Search
Search Results
-
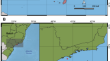
Unlocking the jar: revealing gastric content in Ceriantharia (Cnidaria, Anthozoa) through whole-genome shotgun sequencing
This study focuses on the analyses of the gastral cavity contents of two species of Ceriantharia, namely Isarachnanthus nocturnus Hartog, 1977,...
-
Microbiome signatures associated with clinical stages of gastric Cancer: whole metagenome shotgun sequencing study
BackgroundGastric cancer is one of the global health concerns. A series of studies on the stomach have confirmed the role of the microbiome in...

-
Shotgun metagenomics and computational profiling of the plastisphere microbiome: unveiling the potential of enzymatic production and plastic degradation
Plastic pollution is one of the most resilient types of pollution and is considered a global environmental threat, particularly in the marine...

-
Unlocking gut microbiota potential of dairy cows in varied environmental conditions using shotgun metagenomic approach
Food security and environmental pollution are major concerns for the expanding world population, where farm animals are the largest source of dietary...

-
Unveiling the microbiome during post-partum uterine infection: a deep shotgun sequencing approach to characterize the dairy cow uterine microbiome
BackgroundThe goal of this study was to assess the microbial ecology and diversity present in the uterus of post-partum dairy cows with and without...

-
Unveiling the microbial diversity and functional dynamics of Shiv Kund, Sohna hot spring, India through a shotgun metagenomics approach
In this research, we examined the microbial diversity in Sohna hot spring, Haryana, India using shotgun metagenome sequencing based on the Illumina...

-
Toward efficient and high-fidelity metagenomic data from sub-nanogram DNA: evaluation of library preparation and decontamination methods
BackgroundShotgun metagenomic sequencing has greatly expanded the understanding of microbial communities in various biological niches. However, it is...

-
Simple, efficient and thorough shotgun proteomic analysis with PatternLab V
Shotgun proteomics aims to identify and quantify the thousands of proteins in complex mixtures such as cell and tissue lysates and biological fluids....

-
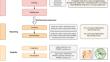
Wochenende — modular and flexible alignment-based shotgun metagenome analysis
BackgroundShotgun metagenome analysis provides a robust and verifiable method for comprehensive microbiome analysis of fungal, viral, archaeal and...
-
HOME-BIO (sHOtgun MEtagenomic analysis of BIOlogical entities): a specific and comprehensive pipeline for metagenomic shotgun sequencing data analysis
BackgroundNext-Generation-Sequencing (NGS) enables detection of microorganisms present in biological and other matrices of various origin and nature,...

-
MicroPredict: predicting species-level taxonomic abundance of whole-shotgun metagenomic data using only 16S amplicon sequencing data
BackgroundThe importance of the human microbiome in the analysis of various diseases is emerging. The two main methods used to profile the human...

-
Microbiota and environmental health monitoring of mouse colonies by metagenomic shotgun sequencing
Metagenomic next-generation sequencing (mNGS) allows the monitoring of microbiota composition of murine colonies employed for scientific purposes in...

-
Evaluation of taxonomic classification and profiling methods for long-read shotgun metagenomic sequencing datasets
BackgroundLong-read shotgun metagenomic sequencing is gaining in popularity and offers many advantages over short-read sequencing. The higher...

-
Paleo-diatom composition from Santa Barbara Basin deep-sea sediments: a comparison of 18S-V9 and diat-rbcL metabarcoding vs shotgun metagenomics
Sedimentary ancient DNA ( sed aDNA) analyses are increasingly used to reconstruct marine ecosystems. The majority of marine sed aDNA studies use a...

-
MaxDIA enables library-based and library-free data-independent acquisition proteomics
MaxDIA is a software platform for analyzing data-independent acquisition (DIA) proteomics data within the MaxQuant software environment. Using...

-
Host DNA Depletion in Saliva Samples for Improved Shotgun Metagenomics
Host DNA makes up the majority of DNA in a saliva sample. Therefore, shotgun metagenomics can be an inefficient way to evaluate the microbial...
-
Recovering prokaryotic genomes from host-associated, short-read shotgun metagenomic sequencing data
Recovering genomes from shotgun metagenomic sequence data allows detailed taxonomic and functional characterization of individual species or strains...

-
Comparison of 6 DNA extraction methods for isolation of high yield of high molecular weight DNA suitable for shotgun metagenomics Nanopore sequencing to detect bacteria
BackgroundOxford Nanopore Technologies (ONT) offers an accessible platform for long-read sequencing, which improves the reconstruction of genomes and...

-
Impact of crop** systems on the functional diversity of rhizosphere microbial communities associated with maize plant: a shotgun approach
Understanding the functions carried out by rhizosphere microbiomes will further explore their importance in biotechnological improvement and...

-
Qualitative and Quantitative Shotgun Proteomics Data Analysis from Data-Dependent Acquisition Mass Spectrometry
Shotgun proteomics is the inferential analysis of proteoforms using peptide proxies produced by enzyme-catalyzed hydrolysis of entire proteomes. Such...
