Search
Search Results
-
Protein–protein interaction and non-interaction predictions using gene sequence natural vector
Predicting protein–protein interaction and non-interaction are two important different aspects of multi-body structure predictions, which provide...

-
Building, Visualizing, and Analyzing Glycosaminoglycan–Protein Interaction Networks
This chapter describes how to generate, visualize, and analyze interaction networks of glycosaminoglycans (GAGs), which are linear polyanionic...
-
Efficient link prediction in the protein–protein interaction network using topological information in a generative adversarial network machine learning model
BackgroundThe investigation of possible interactions between two proteins in intracellular signaling is an expensive and laborious procedure in the...

-
Normalized L3-based link prediction in protein–protein interaction networks
BackgroundProtein–protein interaction (PPI) data is an important type of data used in functional genomics. However, high-throughput experiments are...

-
Protein–protein interaction site prediction by model ensembling with hybrid feature and self-attention
BackgroundProtein–protein interactions (PPIs) are crucial in various biological functions and cellular processes. Thus, many computational approaches...

-
A seed expansion-based method to identify essential proteins by integrating protein–protein interaction sub-networks and multiple biological characteristics
BackgroundThe identification of essential proteins is of great significance in biology and pathology. However, protein–protein interaction (PPI) data...
-
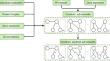
Molecular complex detection in protein interaction networks through reinforcement learning
BackgroundProteins often assemble into higher-order complexes to perform their biological functions. Such protein–protein interactions (PPI) are...

-
DisoFLAG: accurate prediction of protein intrinsic disorder and its functions using graph-based interaction protein language model
Intrinsically disordered proteins and regions (IDPs/IDRs) are functionally important proteins and regions that exist as highly dynamic conformations...

-
Computational prediction of protein–protein interactions’ network in Arabidopsis thaliana
The study of protein–protein interactions (PPIs) has been a major factor in understanding the function of proteins. The development of diverse...

-
Densest subgraph-based methods for protein-protein interaction hot spot prediction
BackgroundHot spots play an important role in protein binding analysis. The residue interaction network is a key point in hot spot prediction, and...

-

-
LINC02015 modulates the cell proliferation and apoptosis of aortic vascular smooth muscle cells by transcriptional regulation and protein interaction network
Long intergenic nonprotein coding RNA 2015 (LINC02015) is a long non-coding RNA that has been found elevated in various cell proliferation-related...

-
Systematic comparison of the protein-protein interaction network of bacterial Universal stress protein A (UspA): an insight into its discrete functions
Universal stress protein A (UspA) is ubiquitously over-expressed in varied species of pathogenic bacteria under stress conditions and helps in cell...

-
Protein–Protein Interaction Network Map** by Affinity Purification Cross-Linking Mass Spectrometry (AP-XL-MS) based Proteomics
Protein–protein interactions (PPIs) are the physical interactions formed among proteins. These interactions are primarily functional, i.e., they...
-
Computational Methods and Deep Learning for Elucidating Protein Interaction Networks
Protein interactions play a critical role in all biological processes, but experimental identification of protein interactions is a time- and...
-
Predicting Drug-Target Affinity Using Protein Pocket and Graph Convolution Network
Drug–target affinity (DTA) plays a crucial role in the discovery and development of pharmaceuticals. The localized structure of protein pockets plays...
-
Struct2Graph: a graph attention network for structure based predictions of protein–protein interactions
BackgroundDevelopment of new methods for analysis of protein–protein interactions (PPIs) at molecular and nanometer scales gives insights into...

-
PCPI: Prediction of circRNA and Protein Interaction Using Machine Learning Method
Circular RNA (circRNA) is an RNA molecule different from linear RNA with covalently closed loop structure. CircRNAs can act as sponging miRNAs and...
-
Protein–Protein Interactions in Neurodegenerative Diseases
Neurodegeneration is a state of progressive decay of neuronal structure and function. It has gained scientific attention owing to the fact that...
-
Hsp104p: a protein disaggregase
All newly synthesized proteins must fold to their correct native conformation in order to function. That protein folding in the crowded...
