Abstract
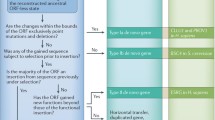
De novo genes, that is, protein-coding genes originating from previously noncoding sequence, have gone from being considered impossibly unlikely to being recognized as an important source of genetic novelty in eukaryotic genomes. It is clear that de novo gene evolution is a rare but consistent feature of eukaryotic genomes, being detected in every genome studied. However, different studies often use different computational methods, and the numbers and identities of the detected genes vary greatly. Here we present a coherent protocol for the computational identification of de novo genes by comparative genomics. The method described uses homology searches, identification of syntenic regions, and ancestral sequence reconstruction to produce high-confidence candidates with robust evidence of de novo emergence. It is designed to be easily applicable given the basic knowledge of bioinformatic tools and scalable so that it can be applied on large and small datasets.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Long M, Betrán E, Thornton K et al (2003) The origin of new genes: glimpses from the young and old. Nat Rev Genet 4:865–875
Andersson DI, Jerlström-Hultqvist J, Näsvall J (2015) Evolution of new functions de novo and from preexisting genes. Cold Spring Harb Perspect Biol 7:a017996
McLysaght A, Hurst LD (2016) Open questions in the study of de novo genes: what, how and why. Nat Rev Genet 17:567–578
Schlötterer C (2015) Genes from scratch—the evolutionary fate of de novo genes. Trends Genet 31:215–219
McLysaght A, Guerzoni D (2015) New genes from non-coding sequence: the role of de novo protein-coding genes in eukaryotic evolutionary innovation. Philos Trans R Soc Lond B Biol Sci 370:20140332
Li D, Dong Y, Jiang Y et al (2010) A de novo originated gene depresses budding yeast mating pathway and is repressed by the protein encoded by its antisense strand. Cell Res 20:408–420
Vakirlis N, Sarilar V, Drillon G et al (2016) Reconstruction of ancestral chromosome architecture and gene repertoire reveals principles of genome evolution in a model yeast genus. Genome Res 26:918–932
Tautz D, Domazet-Lošo T (2011) The evolutionary origin of orphan genes. Nat Rev Genet 12:692–702
Cai J, Zhao R, Jiang H et al (2008) De novo origination of a new protein-coding gene in Saccharomyces cerevisiae. Genetics 179:487–496
Heinen TJAJ, Staubach F, Häming D et al (2009) Emergence of a new gene from an intergenic region. Curr Biol 19:1527–1531
Knowles DG, McLysaght A (2009) Recent de novo origin of human protein-coding genes. Genome Res 9:1752–1759
Levine MT, Jones CD, Kern AD et al (2006) Novel genes derived from noncoding DNA in Drosophila melanogaster are frequently X-linked and exhibit testis-biased expression. Proc Natl Acad Sci 103:9935–9939
Carvunis A-R, Rolland T, Wapinski I et al (2012) Proto-genes and de novo gene birth. Nature 487:370–374
Domazet-Lošo T, Carvunis A-R, Albà MM et al (2017) No evidence for phylostratigraphic bias impacting inferences on patterns of gene emergence and evolution. Mol Biol Evol 34:843–856
Moyers BA, Zhang J (2014) Phylostratigraphic bias creates spurious patterns of genome evolution. Mol Biol Evol 32:258–267
Moyers BA, Zhang J (2016) Evaluating phylostratigraphic evidence for widespread de novo gene birth in genome evolution. Mol Biol Evol 33:1245–1256
Vakirlis N, Hebert AS, Opulente DA et al (2018) A molecular portrait of de novo genes in yeast. Mol Biol Evol 35:631–645
Altschul SF, Madden TL, Schäffer AA et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Pearson WR, Wood T, Zhang Z et al (1997) Comparison of DNA sequences with protein sequences. Genomics 46:24–36
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Löytynoja A, Goldman N (2008) Phylogeny-aware gap placement prevents errors in sequence alignment and evolutionary analysis. Science 320:1632–1635
She R, Chu JS-C, Wang K et al (2009) GenBlastA: enabling BLAST to identify homologous gene sequences. Genome Res 19:143–149
Guindon S, Delsuc F, Dufayard J-F et al (2009) Estimating maximum likelihood phylogenies with PhyML. Methods Mol Biol 537:113–137
Frith MC (2011) A new repeat-masking method enables specific detection of homologous sequences. Nucleic Acids Res 39:e23–e23
Clark MB, Amaral PP, Schlesinger FJ et al (2011) The reality of pervasive transcription. PLoS Biol 9:e1000625
Ingolia NT, Lareau LF, Weissman JS (2011) Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147:789–802
Chen T, Zhao J, Ma J et al (2015) Web resources for mass spectrometry-based proteomics. Genomics Proteomics Bioinformatics 13:36–39
Wang H, Wang Y, **e Z (2017) Computational resources for ribosome profiling: from database to Web server and software. Brief Bioinform. https://doi.org/10.1093/bib/bbx093
Ruiz-Orera J, Messeguer X, Subirana JA et al (2014) Long non-coding RNAs as a source of new peptides. Elife 3:e03523
Scannell DR, Zill OA, Rokas A et al (2011) The awesome power of yeast evolutionary genetics: new genome sequences and strain resources for the Saccharomyces sensu stricto genus. G3 (Bethesda) 1:11–25
Wang L, Park HJ, Dasari S et al (2013) CPAT: coding-potential assessment tool using an alignment-free logistic regression model. Nucleic Acids Res 41:e74
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Vakirlis, N., McLysaght, A. (2019). Computational Prediction of De Novo Emerged Protein-Coding Genes. In: Sikosek, T. (eds) Computational Methods in Protein Evolution. Methods in Molecular Biology, vol 1851. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-8736-8_4
Download citation
DOI: https://doi.org/10.1007/978-1-4939-8736-8_4
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-8735-1
Online ISBN: 978-1-4939-8736-8
eBook Packages: Springer Protocols