Search
Search Results
-
Specific DNMT3C flanking sequence preferences facilitate methylation of young murine retrotransposons
The DNA methyltransferase DNMT3C appeared as a duplication of the DNMT3B gene in muroids and is required for silencing of young retrotransposons in...

-
Flanking sequences influence the activity of TET1 and TET2 methylcytosine dioxygenases and affect genomic 5hmC patterns
TET dioxygenases convert 5-methylcytosine (5mC) preferentially in a CpG context into 5-hydroxymethylcytosine (5hmC) and higher oxidized forms,...

-
Interconnected Codons: Unravelling the Epigenetic Significance of Flanking Sequences in CpG Dyads
Hypothesizing that CpG codon dyads, formed by consecutive codons containing a cytosine-guanine pair (NNC-GNN), may play a crucial role in gene...

-
DAR-PCR: a new tool for efficient retrieval of unknown flanking genomic DNA
Various PCR-based genome-walking methods have been developed to acquire unknown flanking DNA sequences. However, the specificity and efficacy levels,...

-
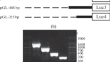
Structural Analysis of the 5'-Flanking Region of Human Alpha-Fetoprotein Encoding Gene
AbstractAlpha fetoprotein (AFP) is a growth factor and a signaling molecule that promotes the development of HCC. However, the mechanism of the...
-
Deep embedding and alignment of protein sequences
Protein sequence alignment is a key component of most bioinformatics pipelines to study the structures and functions of proteins. Aligning highly...

-
ISOGlyP: O-Glycosylation Site Prediction Using Peptide Sequences and GALNTs
Mucin-type O-glycosylation is one of the most common posttranslational modifications of proteins. The abnormal expression of various polypeptide...
-
Deep learning models incorporating endogenous factors beyond DNA sequences improve the prediction accuracy of base editing outcomes
Adenine base editors (ABEs) and cytosine base editors (CBEs) enable the single nucleotide editing of targeted DNA sites avoiding generation of double...

-
Efficient identification of genomic insertions and flanking regions through whole-genome sequencing in three transgenic soybean events
Genomic insertions and flanking regions of transgenes in host genomes constitute a critical component of precise molecular characterization and...

-
An integrated strategy for target SSR genoty** with toleration of nucleotide variations in the SSRs and flanking regions
BackgroundWith the broad application of high-throughput sequencing and its reduced cost, simple sequence repeat (SSR) genoty** by sequencing...

-
G4Bank: A database of experimentally identified DNA G-quadruplex sequences
G-quadruplex (G4), a non-canonical nucleic acid structure, has been suggested to play a key role in important cellular processes including...

-
Secretion of the fungal toxin candidalysin is dependent on conserved precursor peptide sequences
The opportunistic fungal pathogen Candida albicans damages host cells via its peptide toxin, candidalysin. Before secretion, candidalysin is embedded...

-
Detecting gene breakpoints in noisy genome sequences using position-annotated colored de-Bruijn graphs
BackgroundIdentifying the locations of gene breakpoints between species of different taxonomic groups can provide useful insights into the underlying...

-
Mitochondrial genome plasticity of mammalian species
There is an ongoing process in which mitochondrial sequences are being integrated into the nuclear genome. The importance of these sequences has...

-

-
Reporter gene assays and chromatin-level assays define substantially non-overlap** sets of enhancer sequences
BackgroundTranscriptional enhancers are essential for gene regulation, but how these regulatory elements are best defined remains a significant...

-
Four classic “de novo” genes all have plausible homologs and likely evolved from retro-duplicated or pseudogenic sequences
Despite being previously regarded as extremely unlikely, the idea that entirely novel protein-coding genes can emerge from non-coding sequences has...

-
Intronic small nucleolar RNAs regulate host gene splicing through base pairing with their adjacent intronic sequences
BackgroundSmall nucleolar RNAs (snoRNAs) are abundant noncoding RNAs best known for their involvement in ribosomal RNA maturation. In mammals, most...

-
Microsatellite Markers from Whole Genome and Transcriptomic Sequences
Microsatellites (MS) or simple sequence repeats (SSRs) is a DNA sequence set comprising of tandemly repeated motifs. SSRs with codominant...
-
Principles of Identification of Nucleotide Sequences in ISSR Marker Spectra
Abstract —An algorithm for theoretical prediction of nucleotide sequences obtained in ISSR-PCR is developed using a sequenced horse genome as an...

