Search
Search Results
-
GCNCPR-ACPs: a novel graph convolution network method for ACPs prediction
BackgroundAnticancer peptide (ACP) inhibits and kills tumor cells. Research on ACP is of great significance for the development of new drugs, and the...

-
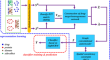
Drug-target interaction prediction based on spatial consistency constraint and graph convolutional autoencoder
BackgroundDrug-target interaction (DTI) prediction plays an important role in drug discovery and repositioning. However, most of the computational...
-
Small networks of expressed genes in the whole blood and relationships to profiles in circulating metabolites provide insights in inter-individual variability of feed efficiency in growing pigs
BackgroundFeed efficiency is a research priority to support a sustainable meat production. It is recognized as a complex trait that integrates...

-
Equivariant score-based generative diffusion framework for 3D molecules
BackgroundMolecular biology is crucial for drug discovery, protein design, and human health. Due to the vastness of the drug-like chemical space,...

-
Robust, scalable, and informative clustering for diverse biological networks
Clustering molecular data into informative groups is a primary step in extracting robust conclusions from big data. However, due to foundational...

-
PWN: enhanced random walk on a warped network for disease target prioritization
BackgroundExtracting meaningful information from unbiased high-throughput data has been a challenge in diverse areas. Specifically, in the early...

-
ScMOGAE: A Graph Convolutional Autoencoder-Based Multi-omics Data Integration Framework for Single-Cell Clustering
The integration of single-cell multi-omics data is a significant step forward in our understanding of the complex biological systems at the cellular...
-
Revisiting a trophic overlap-based measure for species uniqueness in ecological networks
Species uniqueness can be measured from the network perspective offering an alternative view to species importance. Direct and indirect effects...

-
De novo reconstruction of cell interaction landscapes from single-cell spatial transcriptome data with DeepLinc
Based on a deep generative model of variational graph autoencoder (VGAE), we develop a new method, DeepLinc (deep learning framework for Landscapes...

-
Development of performance and learning rate evaluation models in robot-assisted surgery using electroencephalography and eye-tracking
The existing performance evaluation methods in robot-assisted surgery (RAS) are mainly subjective, costly, and affected by shortcomings such as the...

-
Macroecological Data
Following the overall discussion on conceptual and methodological issues, it is essential to start thinking about how to get data to evaluate...
-
Degree-Normalization Improves Random-Walk-Based Embedding Accuracy in PPI Graphs
Among the many proposed solutions in graph embedding, traditional random walk-based embedding methods have shown their promise in several fields....
-
Network-based restoration strategies maximize ecosystem recovery
Redressing global patterns of biodiversity loss requires quantitative frameworks that can predict ecosystem collapse and inform restoration...

-
AMEND: active module identification using experimental data and network diffusion
BackgroundMolecular interaction networks have become an important tool in providing context to the results of various omics experiments. For example,...

-
DisoFLAG: accurate prediction of protein intrinsic disorder and its functions using graph-based interaction protein language model
Intrinsically disordered proteins and regions (IDPs/IDRs) are functionally important proteins and regions that exist as highly dynamic conformations...

-
FFP: joint Fast Fourier transform and fractal dimension in amino acid property-aware phylogenetic analysis
BackgroundAmino acid property-aware phylogenetic analysis (APPA) refers to the phylogenetic analysis method based on amino acid property encoding,...

-
SpaDecon: cell-type deconvolution in spatial transcriptomics with semi-supervised learning
Spatially resolved transcriptomics (SRT) has advanced our understanding of the spatial patterns of gene expression, but the lack of single-cell...

-
Additional Neural Matrix Factorization model for computational drug repositioning
BackgroundComputational drug repositioning, which aims to find new applications for existing drugs, is gaining more attention from the pharmaceutical...

-
Evaluating spatially variable gene detection methods for spatial transcriptomics data
BackgroundThe identification of genes that vary across spatial domains in tissues and cells is an essential step for spatial transcriptomics data...

-
Predicting abiotic stress-responsive miRNA in plants based on multi-source features fusion and graph neural network
BackgroundMore and more studies show that miRNA plays a crucial role in plants' response to different abiotic stresses. However, traditional...

