Search
Search Results
-
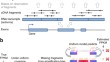
Performance assessment of DNA sequencing platforms in the ABRF Next-Generation Sequencing Study
Assessing the reproducibility, accuracy and utility of massively parallel DNA sequencing platforms remains an ongoing challenge. Here the Association...

-
Bear biometrics: develo** an individual recognition technique for sloth bears
Identifying individual animals, especially in large mammals, is an important goal for wildlife biologists and managers. Bears, occupying diverse...

-
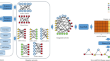
CircWalk: a novel approach to predict CircRNA-disease association based on heterogeneous network representation learning
BackgroundSeveral types of RNA in the cell are usually involved in biological processes with multiple functions. Coding RNAs code for proteins while...
-
aFold – using polynomial uncertainty modelling for differential gene expression estimation from RNA sequencing data
BackgroundData normalization and identification of significant differential expression represent crucial steps in RNA-Seq analysis. Many available...

-

-
Accumulated Endogenous Abscisic Acid Contributes to the Cold Tolerance of Pre-planted Cultivated Tobacco
Pre-planted cultivation is widely used in tobacco seedlings in south China to avoid damage from the cold. However, the underlying mechanisms of...

-
Physiological and Molecular Bases of Drought and Heat Tolerance in Pearl Millet
Pearl millet is one of the most important sources of nutrition for millions of people in arid and semi-arid areas in Africa and Asia. Farmers have...
-

-
Omics-Based Approaches in Improving Drought Stress Tolerance in Pearl Millet
Pearl millet (Pennisetum glaucum (L.) R. Br), a climate-resilient Nutri-cereal grown primarily in Africa and South Asia’s arid and semi-arid regions,...
-
The intestinal digesta microbiota of tropical marine fish is largely uncultured and distinct from surrounding water microbiota
Studying the gut microbes of marine fishes is an important part of conservation as many fish species are increasingly threatened by extinction. The...

-
Multi-drug resistant (MDR), extended spectrum beta-lactamase (ESBL) producing and carbapenem resistant Escherichia coli in rescued Sloth bears (Melursus ursinus), India
The study reports the multi-drug resistant (MDR), extended spectrum beta-lactamase (ESBL) producing and carbapenem resistant Escherichia coli (CRE)...

-
Towards accurate and reliable resolution of structural variants for clinical diagnosis
Structural variants (SVs) are a major source of human genetic diversity and have been associated with different diseases and phenotypes. The...

-
Systematic evaluation of RNA-Seq preparation protocol performance
BackgroundRNA-Seq is currently the most widely used tool to analyze whole-transcriptome profiles. There are numerous commercial kits available to...

-
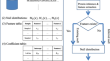
A new estimation of protein-level false discovery rate
BackgroundIn mass spectrometry-based proteomics, protein identification is an essential task. Evaluating the statistical significance of the protein...
-
Best practices and tools for reporting reproducible fluorescence microscopy methods
Although fluorescence microscopy is ubiquitous in biomedical research, microscopy methods reporting is inconsistent and perhaps undervalued. We...

-
Comparative performance of the BGISEQ-500 and Illumina HiSeq4000 sequencing platforms for transcriptome analysis in plants
BackgroundThe next-generation sequencing (NGS) technology has greatly facilitated genomic and transcriptomic studies, contributing significantly in...

-
Cross-site comparison of ribosomal depletion kits for Illumina RNAseq library construction
BackgroundRibosomal RNA (rRNA) comprises at least 90% of total RNA extracted from mammalian tissue or cell line samples. Informative transcriptional...

-
Human splicing diversity and the extent of unannotated splice junctions across human RNA-seq samples on the Sequence Read Archive
BackgroundGene annotations, such as those in GENCODE, are derived primarily from alignments of spliced cDNA sequences and protein sequences. The...

-
Modeling of RNA-seq fragment sequence bias reduces systematic errors in transcript abundance estimation
We find that current computational methods for estimating transcript abundance from RNA-seq data can lead to hundreds of false-positive results. We...