Search
Search Results
-
Comparative evaluation of SNVs, indels, and structural variations detected with short- and long-read sequencing data
Short- and long-read sequencing technologies are routinely used to detect DNA variants, including SNVs, indels, and structural variations (SVs)....

-
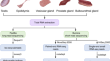
Long- and short-read RNA sequencing from five reproductive organs of boar
The production of semen in boars involves multiple reproductive glands, including the testis (Tes), epididymis (Epi), vesicular gland (VG), prostate...
-
INSurVeyor: improving insertion calling from short read sequencing data
Insertions are one of the major types of structural variations and are defined as the addition of 50 nucleotides or more into a DNA sequence. Several...

-
Comparing genomes recovered from time-series metagenomes using long- and short-read sequencing technologies
BackgroundOver the past years, sequencing technologies have expanded our ability to examine novel microbial metabolisms and diversity previously...

-
Short-Read RNA-Seq
RNA sequencing (RNA-Seq) has emerged as a powerful and versatile tool for the comprehensive analysis of transcriptomes and has been widely used to...
-
High diagnostic potential of short and long read genome sequencing with transcriptome analysis in exome-negative developmental disorders
Exome sequencing (ES) has become the method of choice for diagnosing rare diseases, while the availability of short-read genome sequencing (SR-GS) in...

-
Linked-read sequencing for detecting short tandem repeat expansions
Detection of short tandem repeat (STR) expansions with standard short-read sequencing is challenging due to the difficulty in map** multicopy...

-
MetaCC allows scalable and integrative analyses of both long-read and short-read metagenomic Hi-C data
Metagenomic Hi-C (metaHi-C) can identify contig-to-contig relationships with respect to their proximity within the same physical cell. Shotgun...

-
Assembly and Annotation of Viral Metagenomes from Short-Read Sequencing Data
Viral metagenomics enables the detection, characterization, and quantification of viral sequences present in shotgun-sequenced datasets of purified...
-
Oxford Nanopore R10.4 long-read sequencing enables the generation of near-finished bacterial genomes from pure cultures and metagenomes without short-read or reference polishing
Long-read Oxford Nanopore sequencing has democratized microbial genome sequencing and enables the recovery of highly contiguous microbial genomes...

-
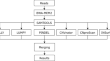
ProcaryaSV: structural variation detection pipeline for bacterial genomes using short-read sequencing
BackgroundStructural variations play an important role in bacterial genomes. They can mediate genome adaptation quickly in response to the external...
-
Whole-genome sequencing of Ganoderma boninense, the causal agent of basal stem rot disease in oil palm, via combined short- and long-read sequencing
The hemibiotrophic Basidiomycete pathogen Ganoderma boninense ( Gb ) is the dominant causal agent of oil palm basal stem rot disease. Here, we report a...

-

-
Short-read full-length 16S rRNA amplicon sequencing for characterisation of the respiratory bacteriome of captive and free-ranging African elephants (Loxodonta africana)
Hypervariable region sequencing of the 16S ribosomal RNA (rRNA) gene plays a critical role in microbial ecology by offering insights into bacterial...

-
Readsynth: short-read simulation for consideration of composition-biases in reduced metagenome sequencing approaches
BackgroundThe application of reduced metagenomic sequencing approaches holds promise as a middle ground between targeted amplicon sequencing and...

-
Short-read and long-read RNA sequencing of mouse hematopoietic stem cells at bulk and single-cell levels
Hematopoietic stem cells (HSCs) lie at the top of the differentiation hierarchy. Although HSC and their immediate downstream, multipotent progenitors...

-
Comparative Analysis of Structural Variant Callers on Short-Read Whole-Genome Sequencing Data
AbstractThree structural variant callers (Manta, Smoove, Delly) are analyzed on whole-genome sequencing data using four different alignment...

-
Performance evaluation of six popular short-read simulators
High-throughput sequencing data enables the comprehensive study of genomes and the variation therein. Essential for the interpretation of this...

-
Detecting Tandem Repeat Expansions Using Short-Read Sequencing for Clinical Use
Repeat expansion disorders are a unique class of genetic diseases caused by expansions of short tandem repeats. Until recently, these pathogenic...
-
Integration of Hi-C with short and long-read genome sequencing reveals the structure of germline rearranged genomes
Structural variants are a common cause of disease and contribute to a large extent to inter-individual variability, but their detection and...

