Search
Search Results
-
Ubiquitin-specific proximity labeling for the identification of E3 ligase substrates
Protein ubiquitylation controls diverse processes within eukaryotic cells, including protein degradation, and is often dysregulated in disease....

-
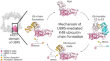
Structural snapshots along K48-linked ubiquitin chain formation by the HECT E3 UBR5
Ubiquitin (Ub) chain formation by homologous to E6AP C-terminus (HECT)-family E3 ligases regulates vast biology, yet the structural mechanisms remain...
-
CUL5-ARIH2 E3-E3 ubiquitin ligase structure reveals cullin-specific NEDD8 activation
An emerging mechanism of ubiquitylation involves partnering of two distinct E3 ligases. In the best-characterized E3-E3 pathways, ARIH-family...

-
A Computational Tool for Analysis of Mass Spectrometry Data of Ubiquitin-Enriched Samples
Mass spectrometry data on ubiquitin and ubiquitin-like modifiers are becoming increasingly more accessible, and the coverage progressively deepen as...
-
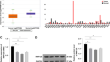
E3 Ubiquitin Ligase RNF125 Suppresses Immune Escape in Head and Neck Squamous Cell Carcinoma by Regulating PD-L1 Expression
Head and neck squamous cell carcinoma (HNSCC) is a lethal malignancy. Given the essential roles of E3 ligases in cancer immunotherapies, this paper...
-
cIAP1-based degraders induce degradation via branched ubiquitin architectures
Targeted protein degradation through chemical hijacking of E3 ubiquitin ligases is an emerging concept in precision medicine. The ubiquitin code is a...

-
Functional E3 ligase hotspots and resistance mechanisms to small-molecule degraders
Targeted protein degradation is a novel pharmacology established by drugs that recruit target proteins to E3 ubiquitin ligases. Based on the...

-
Thioester and Oxyester Linkages in the Ubiquitin System
The traditional textbook describes ubiquitylation as the conjugation of ubiquitin to a target by forming a covalent bond connecting ubiquitin’s...
-
Evaluating Ligands for Ubiquitin Ligases Using Affinity Beads
Proteolysis-targeting chimera (PROTAC®) protein degraders are heterobifunctional small molecules that bind a specific target protein on one end and a...
-
Simulation and Computational Study of RING Domain Mutants of BRCA1 and Ube2k in AD/PD Pathophysiology
Lysine-based post-translational modification (PTM) such as acylation, acetylation, deamination, methylation, SUMOylation, and ubiquitination has...

-
Activity-based profiling of cullin–RING E3 networks by conformation-specific probes
The cullin–RING ubiquitin ligase (CRL) network comprises over 300 unique complexes that switch from inactive to activated conformations upon...

-

-
A Mass Spectrometry-Based Strategy for Map** Modification Sites for the Ubiquitin-Like Modifier NEDD8
The identification of modification sites for ubiquitin and ubiquitin-like modifiers is an essential step in the elucidation of controlled processes....
-
A CRISPR activation screen identifies FBXO22 supporting targeted protein degradation
Targeted protein degradation (TPD) represents a potent chemical biology paradigm that leverages the cellular degradation machinery to...

-
Identification of Ubiquitination-Associated Proteins Using 2D-DIGE
Ubiquitination is a post-translational modification, in which a small regulatory protein (~8.6 kDa) is tagged as a single moiety or as a chain to...
-
HUWE1 employs a giant substrate-binding ring to feed and regulate its HECT E3 domain
HUWE1 is a universal quality-control E3 ligase that marks diverse client proteins for proteasomal degradation. Although the giant HECT enzyme is an...

-
Monitoring HECT Ubiquitination Activity In Vitro
In vitro ubiquitination tools have been employed to mechanistically study the ubiquitin enzymatic cascade. Here, we describe an assay capable to...
-
Recruitment of FBXO22 for targeted degradation of NSD2
Targeted protein degradation (TPD) is an emerging therapeutic strategy that would benefit from new chemical entities with which to recruit a wider...

-
Structure of UBE2K–Ub/E3/polyUb reveals mechanisms of K48-linked Ub chain extension
Ubiquitin (Ub) chain types govern distinct biological processes. K48-linked polyUb chains target substrates for proteasomal degradation, but the...

-
Targeted degradation of oncogenic BCR-ABL by silencing the gene of NEDD8 E3 ligase RAPSYN
Tyrosine kinase inhibitors have been the standard treatment for patients with Philadelphia chromosome-positive (Ph + ) leukemia. However, a series of...

