Search
Search Results
-
The landscape of long noncoding RNAs in the human transcriptome
Long noncoding RNAs (lncRNAs) are emerging as important regulators of tissue physiology and disease processes including cancer. To delineate...

-
Whole genome sequencing and de novo assembly identifies Sydney-like variant noroviruses and recombinants during the winter 2012/2013 outbreak in England
BackgroundNorovirus is the commonest cause of epidemic gastroenteritis among people of all ages. Outbreaks frequently occur in hospitals and the...

-
Genome sequence of the cultivated cotton Gossypium arboreum
The complex allotetraploid nature of the cotton genome (AADD; 2 n = 52) makes genetic, genomic and functional analyses extremely challenging. Here we...

-
Identification and characterization of alternative splicing in parasitic nematode transcriptomes
BackgroundAlternative splicing (AS) of mRNA is a vital mechanism for enhancing genomic complexity in eukaryotes. Spliced isoforms of the same gene...

-
Whole-genome re-sequencing of non-model organisms: lessons from unmapped reads
Unmapped reads are often discarded from the analysis of whole-genome re-sequencing, but new biological information and insights can be uncovered...

-
De novo likelihood-based measures for comparing genome assemblies
BackgroundThe current revolution in genomics has been made possible by software tools called genome assemblers, which stitch together DNA fragments...

-
mPUMA: a computational approach to microbiota analysis by de novo assembly of operational taxonomic units based on protein-coding barcode sequences
BackgroundFormation of operational taxonomic units (OTU) is a common approach to data aggregation in microbial ecology studies based on amplification...

-
Discovery and Classification of Homeobox Genes in Animal Genomes
The diversification of homeobox genes is of great interest to evolutionary and developmental biology. To generate a catalogue of all homeobox genes...
-
De Novo Assembly Algorithms
In an ideal case, an assembly algorithm should merge overlapped reads to one long continuous sequence, called contig, which is a chromosome in the...
-
Minke whale genome and aquatic adaptation in cetaceans
The shift from terrestrial to aquatic life by whales was a substantial evolutionary event. Here we report the whole-genome sequencing and de novo ...

-
The Impact of Next-Generation Sequencing Technology on Bacterial Genomics
For many decades, genomic studies were based on Sanger sequencing or the dideoxy chain termination method of sequencing DNA, along with microarray...
-
Monoaminergic modulation of behavioural and electrophysiological indices of error processing
RationaleError processing is a critical executive function that is impaired in a large number of clinical populations. Although the neural...

-
Genomic analyses identify distinct patterns of selection in domesticated pigs and Tibetan wild boars
We report the sequencing at 131× coverage, de novo assembly and analyses of the genome of a female Tibetan wild boar. We also resequenced the whole...

-
De novo assembly and genoty** of variants using colored de Bruijn graphs
Detecting genetic variants that are highly divergent from a reference sequence remains a major challenge in genome sequencing. We introduce de novo ...

-
ncRNA–Protein Interactions in Development and Disease from the Perspective of High-Throughput Studies
Genome-scale studies have provided strong support to the prevalent transcription of nonprotein-coding RNAs (ncRNAs) in various organisms. The immense...
-
Computational solutions for omics data
High-throughput experimental technologies are generating increasingly massive and complex genomic data sets. The sheer enormity and heterogeneity of...
-
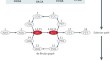
Sequence assembly demystified
Advances in sequencing technologies and increased access to sequencing services have led to renewed interest in sequence and genome assembly....

-
Comparative Genomics and Pathogenicity Islands of Corynebacterium diphtheriae, Corynebacterium ulcerans, and Corynebacterium pseudotuberculosis
The systematic application of next-generation DNA sequencing technologies has provided detailed insights into the genomics of corynebacteria. The...
-
Arapan-S: a fast and highly accurate whole-genome assembly software for viruses and small genomes
BackgroundGenome assembly is considered to be a challenging problem in computational biology, and has been studied extensively by many researchers....

-
Genome Assembler for Repetitive Sequences
The article presents a new algorithm for random DNA fragment assembly. The algorithm uses an extended de Bruijn graph that stores information of...
