Search
Search Results
-
Investigating the role of glycans in Omicron sub-lineages XBB.1.5 and XBB.1.16 binding to host receptor using molecular dynamics and binding free energy calculations
Omicron derived lineages viz. BA.2, BA.3, BA.4 BA.5, BF.7 and XBBs show prominence with improved immune escape, transmissibility, infectivity, and...

-
The maximal and current accuracy of rigorous protein-ligand binding free energy calculations
Computational techniques can speed up the identification of hits and accelerate the development of candidate molecules for drug discovery. Among...

-
Automated relative binding free energy calculations from SMILES to ΔΔG
In drug discovery, computational methods are a key part of making informed design decisions and prioritising experiments. In particular, optimizing...

-
pH-Dependent binding energy-induced inflection-point behaviors for pH-universal hydrogen oxidation reaction
The kinetics of hydrogen oxidation reaction (HOR) declines with orders of magnitude when the electrolyte varies from acid to base. Therefore,...

-
Study of SQ109 analogs binding to mycobacterium MmpL3 transporter using MD simulations and alchemical relative binding free energy calculations
N -geranyl- N ΄-(2-adamantyl)ethane-1,2-diamine (SQ109) is a tuberculosis drug that has high potency against Mycobacterium tuberculosis (Mtb) and may...

-
Exploring binding positions and backbone conformations of peptide ligands of proteins with a backbone-centred statistical energy function
When designing peptide ligands based on the structure of a protein receptor, it can be very useful to narrow down the possible binding positions and...

-
The slow but steady rise of binding free energy calculations in drug discovery
Binding free energy calculations are increasingly used in drug discovery research to predict protein-ligand binding affinities and to prioritize...

-
Decoding the deactivation mechanism of R192W mutation of ZAP-70 using molecular dynamics simulations and binding free energy calculations
ContextZAP-70 (zeta-chain-associated protein of 70 kDa), serving as a critical regulator for T cell antigen receptor signaling, represents an...

-
Computational Protein Binding
Why it is important to know this material? The computational approach to protein binding has become a crucial methodology in drug design and...
-
Evaluating the use of absolute binding free energy in the fragment optimisation process
Key to the fragment optimisation process within drug design is the need to accurately capture the changes in affinity that are associated with a...

-
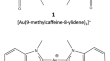
The FMO2 analysis of the ligand-receptor binding energy: the Biscarbene-Gold(I)/DNA G-Quadruplex case study
In this work, the ab initio fragment molecular orbital (FMO) method was applied to calculate and analyze the binding energy of two biscarbene-Au(I)...
-
Quantum-Chemical Study of the Energy of Binding Lithium Endofullerene Ion Li+@C60 to Anions
AbstractThe optimal geometry and binding energy (Δ E ) of Li + @C 60 A ‒ ion pairs (A = BF 4 , AsF 6 , PF 6 , FSI, TFSI, and 4F-BB) in vacuum and...

-
Protein-Ligand Interactions and Binding
Why it is important to know this material? Protein binding is implied in all biological mechanisms, and the definition of quantitative models and...
-
Comprehensive evaluation of end-point free energy techniques in carboxylated-pillar[6]arene host–guest binding: I. Standard procedure
Despite the massive application of end-point free energy methods in protein–ligand and protein–protein interactions, computational understandings...

-
Comprehensive evaluation of end-point free energy techniques in carboxylated-pillar[6]arene host–guest binding: II. regression and dielectric constant
End-point free energy calculations as a powerful tool have been widely applied in protein–ligand and protein–protein interactions. It is often...

-
Evaluation of interactions between the hepatitis C virus NS3/4A and sulfonamidobenzamide based molecules using molecular docking, molecular dynamics simulations and binding free energy calculations
The Hepatitis C Virus (HCV) NS3/4A is an attractive target for the treatment of Hepatitis C infection. Herein, we present an investigation of HCV...

-
Difference in the binding mechanisms of ABT-263/43b with Bcl-xL/Bcl-2: computational perspective on the accurate binding free energy analysis
B-cell lymphoma/leukemia gene-2(Bcl-2) protein family known for regulating cell cycle arrest and subsequent cell death is highly expressed in a...

-
Identification of some dietary flavonoids as potential inhibitors of TMPRSS2 through protein–ligand interaction studies and binding free energy calculations
The continuing threat of COVID-19 and deaths need an urgent cost-effective pharmacological approach. Here, we examine the inhibitory activity of a...

-
Multiscale Molecular Dynamics Simulations of Ice-Binding Proteins
Ice-binding proteins (IBPs) are a diverse class of proteins that are essential for the survival of organisms in cold conditions. IBPs are diverse in...
-
Discovery of novel indoleamine 2,3-dioxygenase-1 (IDO-1) inhibitors: pharmacophore-based 3D-QSAR, Gaussian field-based 3D-QSAR, docking, and binding free energy studies
Indoleamine 2,3-dioxygenase-1 (IDO-1) is a heme-containing enzyme that initiates the kynurenine pathway by catalyzing the first step in l-tryptophan...

