Search
Search Results
-
Base Editing and Prime Editing
The development of new adaptations of CRISPR-based genome editing platforms, such as base editing and prime editing, made it possible to broaden the...
-
Base editing of organellar DNA with programmable deaminases
Mitochondria and chloroplasts are organelles that include their own genomes, which encode key genes for ATP production and carbon dioxide fixation,...

-
Precision RNA base editing with engineered and endogenous effectors
RNA base editing refers to the rewriting of genetic information within an intact RNA molecule and serves various functions, such as evasion of the...

-
Strand-preferred base editing of organellar and nuclear genomes using CyDENT
Transcription-activator-like effector (TALE)-based tools for base editing of nuclear and organellar DNA rely on double-stranded DNA deaminases, which...

-
Cytosine Base Editing in Bacteria
Base editing is a new genome editing technology that enables DNA base mutations without requiring double-stranded DNA backbone cleavage or a donor...
-
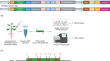
Cytosine base editors (CBEs) for inducing targeted DNA base editing in Nicotiana benthamiana
BackgroundThe base editors can introduce point mutations accurately without causing double-stranded DNA breaks or requiring donor DNA templates....
-
Strand-selective base editing of human mitochondrial DNA using mitoBEs
A number of mitochondrial diseases in humans are caused by point mutations that could be corrected by base editors, but delivery of CRISPR guide RNAs...

-
Multiplexed base editing through Cas12a variant-mediated cytosine and adenine base editors
Cas12a can process multiple sgRNAs from a single transcript of CRISPR array, conferring advantages in multiplexed base editing when incorporated into...

-
Deep learning models incorporating endogenous factors beyond DNA sequences improve the prediction accuracy of base editing outcomes
Adenine base editors (ABEs) and cytosine base editors (CBEs) enable the single nucleotide editing of targeted DNA sites avoiding generation of double...

-
Generation of precision preclinical cancer models using regulated in vivo base editing
Although single-nucleotide variants (SNVs) make up the majority of cancer-associated genetic changes and have been comprehensively catalogued, little...

-
Programmable A-to-Y base editing by fusing an adenine base editor with an N-methylpurine DNA glycosylase
Here we developed an adenine transversion base editor, AYBE, for A-to-C and A-to-T transversion editing in mammalian cells by fusing an adenine base...

-
Targeted C•G-to-T•A base editing with TALE-cytosine deaminases in plants
BackgroundTALE-derived DddA-based cytosine base editors (TALE-DdCBEs) can perform efficient base editing of mitochondria and chloroplast genomes....

-
Glycosylase-based base editors for efficient T-to-G and C-to-G editing in mammalian cells
Base editors show promise for treating human genetic diseases, but most current systems use deaminases, which cause off-target effects and are...

-
Fusion of a rice endogenous N-methylpurine DNA glycosylase to a plant adenine base transition editor ABE8e enables A-to-K base editing in rice plants
Engineering of a new type of plant base editor for simultaneous adenine transition and transversion within the editing window will greatly expand the...

-
Genotoxic effects of base and prime editing in human hematopoietic stem cells
Base and prime editors (BEs and PEs) may provide more precise genetic engineering than nuclease-based approaches because they bypass the dependence...

-
Base Editing of Human Hematopoietic Stem Cells
Base editing by nucleotide deaminases linked to programmable DNA-binding proteins represents a promising approach to remedy blood disorders. Here we...
-
Engineered domain-inlaid Nme2Cas9 adenine base editors with increased on-target DNA editing and targeting scope
BackgroundNme2ABE8e has been constructed and characterized as a compact, accurate adenine base editor with a less restrictive dinucleotide...

-
Deep learning models to predict the editing efficiencies and outcomes of diverse base editors
Applications of base editing are frequently restricted by the requirement for a protospacer adjacent motif (PAM), and selecting the optimal base...

-
Cas9 variants expand the targeting scope of base editing systems in bacteria
Base editors (BEs) consist of partially active Cas9 homologs fused to nucleobase-converting DNA deaminases along with certain accessory proteins. The...

-
Map** variant effects on anti-tumor hallmarks of primary human T cells with base-editing screens
Single-nucleotide variants (SNVs) in key T cell genes can drive clinical pathologies and could be repurposed to improve cellular cancer...

