{\theta }\) consists of \(\theta \) occurrences of \(\bullet \).
Our grammar \(G_1\) is equivalent to the “tree grammar nussinov78” from Steffen and Giegerich (2005).
It is clear that the number of structures equals the partition function \(\sum _{S} \exp (-E(S)/RT)\) provided that \(E(S)=0\).
Alternatively, and more simply, we could have produced this curve from the Taylor coefficients of the expressions to the right of the limit in equations (3) and (6), after first solving for \(S(z,u)\) [resp. \(S(z,u,v)\)] in equation (1) [resp. (4)].
Fig. 2 Theoretical melting curve for two simple energy models of RNA secondary structure. Temperature in Celsius is given on the \(x\)-axis, while expected number of base pairs is given on the \(y\)-axis. We implemented an algorithm, using dynamic programming, with run time \(O(n^5)\) and space \(O(n^3)\), to compute the partition function \(Z_k = \sum _{S \in \mathbb S _k} \exp (-E(S)/RT)\), where \(\mathbb ( S)_k\) denotes the set of all secondary structures for a homopolymer of length 100 nt, having exactly \(k\) base pairs. The expected number of base pairs is thus \(\sum _k k \cdot p_k\), where \(p_k = \frac{Z_k}{Z}\) denotes the probability that a secondary structure has \(k\) base pairs, and \(Z\) denotes the full partition function \(Z = \sum _{S} \exp (-E(S)/RT) = \sum _k Z_k\). (Alternatively, and more simply, we could have produced this curve from the Taylor coefficients of the expressions to the right of the limit in equations (3) and (6), after first solving for \(S(z,u)\) [resp. \(S(z,u,v)\)] in equation (1) [resp. (4)].) In the Nussinov–Jacobson energy model (Nussinov and Jacobson 1980), \(E(S)\) is defined to be \(-1\cdot |S|\); i.e. \(-1\) times the number of base pairs of \(S\). In the base stacking energy model, \(E(S)\) is defined to be \(-1\) times the number of stacked base pairs of \(S\). Although both models are quite similar, we see that the melting curves are indeed different, where the base stacking model entails more cooperative folding (see Dill and Bromberg 2002 for discussion of cooperative folding)
We use the subscript notation for partial derivatives.
Exact base stacking parameters are ignored as is entropy; however, the context-free grammar allows the separate marking of distinct features, such as stacked base pairs, hairpins, bulges, internal loops, multiloops.
References
Bender EA (1973) Central and local limit theorem applied to asymptotic enumeration. J Combin Theory Ser A 15:91–111
Clote P (2005) An efficient algorithm to compute the landscape of locally optimal RNA secondary structures with respect to the Nussinov-Jacobson energy model. J Comput Biol 12(1):83–101
Clote P (2006) Combinatorics of saturated secondary structures of RNA. J Comput Biol 13(9):1640–1657
Clote P, Kranakis E, Krizanc D, Salvy B (2009) Asymptotics of canonical and saturated RNA secondary structures. J Bioinform Comput Biol 7(5):869–893
Clote P, Dobrev S, Dotu I, Kranakis E, Krizanc D, Urrutia J (2012) On the page number of RNA secondary structures with pseudoknots. J Math Biol 65:1337–1257
Dill KA, Bromberg S (2002) Molecular driving forces: statistical thermodynamics in chemistry and biology. Garland Publishing Inc., New York
Drmota M (1997) Systems of functional equations. Random Struct Algorithms 10(1–2):103–124
Drmota M, Fusy É, Jué J, Kang M, Kraus V (2011) Asymptotic study of subcritical graph classes. SIAM J Discrete Math 25(4):1615–1651
Evers DJ, Giegerich R (2001) Reducing the conformation space in RNA structure prediction. In: German conference on bioinformatics (GCB’01), pp 1–6
Flajolet P, Odlyzko A (1990) Singularity analysis of generating functions. SIAM J Discrete Math 3(2):216–240
Flajolet P, Sedgewick R (2009) Analytic combinatorics. Cambridge University Press, Cambridge
Harer J, Zagier D (1986) The Euler characteristic of the moduli space of curves. Invent Math 85(3):457–485
Haslinger C, Stadler PF (1999) RNA structures with pseudo-knots: graph-theoretical, combinatorial, and statistical properties. Bull Math Biol 61(3):437–467
Henrici P (1991) Applied and computational complex analysis, vol 2. Wiley Classics Library, Wiley, New York
Hofacker I (2003) Vienna RNA secondary structure server. Nucleic Acids Res 31(13):3429–3431
Hofacker IL, Fontana W, Stadler PF, Bonhoeffer LS, Tacker M, Schuster P (1994) Fast folding and comparison of RNA secondary structures. Monatsch Chem 125:167–188
Hofacker IL, Schuster P, Stadler PF (1998) Combinatorics of RNA secondary structures. Discr Appl Math 88:207–237
Knudsen B, Hein J (2003) Pfold: RNA secondary structure prediction using stochastic context-free grammars. Nucleic Acids Res 31(13):3423–3428
Li TJ, Reidys CM (2012) Combinatorics of RNA-RNA interaction. J Math Biol 64(3):529–556
Lorenz WA, Ponty Y, Clote P (2008) Asymptotics of RNA shapes. J Comput Biol 15(1):31–63
Lyngso RB, Pedersen CN (2000) RNA pseudoknot prediction in energy-based models. J Comput Biol 7(3–4):409–427
Markham NR (2006) Algorithms and software for nucleic acid sequences. PhD, Rensselaer Polytechnic Institute, under the direction of M. Zuker
Markham NR, Zuker M (2008) UNAFold: software for nucleic acid folding and hybridization. Methods Mol Biol 453:3–31
Matthews DH, Sabina J, Zuker M, Turner DH (1999) Expanded sequence dependence of thermodynamic parameters improves prediction of RNA secondary structure. J Mol Biol 288:911–940
McCaskill JS (1990) The equilibrium partition function and base pair binding probabilities for RNA secondary structure. Biopolymers 29:1105–1119
Meir A, Moon JW (1989) On an asymptotic method in enumeration. J Combin Theory Ser A 51:77–89
Nussinov R, Jacobson AB (1980) Fast algorithm for predicting the secondary structure of single stranded RNA. Proc Natl Acad Sci USA 77(11):6309–6313
Pemantle R, Wilson MC (2008) Twenty combinatorial examples of asymptotics derived from multivariate generating functions. SIAM Rev 50(2):199–272
Rodland EA (2006) Pseudoknots in RNA secondary structures: representation, enumeration, and prevalence. J Comput Biol 13(6):1197–1213
Saule C, Regnier M, Steyaert JM, Denise A (2011) Counting RNA pseudoknotted structures. J Comput Biol 18(10):1339–1351
Sheikh S, Backofen R, Ponty Y (2012) Impact of the energy model on the complexity of RNA folding with pseudoknots. In: Lecture notes in computer science, vol 7354. 23rd Annual symposium on combinatorial pattern matching, CPM 2012, Helsinki, Finland, pp 321–333
Steffen P, Giegerich R (2005) Versatile and declarative dynamic programming using pair algebras. BMC Bioinformatics 6:224
Steffen P, Voss B, Rehmsmeier M, Reeder J, Giegerich R (2006) RNAshapes: an integrated RNA analysis package based on abstract shapes. Bioinformatics 22(4):500–503
Stein PR, Waterman MS (1978) On some new sequences generalizing the Catalan and Motzkin numbers. Discrete Math 26:261–272
Tabaska JE, Cary RE, Gabow HN, Stormo GD (1998) An RNA folding method capable of identifying pseudoknots and base triples. Bioinformatics 14:691–699
Vernizzi G, Orland H, Zee A (2005) Enumeration of RNA structures by matrix models. Phys Rev Lett 94(16):168103
Voss B, Giegerich R, Rehmsmeier M (2006) Complete probabilistic analysis of RNA shapes. BMC Biol 4(1):5–27
Waterman MS (1978) Secondary structure of single-stranded nucleic acids. Stud Found Combin: Adv Math Supplem Stud 1:167–212
**a T Jr, SantaLucia J, Burkard ME, Kierzek R, Schroeder SJ, Jiao X, Cox C, Turner DH (1999) Thermodynamic parameters for an expanded nearest-neighbor model for formation of RNA duplexes with Watson-Crick base pairs. Biochemistry 37:14719–14735
Yoffe AM, Prinsen P, Gelbart WM, Ben-Shaul A (2011) The ends of a large RNA molecule are necessarily close. Nucleic Acids Res 39(1):292–299
Zuker M (1986) RNA folding prediction: the continued need for interaction between biologists and mathematicians. Lect Math Life Sci 17:87–124
Zuker M (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31(13):3406–3415
Zuker M, Stiegler P (1981) Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res 9:133–148
Zuker M, Mathews DH, Turner DH (1999) Algorithms and thermodynamics for RNA secondary structure prediction: a practical guide. In: Barciszewski J, Clark BFC (eds) RNA biochemistry and biotechnology. NATO ASI series. Kluwer, Dordrecht, pp 11–43
Acknowledgments
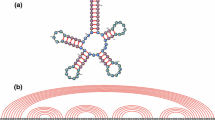
Figure 1 was created by W.A. Lorenz and H. Jabbari. We would like to thank the anonymous referees for their helpful comments. É. Fusy is supported by the European project ExploreMaps—-ERC StG 208471. P. Clote is supported by the National Science Foundation under grants DBI-0543506 and DMS-0817971, and by Digiteo Foundation project RNAomics. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of the National Science Foundation.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Fusy, É., Clote, P. Combinatorics of locally optimal RNA secondary structures. J. Math. Biol. 68, 341–375 (2014). https://doi.org/10.1007/s00285-012-0631-9
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00285-012-0631-9