Search
Search Results
-
G-Aligner: a graph-based feature alignment method for untargeted LC–MS-based metabolomics
BackgroundLiquid chromatography–mass spectrometry is widely used in untargeted metabolomics for composition profiling. In multi-run analysis...

-
EMMA: a new method for computing multiple sequence alignments given a constraint subset alignment
BackgroundAdding sequences into an existing (possibly user-provided) alignment has multiple applications, including updating a large alignment with...

-
Fast Alignment
We’ve seen that optimal alignment is slow but accurate while exact matching is fast but cannot account for mutations. In this Chapter we tweak exact...
-
Protein embedding based alignment
PurposeDespite the many progresses with alignment algorithms, aligning divergent protein sequences with less than 20–35% pairwise identity (so called...

-
Achieving quantitative reproducibility in label-free multisite DIA experiments through multirun alignment
DIA is a mainstream method for quantitative proteomics, but consistent quantification across multiple LC-MS/MS instruments remains a bottleneck in...

-
Optimal Alignment
When we align two sequences, we in fact propose an evolutionary history for them. A history, where we account for three kinds of events, mutation,...
-
POSMM: an efficient alignment-free metagenomic profiler that complements alignment-based profiling
We present here POSMM (pronounced ‘Possum’), Python-Optimized Standard Markov Model classifier, which is a new incarnation of the Markov model...

-
Protein remote homology detection and structural alignment using deep learning
Exploiting sequence–structure–function relationships in biotechnology requires improved methods for aligning proteins that have low sequence...

-
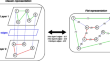
Multilayer network alignment based on topological assessment via embeddings
BackgroundNetwork graphs allow modelling the real world objects in terms of interactions. In a multilayer network, the interactions are distributed...
-
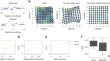
Alignment of spatial genomics data using deep Gaussian processes
Spatially resolved genomic technologies have allowed us to study the physical organization of cells and tissues, and promise an understanding of...

-
Deep embedding and alignment of protein sequences
Protein sequence alignment is a key component of most bioinformatics pipelines to study the structures and functions of proteins. Aligning highly...

-
ACMGA: a reference-free multiple-genome alignment pipeline for plant species
BackgroundThe short-read whole-genome sequencing (WGS) approach has been widely applied to investigate the genomic variation in the natural...

-
Sensitive inference of alignment-safe intervals from biodiverse protein sequence clusters using EMERALD
Sequence alignments are the foundations of life science research, but most innovation so far focuses on optimal alignments, while information derived...

-
Kinesin-7 CENP-E mediates chromosome alignment and spindle assembly checkpoint in meiosis I
In eukaryotes, meiosis is the genetic basis for sexual reproduction, which is important for chromosome stability and species evolution. The defects...

-
Machine learning on alignment features for parent-of-origin classification of simulated hybrid RNA-seq
BackgroundParent-of-origin allele-specific gene expression (ASE) can be detected in interspecies hybrids by virtue of RNA sequence variants between...

-
Sequence Alignment
New biological sequences do not emerge de novo in nature but rather are derived from pre-existing sequences. This foundational principle underlies...
-
Alignment and integration of spatial transcriptomics data
Spatial transcriptomics (ST) measures mRNA expression across thousands of spots from a tissue slice while recording the two-dimensional (2D)...

-
Pairwise Alignment, Multiple Alignment, and BLAST
Quantitative comparison of sequences can be performed pairwise by aligning two sequences based on considerations of gaps representing insertions or...
-
KAGE: fast alignment-free graph-based genoty** of SNPs and short indels
Genoty** is a core application of high-throughput sequencing. We present KAGE, a genotyper for SNPs and short indels that is inspired by recent...

-
cnnLSV: detecting structural variants by encoding long-read alignment information and convolutional neural network
BackgroundGenomic structural variant detection is a significant and challenging issue in genome analysis. The existing long-read based structural...

