Search
Search Results
-
A survey of map** algorithms in the long-reads era
It has been over a decade since the first publication of a method dedicated entirely to map** long-reads. The distinctive characteristics of long...

-
Retained introns in long RNA-seq reads are not reliably detected in sample-matched short reads
BackgroundThere is growing interest in retained introns in a variety of disease contexts including cancer and aging. Many software tools have been...

-
NextDenovo: an efficient error correction and accurate assembly tool for noisy long reads
Long-read sequencing data, particularly those derived from the Oxford Nanopore sequencing platform, tend to exhibit high error rates. Here, we...

-
High-quality metagenome assembly from long accurate reads with metaMDBG
We introduce metaMDBG, a metagenomics assembler for PacBio HiFi reads. MetaMDBG combines a de Bruijn graph assembly in a minimizer space with an...

-
Hybrid-hybrid correction of errors in long reads with HERO
Although generally superior, hybrid approaches for correcting errors in third-generation sequencing (TGS) reads, using next-generation sequencing...

-
Highly accurate long reads are crucial for realizing the potential of biodiversity genomics
BackgroundGenerating the most contiguous, accurate genome assemblies given available sequencing technologies is a long-standing challenge in genome...

-
CoLoRd: compressing long reads
The cost of maintaining exabytes of data produced by sequencing experiments every year has become a major issue in today’s genomic research. In spite...

-
Metagenome assembly of high-fidelity long reads with hifiasm-meta
De novo assembly of metagenome samples is a common approach to the study of microbial communities. Current metagenome assemblers developed for short...

-
Binning long reads in metagenomics datasets using composition and coverage information
BackgroundAdvancements in metagenomics sequencing allow the study of microbial communities directly from their environments. Metagenomics binning is...

-
Performance analysis of conventional and AI-based variant callers using short and long reads
BackgroundThe accurate detection of variants is essential for genomics-based studies. Currently, there are various tools designed to detect genomic...

-
Detecting haplotype-specific transcript variation in long reads with FLAIR2
BackgroundRNA-seq has brought forth significant discoveries regarding aberrations in RNA processing, implicating these RNA variants in a variety of...

-
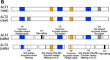
Benchmarking datasets for assembly-based variant calling using high-fidelity long reads
BackgroundRecent advances in long-read sequencing technologies have enabled accurate identification of all genetic variants in individuals or cells;...
-
Accurate isoform discovery with IsoQuant using long reads
Annotating newly sequenced genomes and determining alternative isoforms from long-read RNA data are complex and incompletely solved problems. Here we...

-
Multiplex de Bruijn graphs enable genome assembly from long, high-fidelity reads
Although most existing genome assemblers are based on de Bruijn graphs, the construction of these graphs for large genomes and large k -mer sizes has...

-
Characterization of telomere variant repeats using long reads enables allele-specific telomere length estimation
Telomeres are regions of repetitive DNA at the ends of linear chromosomes which protect chromosome ends from degradation. Telomere lengths have been...

-
SVDSS: structural variation discovery in hard-to-call genomic regions using sample-specific strings from accurate long reads
Structural variants (SVs) account for a large amount of sequence variability across genomes and play an important role in human genomics and...

-
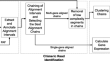
Genion, an accurate tool to detect gene fusion from long transcriptomics reads
BackgroundThe advent of next-generation sequencing technologies empowered a wide variety of transcriptomics studies. A widely studied topic is gene...
-
KFinger: Capturing Overlaps Between Long Reads by Using Lyndon Fingerprints
Detecting common regions and overlaps between DNA sequences is crucial in many Bioinformatics tasks. One of them is genome assembly based on the use...
-
NPGREAT: assembly of human subtelomere regions with the use of ultralong nanopore reads and linked-reads
BackgroundHuman subtelomeric DNA regulates the length and stability of adjacent telomeres that are critical for cellular function, and contains many...

-
BreakNet: detecting deletions using long reads and a deep learning approach
BackgroundStructural variations (SVs) occupy a prominent position in human genetic diversity, and deletions form an important type of SV that has...

