Search
Search Results
-
Transposition Distance Considering Intergenic Regions for Unbalanced Genomes
In seminal works of genome rearrangements, the distance between two genomes is measured by the minimum number of rearrangements (e.g., reversals,...
-
An improved approximation algorithm for the reversal and transposition distance considering gene order and intergenic sizes
BackgroundIn the comparative genomics field, one of the goals is to estimate a sequence of genetic changes capable of transforming a genome into...

-
Spatially resolved epigenome sequencing via Tn5 transposition and deterministic DNA barcoding in tissue
Spatial epigenetic map** of tissues enables the study of gene regulation programs and cellular functions with the dependency on their local tissue...

-
Block Interchange and Reversal Distance on Unbalanced Genomes
One method for inferring the evolutionary distance between two organisms is to find the rearrangement distance, which is defined as the minimum...
-
Approximation algorithm for rearrangement distances considering repeated genes and intergenic regions
The rearrangement distance is a method to compare genomes of different species. Such distance is the number of rearrangement events necessary to...

-
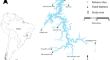
Beyond just a dam blockage problem: larger artificial reservoirs are additional obstacles to reproductive fish migration in the Neotropics
One of the most conspicuous impacts of dam construction on fish is the blocking of their migratory routes. However, the formation of the reservoir, a...
-

-
From parasites to partners: exploring the intricacies of host-transposon dynamics and coevolution
Transposable elements, often referred to as “jum** genes,” have long been recognized as genomic parasites due to their ability to integrate and...

-
First Biochemical Steps on Bacterial Transposition Pathways
Transposons are found in a wide variety of forms throughout the prokaryotic world where they actively contribute to the adaptive strategies of...
-
A 1.375-Approximation Algorithm for Sorting by Transpositions with Faster Running Time
Sorting Permutations by Transpositions is a famous problem in the Computational Biology field. This problem is NP-Hard, and the best approximation...
-
String correction using the Damerau-Levenshtein distance
BackgroundIn the string correction problem, we are to transform one string into another using a set of prescribed edit operations. In string...

-
Approximating Rearrangement Distances with Replicas and Flexible Intergenic Regions
Many tools from Computational Biology compute distances between genomes by counting the number of genome rearrangement events, such as reversals of a...
-
On Computing the Jaro Similarity Between Two Strings
Jaro similarity is widely used in computing the similarity (or distance) between two strings of characters. For example, record linkage is an...
-
BioByGANS: biomedical named entity recognition by fusing contextual and syntactic features through graph attention network in node classification framework
BackgroundAutomatic and accurate recognition of various biomedical named entities from literature is an important task of biomedical text mining,...

-
Influence of species invasion, seasonality, and connectivity on fish functional and taxonomic beta-diversity in a Neotropical floodplain
Studies that combine functional and taxonomic beta-diversity are essential for explaining some ecological processes, including the process of species...

-
Retrotransposable Elements: DNA Fingerprinting and the Assessment of Genetic Diversity
Retrotransposable elements (RTEs) are highly common mobile genetic elements that are composed of several classes and make up the majority of...
-
Heuristics for Cycle Packing of Adjacency Graphs for Genomes with Repeated Genes
The Adjacency Graph is a structure used to model genomes in several rearrangement distance problems. In particular, most studies use properties of a...
-
Diversity and evolution of transposable elements in the plant-parasitic nematodes
BackgroundTransposable elements (TEs) are mobile DNA sequences that propagate within genomes, occupying a significant portion of eukaryotic genomes...

-
Transposable Elements II: Insertional Mutagenesis
This chapter is a sequel to the topic of transposable elements discussed in Chap. 5 . Here, we will...
-
Genome Rearrangement Analysis
Genome rearrangements are mutations that change the gene content of a genome or the arrangement of the genes on a genome. Several years of research...
