Abstract
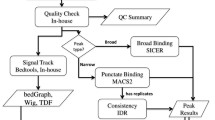
Chromatin immunoprecipitation followed by high-throughput sequencing (ChIP-seq) is invaluable for identifying genome-wide binding of transcription factors and map** of epigenomic profiles. We present a statistical protocol for analyzing ChIP-seq data. We describe guidelines for data preprocessing and quality control and provide detailed examples of identifying ChIP-enriched regions using the Bioconductor package “mosaics.”
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Shen Y, Yue F, McCleary DF, Ye Z, Edsall L, Kuan S, Wagner U, Dixon J, Lee L, Lobanenkov VV, Ren B (2012) A map of cis-regulatory sequences in the mouse genome. Nature 488:116–120
Fujiwara T, O’Geen H, Keles S, Blahnik K, Linnemann AK, Kang Y, Choi K, Farnham PJ, Bresnick EH (2009) Discovering hematopoietic mechanisms through genome-wide analysis of GATA factor chromatin occupancy. Mol Cell 36(4):667–681
Wilbanks EG, Facciotti MT (2010) Evaluation of algorithm performance in ChIP-Seq peak detection. PLoS One 5:e11471
Chen Y, Negre N, Li Q, Mieczkowska JO, Slattery M, Liu T, Zhang T, Kim T-K, He HH, Zieba J, Ruan Y, Bickel PJ, Myers RM, Wold BJ, White KP, Lieb JD, Liu XS (2012) Systematic evaluation of factors influencing ChIP-seq fidelity. Nat Methods 9(6):609–614
Kuan PF, Chung D, Pan G, Thomson JA, Stewart R, Keles S (2011) A statistical framework for the analysis of ChIP-Seq data. J Am Stat Assoc 106(495):891–903
Chung D, Kuan P-F, Li B, SanalKumar R, Liang K, Bresnick E, Dewey C, Keles S (2011) Discovering transcription factor binding sites in highly repetitive regions of genomeswith multi-read analysis of ChIP-Seq data. PLoS Comput Biol 7(7):e1002111
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:R25
Rozowsky J, Euskirchen G, Auerbach R, Zhang Z, Gibson T, Bjornson R, Carriero N, Snyder M, Gerstein M (2009) PeakSeq enables systematic scoring of ChIP-seq experiments relative to controls. Nat Biotechnol 27:66–75
Benjamini Y, Speed TS (2012) Summarizing and correcting the GC content bias in high-throughput sequencing. Nucleic Acids Res 40(10):e72
Liang K, Keles S (2012) Normalization of ChIP-seq data with control. BMC Bioinformatics 13:199
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J Roy Stat Soc B Met 57(1):289–300
Liang K, Keles S (2012) Detecting differential binding of transcription factors with ChIP-seq. Bioinformatics 28(1):121–122
Zeng X, Sanalkumar R, Bresnick EH, Li H, Chang Q, Keles S (2012) jMOSAiCS: joint analysis of multiple ChIP-seq datasets. Submitted. Technical report available at http://www.stat.wisc.edu/~keles/Papers/jmosaics.pdf. R package available at http://www.stat.wisc.edu/~keles/Software/
Acknowledgments
This work is supported by National Institutes of Health Grants (HG0067161, HG003747) to S.K. We thank Audrey Gasch and Jeff Lewis (yeast TFx), John Svaren and Ra**i Srinivasan (Sox10 in rat), and Qiang Chang and Emily Cunningham (human ChIP-seq) for the datasets and useful discussions regarding the analysis.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Appendix: R Script for the Analysis of Yeast TFx ChIP-seq Datasets
Appendix: R Script for the Analysis of Yeast TFx ChIP-seq Datasets
library( mosaics)
library( hexbin)
# construct bin-level files for each replicate ChIP sample #
constructBins(infile = "TFx_EtOH_IP_1_2mis_bowtie_uni.txt", outfileLoc = "/bin_cap0",
fileFormat = "bowtie", byChr = FALSE, fragLen = 200, binSize = 200, cap** = 0, PET = FALSE)
constructBins(infile = "TFx_EtOH_IP_2_2mis_bowtie_uni.txt", outfileLoc = "/bin_cap0",
fileFormat = "bowtie", byChr = FALSE, fragLen = 200, binSize = 200, cap** = 0, PET = FALSE)
constructBins( infile = "TFx_EtOH_IP_3_2mis_bowtie_uni.txt", outfileLoc = "/bin_cap0",
fileFormat = "bowtie", byChr = FALSE, fragLen = 200, binSize = 200, cap** = 0, PET = FALSE)
# construct bin-level files for pooled ChIP (uncapped and capped by 3) and pooled input samples #
constructBins( infile = "TFx_EtOH_input_2mis_bowtie_uni.txt", outfileLoc = "/bin_cap0",
fileFormat = "bowtie", byChr = FALSE, fragLen = 200, binSize = 200, cap** = 0, PET = FALSE)
constructBins( infile = "TFx_EtOH_input_2mis_bowtie_uni.txt", outfileLoc = "/bin_cap3",
fileFormat = "bowtie", byChr = FALSE, fragLen = 200, binSize = 200, cap** = 3, PET = FALSE)
constructBins( infile = "TFx_EtOH_chip_2mis_bowtie_uni.txt", outfileLoc = "/bin_cap0",
fileFormat = "bowtie", byChr = FALSE, fragLen = 200, binSize = 200, cap** = 0, PET = FALSE)
# generate hexbin plots between replicates to decide on pooling #
bin_rep = vector( “list”,3)
bin_rep[[1]] = readBins(type = c("chip","input"), fileName = c( "/bin_cap0/TFx_EtOH_IP_1_2mis_bowtie_uni.txt_fragL200_bin200.txt"," /bin_cap0/TFx_EtOH_IP_2_2mis_bowtie_uni.txt_fragL200_bin200.txt"))
bin_rep[[2]] = readBins(type = c("chip","input"), fileName = c( "/bin_cap0/TFx_EtOH_IP_1_2mis_bowtie_uni.txt_fragL200_bin200.txt","/bin_cap0/TFx_EtOH_IP_3_2mis_bowtie_uni.txt_fragL200_bin200.txt"))
bin_rep[[3]] = readBins(type = c("chip","input"), fileName = c( "/bin_cap0/TFx_EtOH_IP_2_2mis_bowtie_uni.txt_fragL200_bin200.txt"," /bin_cap0/TFx_EtOH_IP_3_2mis_bowtie_uni.txt_fragL200_bin200.txt"))
xlabel = c( “rep1”, ”rep1”, ”rep2”); ylabel = c( “rep2”, ”rep3”, ”rep3”)
for (i in 1:3) {
a = hexbin( bin_rep[[i]]@tagCount, bin_rep[[i]]@input, xbins = 100)
plot( a, trans = log, inv = exp, xlab = xlabel[i], ylab = ylabel[i], colramp = rainbow)
}
# generate hexbin plots of uncapped and capped bin-level read count data #
bin_cap = readBins(type = c("chip","input"), filename = c( "/bin_cap0/TFx_EtOH_IP_2mis_bowtie_uni.txt_fragL200_bin200.txt", "/bin_cap3/TFx_EtOH_IP_2mis_bowtie_uni.txt_fragL200_bin200.txt"))
a = hexbin (bin_cap@ input, bin_cap@ tagCount, xbins = 100)
plot(a, trans = log, inv = exp, xlab = "Capped counts", ylab = "Uncapped counts", colramp = rainbow)
# load bin-level files for MOSAiCS analysis #
bin = readBins(type = c("chip","input"), filename = c("/bin_cap0/TFx_EtOH_IP_2mis_bowtie_uni.txt_fragL200_bin200.txt", "/bin_cap0/TFx_EtOH_input_2mis_bowtie_uni.txt_fragL200_bin200.txt"))
# generate hexbin plot to check ChIP enrichment over Input#
a = hexbin (bin@input, bin@tagCount, xbins = 100)
plot(a, trans = log, inv = exp, xlab = "Input", ylab = "ChIP", colramp = rainbow)
# histogram of bin-level files #
plot(bin)
# use Input-only model to fit data #
fit = mosaicsFit(bin, analysisType = "IO", bgEst = "automatic", truncProb = 0.08)
# goodness-of-fit plot of the model #
plot(fit)
# call peaks #
thres = quantile (fit@tagCount, probs = 0.95)
peak = mosaicsPeak(fit, signalModel = "2S", FDR = 0.05, maxgap = 200, minsize = 50, thres = thres)
# export peaks #
export(peak, type = "txt", filename = "TFx_TSpeakList.txt")
export(peak, type = "bed", filename = "TFx_TSpeakList.bed")
# generate wig files #
generateWig(infile = "TFx_EtOH_IP_2mis_bowtie_uni.txt ", PET = FALSE, fileFormat = "bowtie", outfileLoc = "/bin_cap0/", byChr = FALSE, useChrfile = FALSE, chrfile = NULL, fragLen = 200, span = 200, cap** = 0)
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media New York
About this protocol
Cite this protocol
Sun, G., Chung, D., Liang, K., Keleş, S. (2013). Statistical Analysis of ChIP-seq Data with MOSAiCS. In: Shomron, N. (eds) Deep Sequencing Data Analysis. Methods in Molecular Biology, vol 1038. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-514-9_12
Download citation
DOI: https://doi.org/10.1007/978-1-62703-514-9_12
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-513-2
Online ISBN: 978-1-62703-514-9
eBook Packages: Springer Protocols