Search
Search Results
-
Eukaryotic-driven directed evolution of Cas9 nucleases
BackgroundFurther advancement of genome editing highly depends on the development of tools with higher compatibility with eukaryotes. A multitude of...

-
Construction and Evaluation of Zinc Finger Nucleases
Zinc finger nucleases (ZFNs) are programmable nucleases that have contributed significantly to past genome-editing research. They are now utilized...
-
Background: Genome Editing with Programmable Nucleases
Scientists use genome editing with programmable nucleases to carry out the targeted modification of nucleic acid. The major genome editing...
-
Type II CRISPR–Cas System Nucleases: A Pipeline for Prediction and In Vitro Characterization
AbstractThe use of CRISPR–Cas bacterial adaptive immunity system components for targeted DNA changes has opened broad prospects for programmable...

-
Nucleases are upregulated in potato tubers afflicted with zebra chip disease
Main ConclusionThe defense response of potato tubers afflicted with zebra chip disease involves oxidatively mediated upregulation of nucleases that...

-
Structure and Function of SNM1 Family Nucleases
Three human nucleases, SNM1A, SNM1B/Apollo, and SNM1C/Artemis, belong to the SNM1 gene family. These nucleases are involved in various cellular...
-
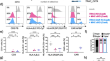
Combining different CRISPR nucleases for simultaneous knock-in and base editing prevents translocations in multiplex-edited CAR T cells
BackgroundMultiple genetic modifications may be required to develop potent off-the-shelf chimeric antigen receptor (CAR) T cell therapies....

-
Rapidly Characterizing CRISPR-Cas13 Nucleases Using Cell-Free Transcription-Translation Systems
Cell-free transcription-translation (TXTL) systems produce RNAs and proteins from added DNA. By coupling their production to a biochemical assay,...
-

-
Evolutionary mining and functional characterization of TnpB nucleases identify efficient miniature genome editors
As the evolutionary ancestor of Cas12 nuclease, the transposon (IS200/IS605)-encoded TnpB proteins act as compact RNA-guided DNA endonucleases. To...

-
Enhanced mitochondrial DNA editing in mice using nuclear-exported TALE-linked deaminases and nucleases
We present two methods for enhancing the efficiency of mitochondrial DNA (mtDNA) editing in mice with DddA-derived cytosine base editors (DdCBEs)....

-
Tag-seq: a convenient and scalable method for genome-wide specificity assessment of CRISPR/Cas nucleases
Genome-wide identification of DNA double-strand breaks (DSBs) induced by CRISPR-associated protein (Cas) systems is vital for profiling the...

-
Homology-based repair induced by CRISPR-Cas nucleases in mammalian embryo genome editing
Recent advances in genome editing, especially CRISPR-Cas nucleases, have revolutionized both laboratory research and clinical therapeutics....

-
Tools for experimental and computational analyses of off-target editing by programmable nucleases
Genome editing using programmable nucleases is revolutionizing life science and medicine. Off-target editing by these nucleases remains a...

-
Identifying genome-wide off-target sites of CRISPR RNA–guided nucleases and deaminases with Digenome-seq
Digested genome sequencing (Digenome-seq) is a highly sensitive, easy-to-carry-out, cell-free method for experimentally identifying genome-wide...

-
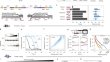
Massively parallel kinetic profiling of natural and engineered CRISPR nucleases
Engineered Sp Cas9s and As Cas12a cleave fewer off-target genomic sites than wild-type (wt) Cas9. However, understanding their fidelity, mechanisms and...
-
Genome editing with CRISPR–Cas nucleases, base editors, transposases and prime editors
The development of new CRISPR–Cas genome editing tools continues to drive major advances in the life sciences. Four classes of CRISPR–Cas-derived...

-
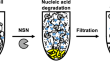
Bacterial non-specific nucleases of the phospholipase D superfamily and their biotechnological potential
Bacterial non-specific nucleases are ubiquitously distributed and involved in numerous intra- and extracellular processes. Although all nucleases...
-
Efficient genome editing in wheat using Cas9 and Cpf1 (AsCpf1 and LbCpf1) nucleases
Genome editing can be used to create new wheat varieties with enhanced performance. Clustered regularly interspaced short palindromic repeat (CRISPR)...

-
Genome editing in plants using the TnpB transposase system
The widely used clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated nuclease (Cas) system is thought to have evolved...

