Abstract
The outbreak of novel coronavirus disease 2019 (COVID-19) caused by severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) virus continually led to infect a large population worldwide. SARS-CoV-2 utilizes its NSP6 and Orf9c proteins to interact with sigma receptors that are implicated in lipid remodeling and ER stress response, to infect cells. The drugs targeting the sigma receptors, sigma-1 and sigma-2, have emerged as effective candidates to reduce viral infectivity, and some of them are in clinical trials against COVID-19. The antipsychotic drug, haloperidol, exerts remarkable antiviral activity, but, at the same time, the sigma-1 benzomorphan agonist, dextromethorphan, showed pro-viral activity. To explore the potential mechanisms of biased binding and activity of the two drugs, haloperidol and dextromethorphan towards NSP6, we herein utilized molecular docking–based molecular dynamics simulation studies. Our extensive analysis of the protein-drug interactions, structural and conformational dynamics, residual frustrations, and molecular switches of NSP6-drug complexes indicates that dextromethorphan binding leads to structural destabilization and increase in conformational dynamics and energetic frustrations. On the other hand, the strong binding of haloperidol leads to minimal structural and dynamical perturbations to NSP6. Thus, the structural insights of stronger binding affinity and favorable molecular interactions of haloperidol towards viral NSP6 suggests that haloperidol can be potentially explored as a candidate drug against COVID-19.
Key messages
•Inhibitors of sigma receptors are considered as potent drugs against COVID-19.
•Antipsychotic drug, haloperidol, binds strongly to NSP6 and induces the minimal changes in structure and dynamics of NSP6.
•Dextromethorphan, agonist of sigma receptors, binding leads to overall destabilization of NSP6.
•These two drugs bind with NSP6 differently and also induce differences in the structural and conformational changes that explain their different mechanisms of action.
•Haloperidol can be explored as a candidate drug against COVID-19.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The current outbreak of corona virus disease 2019 (COVID-19) caused by a novel coronavirus SARS-CoV-2 was first reported from Wuhan, China, in late December 2019 [1], which has subsequently affected the entire world, reporting nearly 26 million of confirmed cases of COVID-19 along with ~ 9.0-lakh deaths as per data recorded in September 1st week, 2020, posing a global threat for human health and economy. With so many novel studies and findings surfaced, since its inception, we are still lagging behind in development of an effective treatment strategy to control the virus spread and prevent the disease [2,3,4,5,6,7].
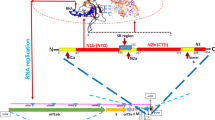
SARS-CoV-2 is an enveloped non-segmented large positive sense, single-stranded RNA virus (~ 30 kb) with 5′-cap structure and 3′-poly-A tail belonging to β-CoV category [8, 9]. Its RNA genome contains 29,891 nucleotides and encoding for ~ 9860 amino acids [9]. The genome codes for both structural proteins like spike (S), envelope (E), membrane (M), and nucleocapsid (N), along with many non-structural proteins (NSPs 1–16) [10]. While these NSPs linked to RNA replication and processing of subgenomic RNAs, the functions of some of the NSPs are not known. A key component, NSP6, is a membrane protein of approximately 34 kDa with eight transmembrane helices and a highly conserved C-terminus. Together with NSP3 and NSP4, NSP6 is involved in the formation of replication-transcription complexes (RTCs) or replication organelles (RO) by stimulating the rearrangement of host cell membranes [11]. These replication complexes serve many important functions during the virus life cycle and play an important role in infection [12, S1). The drug binding thus increases the number of highly frustrated residues and thus increases the local frustration and flexibility of the protein, especially in dextromethorphan binding.
Moreover, the residues involved in drug binding were dominantly neutral to minimal frustration. However, some of the dextromethorphan-binding residues like Ser32 and Leu276 gain frustration upon binding, whereas the frustration of Tyr132, Tyr136, Gly177, Arg236, and Lys270 decreased upon binding. Also, haloperidol binding increases the frustration of Leu231, Leu237, and Thr238 upon binding, and at the same time, it also decreased the frustration of residues like Tyr136, Phe184, and Phe269 (Table S1).
Moreover, to find out how drug binding induced the local frustration changes, we performed a comparative analysis of the spatial distribution of local frustrations mapped onto the secondary structure of the protein (Fig. 9). In particular, highly frustrated residues in the NSP6-dextromethorphan complex increased in the α-helix H1, H7, and H13, while the increase is seen in H11 for the NSP-haloperidol complex (Fig. 9a). The decrease in frustration was also seen for both the complexes mainly in C-terminal H11, H12, S1, and H14 for the NSP6-dextromethorphan complex, and for NSP6-haloperidol, a decrease is observed in H5, H8, and S1 (Fig. 9a). Additionally, the drug binding decreased the minimal frustration in H5, H7, H9, and H11 helices, and also increased the minimal frustration in H2 and H12 helices (Fig. 9b).
Frustration analysis in NSP6-drug complexes. The changes in residual frustration are distributed along the structural regions of NSP6. a The changes in minimal frustration values in NSP6-drug complex. b The changes in highly frustration values in NSP6-drug complex. The secondary structural regions are represented as: H-α-helix and S-β-strands
Thus, the coupling between structurally rigid C-terminal helices H10 and H11, and conformationally flexible helices, H5 and H13, is important for drug binding, and also the frustration index of the regions close to the binding site changes upon association.
Discussion
The all-atom MD simulation shows significant differences in the tertiary structure of the NSP6-drug complexes. The structural snapshots of the protein-drug complex were used to analyze the differences in tertiary structure caused by each drug (Figure S8).
Overall comparison of the binding conformations of haloperidol and dextromethorphan in NSP6
The NSP6-drug complexes show the significant differences in H2, H5, H7, and C-terminal regions comprising H12, H13, β1, and β2 (Figure S8). The haloperidol and dextromethorphan complexes have a kink in H2, H5, and H7 at midway of simulation (50 ns) that causes a change in orientation when the drug binds (Figure S8 A, C). The dextromethorphan binding induces much larger kink at H2 and H5 with a much larger deviation also (RMSD = 3.59) (Figure S8C). At the end of the simulation, the haloperidol and dextromethorphan complexes showed an increase in twist of H12 and H13, and both the strands of the C-termini along with the kinks present in H2, H5, and H7 (Figure S8B, D). Similar to what was observed at ~ 50 ns, dextromethorphan binding induces much larger disruption (RMSD = 3.24). Thus, although both drugs lead to bending of the helices, dextromethorphan binding induces larger disruption of the helices and thus more destabilization observed in the protein.
The protein-drug binding analysis shows that dextromethorphan and haloperidol interact with many key residues and are shared between both the drugs. The residues interacting with each drug during the simulation were compared and visual 2D representations are shown in Figures S9 and S10.
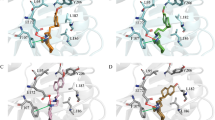
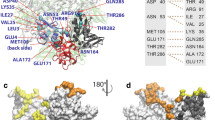
Figure 2 a shows the initial pose of the dextromethorphan molecule in the MD simulation, which is also the best pose from the docking study. As can be seen, the dextromethorphan molecule formed hydrogen bonds with residue Lys 61, and several van der Waals interactions with His62, Asn232, Arg233, Arg236, Thr238, Asp243, Leu245, and Pro282. Figure 2 b shows the haloperidol binding sites in NSP6 which is comprised of helix H7, and H9 residues (i.e., H7: Tyr132, Asp134, Ala136, Arg137, Trp140; H9: Asn174, Tyr175, Ser176, and Val178) and C-terminal positively charged residues Leu231, Leu237, Leu239, and Ser265.
The structural snapshots of the drug-protein complexes have been shown in Fig. 10. It has been seen during simulation that dextromethorphan has drifted to the different locations inside the NSP6 during ~ 20–100 ns of simulation. In the first ∼ 20 ns, dextromethorphan stayed in its initial location (Fig. 10a). After that, it drifted away from its initial location and moved into the water above the binding pocket and the interaction becomes minimal, only with Tyr234, Phe235, and Leu276 (Fig. 10b). Without drifting further into the water, the dextromethorphan molecule re-entered to the enlarged binding pocket, where it interacts mainly with H12 residues (Phe225, Leu230, Arg233, Tyr234, and Arg236) through the hydrophobic interaction (Fig. 10c).
During the second half of the MD simulation (~ 50–100 ns), the dextromethorphan molecule stably remained at a new binding pose with much stronger interactions. At this new location, dextromethorphan coordinates with Ser32 through H-bonds, and hydrophobic interactions by Trp31, Ile189, Val190, Met192, Cys193, Phe200, and Phe201 (H9-H10) (Fig. 10d).
In contrast, the drug haloperidol was stably bound inside the NSP6 pocket and explored two binding poses during the simulation. For the first 50 ns, it remained at its initial docking pose where it forms H-bonds to Ser 176 and Ser265, and several hydrophobic interactions mediated by Arg137, Trp140, Asn174, Tyr175, Val178, and Thr238 (Fig. 10e, f). After that, it drifted to another binding site and remained stable there, until the end of the simulation. At this new site, haloperidol interacts mainly with H7-H9 residues (Trp140, Asn144, Trp165, Ala166, Ile169, Ser170, Ser176, Val178, and Phe184), Leu237, Leu239, and Ile266, Phe269 (H14) (Fig. 10g, h).
Conclusion
To better understand the mechanism of biased activity of SARS-CoV-2 in the presence of two sigma-R1 binding drugs to NSP6, molecular dynamics simulation studies were employed. Our data suggests dextromethorphan binds to C-terminal helices while haloperidol binding sites are in the middle of the protein domain in helices H7 and H9. The disruption of the alpha helix H7 and H8 is significant for the dextromethorphan complex along with a large kink in H5 and H7. The NSP6-haloperidol complex showed a less significant change in the tertiary structure despite major disruption of the H12 helix.
Furthermore, the analyses of RMSD, RMSF, PCA, FEL, and dynamic cross-correlation matrix (DCCM) indicated that dextromethorphan binding leads to destabilization of the protein with the loss of correlated motions and residual frustrations. In contrast, haloperidol binding brings milder alterations in these order parameters and thus showed minimal changes in stability compared with dextromethorphan system. Besides, intermolecular hydrogen bonds were constantly formed in haloperidol with high occupation indicating the more stabilization of the NSP6-haloperidol system. In conclusion, the study elucidated the detailed interaction mechanism of dextromethorphan and haloperidol to NSP6 protein and the associated structural and dynamical changes upon drug binding. These results will significantly enhance our understanding of the working mode of these drugs at the molecular and structural level and will contribute to the future rational drug design for COVID-19.
References
Chen N, Zhou M, Dong X, Qu J, Gong F, Han Y, Qiu Y, Wang J, Liu Y, Wei Y et al (2020) Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet 395:507–513
Hossein-Khannazer N, Shokoohian B, Shpichka A, Aghdaei HA, Timashev P, Vosough M (2020) Novel therapeutic approaches for treatment of COVID-19. J Mol Med (Berl) 98:789–803
Touma M (2020) COVID-19: molecular diagnostics overview. J Mol Med (Berl) 98:947–954
Olwenyi OA, Dyavar SR, Acharya A, Podany AT, Fletcher CV, Ng CL, Reid SP, Byrareddy SN (2020) Immuno-epidemiology and pathophysiology of coronavirus disease 2019 (COVID-19). J Mol Med (Berl). https://doi.org/10.1007/s00109-020-01961-4
Comentale G, Manzo R, Pilato E (2020) Sars-Cov-2 interference in HEME production: is it the time for an early predictive biomarker? J Mol Med (Berl) 98:1053–1054
Khatoon F, Prasad K, Kumar V (2020) Neurological manifestations of COVID-19: available evidences and a new paradigm. J Neurovirol. https://doi.org/10.1007/s13365-020-00895-4
Kalita P, Padhi AK, Zhang KYJ, Tripathi T (2020) Design of a peptide-based subunit vaccine against novel coronavirus SARS-CoV-2. Microb Pathog 145:104236
Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, Zhao X, Huang B, Shi W, Lu R et al (2020) A novel coronavirus from patients with pneumonia in China, 2019. N Engl J Med 382:727–733
Cascella M, Rajnik M, Cuomo A, Dulebohn SC, Di Napoli R (2020) Features, evaluation, and treatment of coronavirus (COVID-19). DOI NBK554776 [bookaccession]
Barnard DL (2008) Coronaviruses: molecular and cellular biology. Futur Virol 3:119–123
van der Hoeven B, Oudshoorn D, Koster AJ, Snijder EJ, Kikkert M, Barcena M (2016) Biogenesis and architecture of arterivirus replication organelles. Virus Res 220:70–90
Zalinger ZB, Elliott R, Rose KM, Weiss SR (2015) MDA5 is critical to host defense during infection with murine coronavirus. J Virol 89:12330–12340
He Y, **e Z, Dai J, Cao Y, Hou J, Zheng Y, Wei T, Mo M, Wei P (2016) Responses of the Toll-like receptor and melanoma differentiation-associated protein 5 signaling pathways to avian infectious bronchitis virus infection in chicks. Virol Sin 31:57–68
Oudshoorn D, Rijs K, Limpens R, Groen K, Koster AJ, Snijder EJ, Kikkert M, Barcena M (2017). Expression and cleavage of middle east respiratory syndrome coronavirus nsp3-4 polyprotein induce the formation of double-membrane vesicles that mimic those associated with coronaviral RNA replication. mBio 8. DOI https://doi.org/10.1128/mBio.01658-17
Knoops K, Kikkert M, Worm SH, Zevenhoven-Dobbe JC, van der Meer Y, Koster AJ, Mommaas AM, Snijder EJ (2008) SARS-coronavirus replication is supported by a reticulovesicular network of modified endoplasmic reticulum. PLoS Biol 6:e226
Maier HJ, Hawes PC, Cottam EM, Mantell J, Verkade P, Monaghan P, Wileman T, Britton P (2013) Infectious bronchitis virus generates spherules from zippered endoplasmic reticulum membranes. mBio 4: e00801-e00813. DOI
Cottam EM, Maier HJ, Manifava M, Vaux LC, Chandra-Schoenfelder P, Gerner W, Britton P, Ktistakis NT, Wileman T (2011) Coronavirus nsp6 proteins generate autophagosomes from the endoplasmic reticulum via an omegasome intermediate. Autophagy 7:1335–1347
Cottam EM, Whelband MC, Wileman T (2014) Coronavirus NSP6 restricts autophagosome expansion. Autophagy 10:1426–1441
Gordon DE, Jang GM, Bouhaddou M, Xu J, Obernier K, White KM, O’Meara MJ, Rezelj VV, Guo JZ, Swaney DL et al (2020) A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature. https://doi.org/10.1038/s41586-020-2286-9
Tulgar S, Ahiskalioglu A, Kok A, Thomas DT (2020) Possible old drugs for repositioning in COVID-19 treatment: combating cytokine storms from haloperidol to anti-interleukin agents. Turk J Anaesthesiol Reanim 48:256–257
Lauterbach EC (2012) An extension of hypotheses regarding rapid-acting, treatment-refractory, and conventional antidepressant activity of dextromethorphan and dextrorphan. Med Hypotheses 78:693–702
Werling LL, Keller A, Frank JG, Nuwayhid SJ (2007) A comparison of the binding profiles of dextromethorphan, memantine, fluoxetine and amitriptyline: treatment of involuntary emotional expression disorder. Exp Neurol 207:248–257
Matsumoto RR, Nguyen L, Kaushal N, Robson MJ (2014) Sigma (sigma) receptors as potential therapeutic targets to mitigate psychostimulant effects. Adv Pharmacol 69:323–386
Fishback JA, Robson MJ, Xu YT, Matsumoto RR (2010) Sigma receptors: potential targets for a new class of antidepressant drug. Pharmacol Ther 127:271–282
Nguyen L, Robson MJ, Healy JR, Scandinaro AL, Matsumoto RR (2014) Involvement of sigma-1 receptors in the antidepressant-like effects of dextromethorphan. PLoS One 9:e89985
Prasad K, Khatoon F, Rashid S, Ali N, AlAsmari AF, Ahmed MZ, Alqahtani AS, Alqahtani MS, Kumar V (2020) Targeting hub genes and pathways of innate immune response in COVID-19: a network biology perspective. Int J Biol Macromol. https://doi.org/10.1016/j.ijbiomac.2020.06.228
Enkirch T, Sauber S, Anderson DE, Gan ES, Kenanov D, Maurer-Stroh S, von Messling V (2019) Identification and in vivo efficacy assessment of approved orally bioavailable human host protein-targeting drugs with broad anti-influenza A activity. Front Immunol 10:1097
Senior AW, Evans R, Jumper J, Kirkpatrick J, Sifre L, Green T, Qin C, Zidek A, Nelson AWR, Bridgland A et al (2020) Improved protein structure prediction using potentials from deep learning. Nature 577:706–710
Zhang C, Zheng W, Huang X, Bell EW, Zhou X, Zhang Y (2020) Protein structure and sequence reanalysis of 2019-nCoV genome refutes snakes as its intermediate host and the unique similarity between its spike protein insertions and HIV-1. J Proteome Res 19:1351–1360
Zheng W, Li Y, Zhang C, Pearce R, Mortuza SM, Zhang Y (2019) Deep-learning contact-map guided protein structure prediction in CASP13. Proteins 87:1149–1164
Sterling T, Irwin JJ (2015) ZINC 15--ligand discovery for everyone. J Chem Inf Model 55:2324–2337
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comput Chem 31:455–461
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791
Seeliger D, de Groot BL (2010) Ligand docking and binding site analysis with PyMOL and Autodock/Vina. J Comput Aided Mol Des 24:417–422
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera--a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Laskowski RA, Swindells MB (2011) LigPlot+: multiple ligand-protein interaction diagrams for drug discovery. J Chem Inf Model 51:2778–2786
Abraham MJ, Murtola T, Schulz R, Páll S, Smith JC, Hess B, Lindahl E (2015) GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1-2:19–25
Zoete V, Cuendet MA, Grosdidier A, Michielin O (2011) SwissParam: a fast force field generation tool for small organic molecules. J Comput Chem 32:2359–2368
Joung IS, Cheatham TE 3rd (2008) Determination of alkali and halide monovalent ion parameters for use in explicitly solvated biomolecular simulations. J Phys Chem B 112:9020–9041
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N·log(N) method for Ewald sums in large systems. J Chem Phys 98:10089–10092
Berendsen HJC, Grigera JR, Straatsma TP (1987) The missing term in effective pair potentials. J Phys Chem 91:6269–6271
Parrinello M, Rahman A (1980) Crystal structure and pair potentials: a molecular-dynamics study. Phys Rev Lett 45:1196–1199
Berk H, Henk B, C. BHJ, M. FJGE (1997) LINCS: a linear constraint solver for molecular simulations. J Comput Chem 18: 1463-1472. DOI:
Kumar V, Pandey P, Idrees D, Prakash A, Lynn AM (2019) Delineating the effect of mutations on the conformational dynamics of N-terminal domain of TDP-43. Biophys Chem 250:106174
Prakash A, Kumar V, Banerjee A, Lynn AM, Prasad R (2020) Structural heterogeneity in RNA recognition motif 2 (RRM2) of TAR DNA-binding protein 43 (TDP-43): clue to amyotrophic lateral sclerosis. J Biomol Struct Dyn:1–11. https://doi.org/10.1080/07391102.2020.1714481
McGibbon RT, Beauchamp KA, Harrigan MP, Klein C, Swails JM, Hernandez CX, Schwantes CR, Wang LP, Lane TJ, Pande VS (2015) MDTraj: a modern open library for the analysis of molecular dynamics trajectories. Biophys J 109:1528–1532
Kabsch W, Sander C (1983) Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22:2577–2637
Kumar N, Srivastava R, Prakash A, Lynn AM (2019) Structure-based virtual screening, molecular dynamics simulation and MM-PBSA toward identifying the inhibitors for two-component regulatory system protein NarL of Mycobacterium Tuberculosis. J Biomol Struct Dyn:1–15. https://doi.org/10.1080/07391102.2019.1657499
Pandey P, Lynn AM, Bandyopadhyay P (2017) Identification of inhibitors against alpha-isopropylmalate synthase of mycobacterium tuberculosis using docking-MM/PBSA hybrid approach. Bioinformation 13:144–148
Ferreiro DU, Hegler JA, Komives EA, Wolynes PG (2007) Localizing frustration in native proteins and protein assemblies. Proc Natl Acad Sci U S A 104:19819–19824
Parra RG, Schafer NP, Radusky LG, Tsai MY, Guzovsky AB, Wolynes PG, Ferreiro DU (2016) Protein Frustratometer 2: a tool to localize energetic frustration in protein molecules, now with electrostatics. Nucleic Acids Res 44:W356–W360
Husain S, Kumar V, Hassan MI (2018) Phosphorylation-induced changes in the energetic frustration in human Tank binding kinase 1. J Theor Biol 449:14–22
Tian W, Chen C, Lei X, Zhao J, Liang J (2018) CASTp 3.0: computed atlas of surface topography of proteins. Nucleic Acids Res 46:W363–W367
Jendele L, Krivak R, Skoda P, Novotny M, Hoksza D (2019) PrankWeb: a web server for ligand binding site prediction and visualization. Nucleic Acids Res 47:W345–W349
Yang J, Roy A, Zhang Y (2013) Protein-ligand binding site recognition using complementary binding-specific substructure comparison and sequence profile alignment. Bioinformatics 29:2588–2595
Kollman PA, Massova I, Reyes C, Kuhn B, Huo S, Chong L, Lee M, Lee T, Duan Y, Wang W et al (2000) Calculating structures and free energies of complex molecules: combining molecular mechanics and continuum models. Acc Chem Res 33:889–897
Kumari R, Kumar R, Lynn A (2014) g_mmpbsa--a GROMACS tool for high-throughput MM-PBSA calculations. J Chem Inf Model 54:1951–1962
Chen J, Wang J, Zhu W, Li G (2013) A computational analysis of binding modes and conformation changes of MDM2 induced by p53 and inhibitor bindings. J Comput Aided Mol Des 27:965–974
Chen J, Liang Z, Wang W, Yi C, Zhang S, Zhang Q (2014) Revealing origin of decrease in potency of darunavir and amprenavir against HIV-2 relative to HIV-1 protease by molecular dynamics simulations. Sci Rep 4:6872
Rodrigues CH, Pires DE, Ascher DB (2018) DynaMut: predicting the impact of mutations on protein conformation, flexibility and stability. Nucleic Acids Res 46:W350–W355
Acknowledgments
The authors sincerely thank the Amity University, Noida, for providing facilities. Authors gratefully acknowledge the computational facility funded by Science and Engineering Research Board (SERB), Government of India (Ref. No.: YSS/2015/000228/LS).
Author information
Authors and Affiliations
Contributions
V.K. designed the study. P.P. and K.P. performed the experiments and calculations. V.K. and A.P. analyzed the data. P.P. and K.P. prepared figures of the results. V.K. wrote the manuscript with the contributions of K.P. and A.P.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Pandey, P., Prasad, K., Prakash, A. et al. Insights into the biased activity of dextromethorphan and haloperidol towards SARS-CoV-2 NSP6: in silico binding mechanistic analysis. J Mol Med 98, 1659–1673 (2020). https://doi.org/10.1007/s00109-020-01980-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-020-01980-1