Abstract
Background
Acquired resistance to sorafenib greatly limits its therapeutic efficiency in the treatment of hepatocellular carcinoma (HCC). Increasing evidence indicates that long noncoding RNAs (lncRNAs) play important roles in the resistance to anti-cancer drugs. The present study aims to explore the involvement of lncRNA SNHG1 (small nucleolar RNA host gene 1) in sorafenib resistance and how SNHG1 is associated with overexpressed microRNA-21 (miR-21) and the activated Akt pathway, which have been demonstrated to mediate this resistance in HCC cells.
Methods
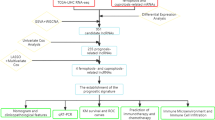
Sorafenib-resistant HCC (SR-HCC) cells were generated and their sorafenib-resistant properties were confirmed by cell viability and apoptosis assays. Potential lncRNAs were screened by using multiple bioinformatics analyses and databases. The expression of genes and proteins was detected by qRT-PCR, Western blot and in situ hybridization. Gene silencing was achieved by specific siRNA or lncRNA Smart Silencer. The effects of anti-SNHG1 were evaluated in vitro and in experimental animals by using quantitative measures of cell proliferation, apoptosis and autophagy. The binding sites of miR-21 and SNHG1 were predicted by using the RNAhybrid algorithm and their interaction was verified by luciferase assays.
Results
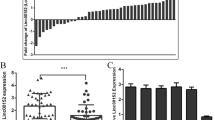
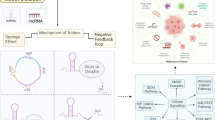
The Akt pathway was highly activated by overexpressed miR-21 in SR-HCC cells compared with parental HCC cells. Among ten screened candidates, SNHG1 showed the largest folds of alteration between SR-HCC and parental cells and between vehicle- and sorafenib-treated cells. Overexpressed SNHG1 contributes to sorafenib resistance by activating the Akt pathway via regulating SLC3A2. Depletion of SNHG1 enhanced the efficacy of sorafenib to induce apoptosis and autophagy of SR-HCC cells by inhibiting the activation of Akt pathway. Sorafenib induced translocation of miR-21 to the nucleus, where it promoted the expression of SNHG1, resulting in upregulation of SLC3A2, leading to the activation of Akt pathway. In contrast, SNHG1 was shown to have little effect on the expression of miR-21, which downregulated the expression of PTEN, leading to the activation of the Akt pathway independently of SNHG1.
Conclusions
The present study has demonstrated that lncRNA SNHG1 contributes to sorafenib resistance by activating the Akt pathway and its nuclear expression is promoted by miR-21, whose nuclear translocation is induced by sorafenib. These results indicate that SNHG1 may represent a potentially valuable target for overcoming sorafenib resistance for HCC.
Similar content being viewed by others
Background
Hepatocellular carcinoma (HCC) has the third highest rate of mortality worldwide [1]. It manifests frequently in patients with cirrhosis and liver dysfunction, limiting interventions with systemic cytotoxic drugs [2]. To date, four drugs targeting tyrosine kinases have been approved for the treatment of HCC. Lenvatinib has recently been used as a first-line drug [3], and regorafenib and cabozantinib have been approved as second-line drugs after the failure of sorafenib [4], but their use is not widely accepted. Sorafenib is the first approved systemic drug and continues to play the most important role in the management of advanced HCC [5]. However, the known resistance to sorafenib greatly minimizes its therapeutic benefits. Therefore, exploring the mechanisms for sorafenib resistance and seeking novel molecular targets are urgently required.
We have previously demonstrated that the Akt pathway is highly activated in sorafenib-resistant HCC (SR-HCC) cells and the inhibition of Akt could reverse this resistance by switching autophagy from a cytoprotective role to a death-promoting mechanism [6]. MicroRNAs (miRNAs) are a group of small noncoding RNAs, which regulate multiple cellular functions and have emerged as potential targets in the anti-cancer campaign [7]. In exploring the miRNA-related mechanisms that regulate the activation of Akt involved in sorafenib resistance, we found that miR-21 was highly expressed in SC-HCC cells and was able to activate the Akt pathway by dysregulating phosphatase and tensin homolog (PTEN) [8].
Long non-coding RNAs (lncRNAs) play crucial roles in controlling gene expression involving numerous biological processes in many diseases [9]. One of the mechanisms of their action involves the formation of regulatory networks with other RNA species, such as miRNAs and mRNAs [10, 11]. Several lncRNAs have been shown to participate in anti-cancer drug resistance [12]. For instance, lncARSR secreted by exosomes regulated sunitinib resistance by acting as competing endogenous RNAs for miR-34 and miR-449 in renal cancer [13], LncRNA MIR-100HG was involved in cetuximab resistance by regulating the wnt/β-catenin pathway via miR-100 and miR-125b in colorectal carcinoma [14], and LncRNA HULC reduced the sensitivity to cytotoxic drugs by stabilizing Sirt1-induced autophagy in HCC cells [37]. The best characterized mechanism for the interaction of lncRNAs and miRNAs pertains to the role of competitive endogenous lncRNAs, which serve as “sponges” for miRNAs [38]. On the other hand, miRNAs can regulate lncRNAs via a RNA-induced silencing complex, which often takes places in the cytoplasm [39]. In the canonical way, miRNAs are egressed from the nucleus and eventually mature in the cytoplasm, where they exert the post-transcriptional gene regulatory roles [40]. However, recent evidence suggests that certain mature cytoplasmic miRNAs can shuttle back to the nucleus, where they regulate gene expression [41,42,43]. The presence of mature miRNAs in the nucleus has recently been confirmed as a general phenomenon in mammalian cells [39, 41]. MiR-21 is the first miRNA with both a cytoplasmic and nuclear location in HeLa cells, with approximately 20% of mature miR-21 in the nucleus [44]. In agreement, the present study shows that 34.7% of miR-21 resides in nuclei of untreated Huh7 cells, which is supported by our microscopic evidence from in situ hybridization analysis. More interestingly, sorafenib induced the nuclear translocation of miR-21, which contributes to sorafenib resistance by upregulating SNHG1 in HCC cells.
MiR-21 is one of the few widely studied miRNAs expressed in many types of malignances including HCC, and plays a central role in cancer progression [27, 45]. The proposed “onco-miR addition” concept has further strengthened its role in carcinogenesis [45]. MiR-21 has also been identified as a key miRNA associated with drug insensitivity including the resistance to sorafenib [46]. In the conventional way, miR-21 exhibits its function by regulating downstream mRNAs [47]. The present study provides strong evidence that miR-21 could re-enter the nucleus, where it binds to the enhancer/promoter region of lncRNA SNHG1, leading to enhanced expression of SNHG1. This novel regulatory function of miRNAs on lncRNAs has been demonstrated in a previously reported study, where mature miR-140 trafficking in the nucleus can interact with lncRNA NEAT1, resulting in an increase in NEAT1 expression [48].
One weak point of the present study is that the results obtained in cell and animal experiments have not been validated in tumor tissues collected from HCC patients who have developed sorafenib resistance. The guideline of American Association for the Study of Liver Diseases (AASLD) indicates that sorafenib therapy is only approved for treating advanced HCC patients, judged clinically unsuitable for curative therapies including surgery and liver transplantation [49]. Due to the anatomic feature, clinical HCC tissues can be obtained by laparotomy, laparoscopy or fine-needle biopsy. However, the chance for harvesting SR-HCC tissues from late-staged patients is rare because such patients are unable to benefit from these invasive procedures in the view of stringent human ethic requirements, the informed consent and potentially risky complications. Postmortem may be another possible procedure for collecting SR-HCC tissues, but the quality of such tissues is hard to control. However, further validation in clinical SR-HCC tissues should be investigated in the future when such tissues are available since it will increase the clinical relevance and translation of the present results.
Conclusions
The present study has demonstrated that lncRNA SNHG1 contributes to sorafenib resistance by activating the Akt pathway through upregulating SLC3A2 and its nuclear overexpression is promoted by miR-21 in HCC cells. Depletion of SNHG1 inhibited the activation of the Akt pathway, enhancing the efficacy of sorafenib in suppressing SR-HCC cells by promoting apoptosis and autophagy in cultured cells and in animal experiments. Despite the fact that two putative miR-21 and SNHG1 binding sites exist, SNHG1 was shown to be incapable of regulating miRNA expression, which is different from the well characterized role of lncRNAs as sponges for miRNAs. On the other hand, miR-21 did not exhibit a negative regulatory effect on SNHG1, which usually takes place in the cytoplasm via a miRNA-induced silencing complex. However, miR-21 was able to promote the expression of SNHG1 in the nucleus, to which the shuttling of miR-21 was induced by sorafenib in HCC cells. The present study has uncovered a novel regulatory pathway by which miR-21 promotes the expression of SNHG1, leading to the activation of Akt pathway. Our results also suggest that SNHG1 may represent a potentially valuable target for overcoming sorafenib resistance in HCC.
Abbreviations
- 4EBP1:
-
Eukaryotic translation initiation factor 4E-binding protein 1
- ANOVA:
-
Analysis of variance
- GSK-3β:
-
glycogen synthase kinase 3β
- HCC:
-
Hepatocellular carcinoma
- LC3:
-
Microtubule-associated protein 1 light chain 3
- lncRNA:
-
Long non-coding RNA
- miR-21:
-
microRNA-21
- mTOR:
-
mammalian target of rapamycin
- NC:
-
Negative control
- PTEN:
-
Phosphatase and tensin homolog
- qRT-PCR:
-
quantitative reverse-transcription polymerase chain reaction
- S6K:
-
ribosomal protein S6 kinase
- SLC3A2:
-
solute carrier family 3 member 2
- SNHG1:
-
small nucleolar RNA host gene 1
- SR-HCC:
-
sorafenib-resistant HCC
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018;68(1):7–30.
Forner A, Reig M, Bruix J. Hepatocellular carcinoma. Lancet. 2018;391(10127):1301-1314.
Kudo M. Lenvatinib may drastically change the treatment landscape of hepatocellular carcinoma. Liver Cancer. 2018;7(1):1–19.
Daher S, et al. Current and future treatment of hepatocellular carcinoma: an updated comprehensive review. J Clin Transl Hepatol. 2018;6(1):69–78.
Llovet JM, et al. Sorafenib in advanced hepatocellular carcinoma. N Engl J Med. 2008;359(4):378–90.
Zhai B, et al. Inhibition of Akt reverses the acquired resistance to sorafenib by switching protective autophagy to autophagic cell death in hepatocellular carcinoma. Mol Cancer Ther. 2014;13(6):1589–98.
Lin S, Gregory RI. MicroRNA biogenesis pathways in cancer. Nat Rev Cancer. 2015;15(6):321–33.
He C, et al. MiR-21 mediates sorafenib resistance of hepatocellular carcinoma cells by inhibiting autophagy via the PTEN/Akt pathway. Oncotarget. 2015;6(30):28867–81.
Guttman M, Rinn JL. Modular regulatory principles of large non-coding RNAs. Nature. 2012;482(7385):339–46.
Alfarouk KO. Tumor metabolism, cancer cell transporters, and microenvironmental resistance. J Enzyme Inhib Med Chem. 2016;31(6):859–66.
Fatica A, Bozzoni I. Long non-coding RNAs: new players in cell differentiation and development. Nat Rev Genet. 2014;15(1):7–21.
Majidinia M, Yousefi B. Long non-coding RNAs in cancer drug resistance development. DNA Repair (Amst). 2016;45:25–33.
Qu L, et al. Exosome-transmitted lncARSR promotes Sunitinib resistance in renal Cancer by acting as a competing endogenous RNA. Cancer Cell. 2016;29(5):653–68.
Lu Y, et al. lncRNA MIR100HG-derived miR-100 and miR-125b mediate cetuximab resistance via Wnt/beta-catenin signaling. Nat Med. 2017;23(11):1331–41.
**ong H, et al. LncRNA HULC triggers autophagy via stabilizing Sirt1 and attenuates the chemosensitivity of HCC cells. Oncogene. 2017;36(25):3528–40.
Shen Y, et al. Nuclear retention of the lncRNA SNHG1 by doxorubicin attenuates hnRNPC-p53 protein interactions. EMBO Rep. 2017;18(4):536–48.
Lu Q, et al. Long noncoding RNA SNHG1 promotes non-small cell lung cancer progression by up-regulating MTDH via sponging miR-145-5p. FASEB J. 2018;32(7):3957–67.
Zhu Y, et al. Up-regulation of lncRNA SNHG1 indicates poor prognosis and promotes cell proliferation and metastasis of colorectal cancer by activation of the Wnt/beta-catenin signaling pathway. Oncotarget. 2017;8(67):111715–27.
Zhang M, et al. Long noncoding RNA SNHG1 predicts a poor prognosis and promotes hepatocellular carcinoma tumorigenesis. Biomed Pharmacother. 2016;80:73–9.
Sun Y, et al. The long noncoding RNA SNHG1 promotes tumor growth through regulating transcription of both local and distal genes. Oncogene. 2017;36(49):6774–83.
Piguet AC, et al. Everolimus augments the effects of sorafenib in a syngeneic orthotopic model of hepatocellular carcinoma. Mol Cancer Ther. 2011;10(6):1007–17.
Zhang PF, et al. Galectin-1 induces hepatocellular carcinoma EMT and sorafenib resistance by activating FAK/PI3K/AKT signaling. Cell Death Dis. 2016;7:e2201.
Han P, et al. Dual inhibition of Akt and c-met as a second-line therapy following acquired resistance to sorafenib in hepatocellular carcinoma cells. Mol Oncol. 2017;11(3):320–34.
Jiang W, et al. Sodium orthovanadate overcomes sorafenib resistance of hepatocellular carcinoma cells by inhibiting Na(+)/K(+)-ATPase activity and hypoxia-inducible pathways. Sci Rep. 2018;8(1):9706.
de Planell-Saguer M, Rodicio MC, Mourelatos Z. Rapid in situ codetection of noncoding RNAs and proteins in cells and formalin-fixed paraffin-embedded tissue sections without protease treatment. Nat Protoc. 2010;5(6):1061–73.
Wu Y, et al. Sodium orthovanadate inhibits growth of human hepatocellular carcinoma cells in vitro and in an orthotopic model in vivo. Cancer Lett. 2014;351(1):108–16.
Meng F, et al. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology. 2007;133(2):647–58.
Yamamura S, et al. Interaction and cross-talk between non-coding RNAs. Cell Mol Life Sci. 2018;75(3):467–84.
Manning BD, Toker A. AKT/PKB signaling: navigating the network. Cell. 2017;169(3):381–405.
Chen Y, et al. MiRNA153 reduces effects of chemotherapeutic agents or small molecular kinase inhibitor in HCC cells. Curr Cancer Drug Targets. 2015;15(3):176–87.
Poettler M, et al. CD98hc (SLC3A2) drives integrin-dependent renal cancer cell behavior. Mol Cancer. 2013;12:169.
Song MS, Salmena L, Pandolfi PP. The functions and regulation of the PTEN tumour suppressor. Nat Rev Mol Cell Biol. 2012;13(5):283–96.
Kim SE, Overholtzer M. Autophagy proteins regulate cell engulfment mechanisms that participate in cancer. Semin Cancer Biol. 2013;23(5):329–36.
Sini P, et al. Simultaneous inhibition of mTORC1 and mTORC2 by mTOR kinase inhibitor AZD8055 induces autophagy and cell death in cancer cells. Autophagy. 2010;6(4):553–4.
Niu L, et al. New insights into sorafenib resistance in hepatocellular carcinoma: responsible mechanisms and promising strategies. Biochim Biophys Acta Rev Cancer. 2017;1868(2):564–70.
Prieto-Dominguez N, et al. Modulation of autophagy by Sorafenib: effects on treatment response. Front Pharmacol. 2016;7:151.
Engreitz JM, et al. RNA-RNA interactions enable specific targeting of noncoding RNAs to nascent pre-mRNAs and chromatin sites. Cell. 2014;159(1):188–99.
Cheng EC, Lin H. Repressing the repressor: a lincRNA as a MicroRNA sponge in embryonic stem cell self-renewal. Dev Cell. 2013;25(1):1–2.
Liang H, et al. Nuclear microRNAs and their unconventional role in regulating non-coding RNAs. Protein Cell. 2013;4(5):325–30.
Ambros V. The functions of animal microRNAs. Nature. 2004;431(7006):350–5.
Jeffries CD, Fried HM, Perkins DO. Nuclear and cytoplasmic localization of neural stem cell microRNAs. Rna. 2011;17(4):675–86.
Hwang HW, Wentzel EA, Mendell JT. A hexanucleotide element directs microRNA nuclear import. Science. 2007;315(5808):97–100.
Castanotto D, et al. CRM1 mediates nuclear-cytoplasmic shuttling of mature microRNAs. Proc Natl Acad Sci U S A. 2009;106(51):21655–9.
Meister G, et al. Human Argonaute2 mediates RNA cleavage targeted by miRNAs and siRNAs. Mol Cell. 2004;15(2):185–97.
Medina PP, Nolde M, Slack FJ. OncomiR addiction in an in vivo model of microRNA-21-induced pre-B-cell lymphoma. Nature. 2010;467(7311):86–90.
Hong L, et al. MicroRNA-21: a therapeutic target for reversing drug resistance in cancer. Expert Opin Ther Targets. 2013;17(9):1073–80.
Vidigal JA, Ventura A. The biological functions of miRNAs: lessons from in vivo studies. Trends Cell Biol. 2015;25(3):137–47.
Gernapudi R, et al. MicroRNA 140 promotes expression of long noncoding RNA NEAT1 in Adipogenesis. Mol Cell Biol. 2016;36(1):30–8.
Bruix J, Sherman M. Management of hepatocellular carcinoma: an update. Hepatology. 2011;53(3):1020–2.
Acknowledgements
We thank Dr. Shiva Reddy from the University of Auckland (New Zealand) for revising and editing the manuscript.
Funding
This study was supported by grants from the National Key Research and Development Program of China (2017YFC1308602), the National Natural Scientific Foundation of China (81472321, 81401975 and 81703055), a Fundamental Research Fund for the Provincial Universities in Heilongjiang Province (2017LCZX06), a Supportive Fund by Heilongjiang Provincial Department of Science and Technology (GX18C010), and Scientific Foundation of Heilongjiang Provincial Health and Family Planning Commission (2016-113), in China.
Availability of data and materials
The datasets used and analyzed during the current study are available from the corresponding author on reasonable request.
Author information
Authors and Affiliations
Contributions
WL performed cell experiments, analyzed the data and drafted the manuscript. XD performed animal experiments and participated in cell culture and assays. CH participated in data analysis and revising the manuscript. GT participated in animal experiments and RT-PCR. ZL and BZ participated in the immunohistochemistry and statistical analysis. JF assisted in collecting and processing data. XJ assisted in cell and animal experiments. CL and HJ contributed in study design and manuscript revision. XS designed the project, supervised the study and finalized the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The animal experiments had been approved (permit SYXK20020009) by the Animal Ethics Committee of Harbin Medical University (Harbin, China).
Consent for publication
All authors agree on publication of the results of the present manuscript.
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Additional information
The original online version of this article was revised: the authors identified some minor errors in Figs. 3a and 4f.
Additional files
Additional file 1:
Supplementary Materials and Methods. (PDF 149 kb)
Additional file 2:
Supplementary tables and figures. (PDF 149 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Li, W., Dong, X., He, C. et al. LncRNA SNHG1 contributes to sorafenib resistance by activating the Akt pathway and is positively regulated by miR-21 in hepatocellular carcinoma cells. J Exp Clin Cancer Res 38, 183 (2019). https://doi.org/10.1186/s13046-019-1177-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13046-019-1177-0