Abstract
Epidermal growth factor receptor (EGFR), the receptor for members of the epidermal growth factor family, regulates cell proliferation and signal transduction; moreover, EGFR is related to the inhibition of tumor cell proliferation, angiogenesis, invasion, metastasis, and apoptosis. Therefore, EGFR has become an important target for the treatment of cancer, including non-small cell lung cancer, head and neck cancer, breast cancer, glioma, cervical cancer, and bladder cancer. First- to third-generation EGFR inhibitors have shown considerable efficacy and have significantly improved disease prognosis. However, most patients develop drug resistance after treatment. The challenge of overcoming intrinsic and acquired resistance in primary and recurrent cancer mediated by EGFR mutations is thus driving the search for alternative strategies in the design of new therapeutic agents. In view of resistance to third-generation inhibitors, understanding the intricate mechanisms of resistance will offer insight for the development of more advanced targeted therapies. In this review, we discuss the molecular mechanisms of resistance to third-generation EGFR inhibitors and review recent strategies for overcoming resistance, new challenges, and future development directions.
Similar content being viewed by others
Introduction
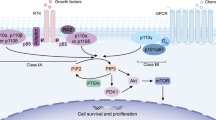
Epidermal growth factor receptor (EGFR) is a member of the receptor tyrosine kinase (RTK) superfamily that consists of exon boundaries and associated extracellular, transmembrane, and intracellular protein domains. EGFR is involved in multiple signaling pathways and regulates numerous cell functions (Fig. 1A). This transmembrane glycoprotein is composed of a cysteine-rich extracellular ligand binding domain, hydrophobic transmembrane domain, cytoplasmic RTK domain, and C-terminal domain. The RTK domain contains an N-lobe consisting of five β-sheet strands and one αC helix and a C-lobe containing the main helices of a highly flexible activation loop (A-loop) [1]. The deep cleft at the junction of these two lobes forms the binding pocket for the adenine ring of ATP. The conformation of three conserved structural elements, namely the Asp-Phe-Gly (DFG) motif, αC helix, and A-loop, critically regulates the activation or inactivation of the catalytic domain. When EGFR is in the active state, the important catalytic residue D855 is located in the ATP binding site, stabilizing the ATP-loaded complex (DFG-in) and αC helix (αC-in). In the inactive state, EGFR forms a Src-like structure, including a closed A-loop, αC-out, and DFG-in [2]. (Fig. 1B). EGFR can dimerize upon binding by ligands, such as amphiregulin, β-cytokines, epidermal growth factor (EGF), heparin-binding EGF-like growth factor (HB-EGF), and transforming growth factor (TGF). The activation of the intracellular tyrosine kinase domain and autophosphorylation, which initiates the Ras/RAF/MEK, signal transducer and activator of transcription (STAT), PI3K/AKT/mTOR and other downstream signaling pathways, are closely related to embryonic development and stem cell division [2,3,4]. Overexpression of wild-type (WT) EGFR protein with or without EGFR gene amplification or a kinase-activating mutation further enhances cell proliferation, migration, survival, and antiapoptotic responses through signaling cascades, and these processes are closely related to the occurrence and development of many types of epithelial-derived cancer, such as non-small cell lung cancer (NSCLC), breast cancer, glioma, head and neck cancer, cervical cancer, and bladder cancer. Among these cancers, lung cancer appears to be the most common and has the characteristics of aberrant proliferation, metastasis, and drug resistance [5,6,7,8]. Thus, EGFR has become a promising target for anticancer drug design and development. EGFR tyrosine kinase inhibitors (EGFR-TKIs) have achieved remarkable results in the clinic [9]. However, most patients develop acquired drug resistance to first- and second-generation EGFR-TKIs after 1–2 years. The mechanism of drug resistance for nearly half of cases relates to the T790M mutation. Third-generation EGFR-TKIs that target EGFR-TKI-sensitive mutations and the T790M mutation have been developed [10].
Structure and functions of EGFR. A EGFR exon boundaries and associated extracellular, transmembrane, and intracellular protein domains. EGFR is involved in multiple signaling pathways and regulates numerous cell functions. B The tyrosine kinase domain of EGFR and the activation or inactivation of the catalytic domain. C EGFR domains and the molecular mechanisms of acquired resistance. The intracellular domain contains a juxtamembrane domain, tyrosine kinase domain, and multiple C-terminal tyrosine residues. Multiple mutations within the tyrosine kinase domain are associated with resistance and sensitivity to EGFR-TKIs
Unfortunately, drug resistance caused by less-common mutations in the EGFR gene and components of signal transduction pathways continues to emerge. In addition to common secondary (T790M) and tertiary (C797S) mutations, other EGFR mutations (such as the L718Q, L796S, and L792H mutations and the exon 20 insertion), MET amplification, phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha (PIK3CA) mutations, HER2 amplification, oncogene fusions, and alterations in cell cycle-related genes have been observed [11] (Fig. 1C). There is an urgent need for better strategies to combat the inevitable molecular-targeted drug resistance associated with third-generation inhibitors. This review aims to provide a comprehensive overview of the mechanisms of resistance to third-generation EGFR-TKIs and to explore new insights and strategies for overcoming acquired resistance.
Third-generation EGFR-TKIs and drug resistance mechanisms
The development of third-generation EGFR-TKIs
The first-generation EGFR-TKIs form hydrogen bonds with Met793 in the ATP binding pocket of EGFR and reversibly compete with ATP for binding. Drug resistance occurs due to the EGFR T790M mutation (Thr790 in the hydrophobic ATP binding site encoded on exon 20 is replaced by methionine), subclonal selection (of a genetically resistant clone), and rare EGFR mutations (such as G719X, S768I, and L861Q). Thereafter, the development of second-generation EGFR-TKIs was reported; these inhibitors have the same quinazoline scaffold as first-generation EGFR-TKIs, but the side chain can irreversibly bind to Cys797 to inhibit the tyrosine kinase activity of EGFR. For example, the anilinoquinazoline derivative forms hydrogen bonds with the backbone of Met793 in the hinge region and interacts with the hydrophobic region. The acrylamide group binds covalently to Cys797 in the active conformation of EGFR, the furanyl group is exposed to solvent, and the 3-chloro-4-fluorophenyl group is situated next to the gatekeeper residue [12,13,14]. However, mutations such as T790M still emerge upon treatment with second-generation EGFR-TKIs, which have limited selectivity against WT-EGFR, resulting in serious side effects [15]. Fortunately, third-generation covalent inhibitors that bind irreversibly to the target and are mutation-selective have been developed. These compounds were designed based on a new aminopyrimidine scaffold and show preferable biological activities [16]. Early clinical trials have proven that these third-generation EGFR-TKIs are effective in patients with double-mutated tumors (EGFR L858R/T790M or ex19del/T790M) and have high selectivity for mutant EGFR, thereby eliminating the side effects in the skin and gastrointestinal system associated with the nonselective inhibition of WT-EGFR [17]. For example, the crystal structures of rociletinib (CO-1686) in complex with EGFR T790M and EGFR L858R have been published; in EGFR T790M, the anilinopyrimidine group of rociletinib forms hydrogen bonds with the Met793 amide and the carbonyl backbone, whereas in EGFR L858R, hydrophobic interactions between rociletinib and the protein were due to hydrogen bonds between nitrogens in the pyrimidine group and between the fluoromethyl and Thr790. In addition, the acrylamide group in rociletinib covalently binds to Cys797 in the DFG-in/αC-in active conformations [18]. The specificity for EGFR T790M may stem from hydrophobic interactions between the large methionine in mutant EGFR and pyrimidines. Drugs that have been approved for marketing include osimertinib (US), almonertinib (China), lazertinib (South Korea), and alflutinib (China) (Fig. 2).
Mechanisms of resistance to third-generation EGFR-TKIs
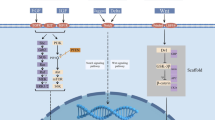
Due to the covalent bond between the acrylamide (Michael acceptor) of third-generation EGFR-TKIs and the active thiol in the EGFR kinase domain, highly selective inhibitory activity has been achieved by targeting Cys797 and irreversible binding EGFR; thus, these compounds show excellent antitumor activity. Targeted therapy for patients with EGFR T790M and EGFR-activating mutations showed good efficacy in both first- and second-line settings. In patients who developed resistance to third-generation EGFR-TKIs as first-line therapy, genetic changes such as MET amplification, EGFR C797X mutation, PIK3CA amplification and mutation, HER2 amplification and mutation, K-RAS mutation, and BRAF mutation, as well as changes in cell cycle-related genes and oncogene fusions, have been reported, but no T790M mutations have been detected. The mechanism of resistance to second-line therapy is more complicated. Acquisition or deletion of the T790M mutation has been detected in patients [19], and other EGFR mutations (such as L718Q, L796S, L792H, and exon 20 insertion) have also been observed (Fig. 1B). In addition, the mechanisms of acquired resistance to third-generation EGFR-TKIs include alternative pathway activation and histologic and phenotypic transformation (Fig. 3); the details will be discussed in the following sections.
Molecular mechanisms of acquired resistance. The mechanisms include target gene modification, parallel alternative pathway activation, downstream pathway activation, and histological/phenotypic transformation. Both amplification and mutation of receptor tyrosine kinases (RTKs) can induce downstream survival signaling pathways. Moreover, direct overexpression and/or mutation of components of downstream pathways can contribute to acquired resistance by promoting cancer cell survival
Primary/intrinsic resistance
The differential sensitivity of TKIs to different EGFR mutations is a cause of primary drug resistance. In NSCLC patients, the in-frame deletion of exon 19 (ex19del) and the L858R point mutation in exon 21 are the most common somatic mutations, occurring in approximately 80% of cases. During EGFR-TKI treatment, patients with longer median survival have presented with more than 20 unique deletions of exon 19. Intrinsic drug resistance can all be triggered by other nonclassical sensitizing mutations (mainly exon 20 insertion) and inherent secondary genetic changes. Drug-resistant clones (for example, T790M) may already exist within the cancer cell population, leading to drug resistance during treatment [20]. Some studies have found that in nearly 1% of lung cancer patients, 2–3 simultaneous driver mutations can be detected before treatment. Some molecular and genetic changes have been reported to relate to intrinsic drug resistance, such as the lack of K-RAS/phosphatase and tensin homolog (PTEN) expression. These preexisting molecular and genetic alterations can stimulate the Ras/Raf/MEK/ERK and PI3K/AKT downstream pathways to promote cancer progression [21].
BIM deletion polymorphism
BIM is a proapoptotic member of the B-cell lymphoma-2 (Bcl-2) family [22]. Recent studies showed that lung cancer cells with the BIM deletion polymorphism and EGFR mutation are resistant to third-generation EGFR-TKIs, suggesting that the BIM deletion polymorphism has potential as a biomarker to predict the efficacy of third-generation EGFR-TKIs in patients [22].
EGFR exon 20 insertion
The molecular mechanism of drug resistance caused by the exon 20 insertion is not fully understood. Eck et al. [23] hypothesized that this mutation prevents binding to EGFR-TKIs due to the addition of residues to the N-lobe of EGFR. The crystal structure of EGFR exon 20 with the D770_N771insNPG insertion shows an unchanged ATP binding pocket and a rigid active conformation, leading to steric hindrance of the drug binding pocket and resistance to EGFR-TKIs.
Acquired resistance
Acquired drug resistance refers to the process by which tumor cells with prior sensitivity to treatment circumvent the inhibitory effects of drugs by changing their metabolic pathways. The mechanisms of acquired resistance to third-generation EGFR-TKIs can be divided into EGFR-dependent resistance and EGFR-independent resistance [24].
EGFR-dependent drug resistance mechanisms
Reappearance of an EGFR mutation
C797S mutation
One point mutation of EGFR (C797S) involves the replacement of Cys797 within the ATP binding site (exon 20) with serine [25]. Osimertinib binds covalently and irreversibly to EGFR T790M by interacting with Cys797. When the C797S mutation occurs, the osimertinib binding efficiency decreases [10], resulting in tumor resistance to all third-generation EGFR-TKIs.
G796R/D mutation
The G796R mutation has been detected in cancer patients who received treatment with a third-generation EGFR-TKI. Molecular docking predictions revealed that G796R sterically hinders the covalent binding of osimertinib. Because the bulky side chain and hydrophilic group hinder the binding of osimertinib to the hydrophobic region, the change in binding energy renders binding unfavorable. Compared with samples containing the double-mutant EGFR L858R/T790M, those harboring the triple-mutant EGFR L858R/T790M/G796R are 110 times more resistant to osimertinib [26]. G796D was reported for the first time in osimertinib-resistant NSCLC patients. In vitro studies have shown that the G796D mutation causes a 50-fold increase in the growth inhibitory 50% (GI50) value of osimertinib. Structural modeling showed that the side chain of the mutated G796D residue collides with the surface of osimertinib, resulting in steric hindrance and energy repulsion and ultimately the loss of binding affinity [27].
L792 mutation
The mutations at Leu792 include L792F, L792Y, and L792H. Structural prediction revealed that these mutations introduce a benzene ring or imidazole ring to the side chain of the residue at 792, which spatially disrupts the orientation of osimertinib, thereby potentially affecting the binding of osimertinib to the EGFR ATP binding site [28].
M766Q mutation
The homology simulation with the T790M and M766Q double mutant showed that M766Q seems to position T790M in the inhibitor binding site, thereby weakening osimertinib binding [29].
Mutations in exon 18
EGFR L718Q/V
EGFR L718Q was reported for the first time in a cell model of resistance to third-generation EGFR-TKIs. Subsequent studies have shown that NSCLC with EGFR L858R/T790M/L718Q is resistant to all EGFR-TKIs, but that with only L858R/L718Q remains sensitive to afatinib [30]. The crystallographic model revealed that the L718Q mutation reduces the efficiency of the formation of covalent bonds between the acrylamide warhead and the Cys797 thiol group, thus interfering with the irreversible binding of osimertinib [31, 32]. In addition, L718V resistance mutations in the kinase domain of EGFR have been detected, and these may interfere with the binding of osimertinib to the kinase domain [33]. Of note, EGFR L718Q/V is still sensitive to afatinib [32].
EGFR G724S
The G724S mutation in the ATP binding loop enriches this loop in glycine, which can lead to the development of resistance to EGFR-TKIs by changing the protein structure, enhancing ATP affinity, and stabilizing activating mutations [34]. However, this mutation does not lead to resistance to second-generation EGFR inhibitors [34].
Compound mutations
A compound mutation refers to the simultaneous detection of two or more different types of EGFR mutations in patient cancer cells [35]. The impact of compound mutations on EGFR-TKI sensitivity is listed in descending order: double classic mutations, compound mutations involving classic mutations and rare mutations, and compound mutations of only rare mutations [36, 37]. These EGFR mutations caused by treatment with third-generation EGFR-TKIs confer resistance to irreversible pyrimidine TKIs but not to quinazoline EGFR inhibitors [42]. After treatment with third-generation EGFR-TKIs, MET gene amplification can promote drug resistance by activating MAPK/ERK, which is independent of EGFR [43].
HER2 amplification
Hus et al. found that H1975 cells expressing HER2D16 were resistant to osimertinib in vitro. HER2D16 can form a heterodimer with EGFR or a disulfide homodimer, which activates downstream signaling to achieve resistance to osimertinib [44]. HER2D16-driven drug resistance occurs in a manner unrelated to the kinase Src. In addition, other mutations in exon 20 of HER2 have been reported, including point mutations (such as G776C and L755S) and insertions that cause downstream activation [45, 46]. HER2 mutation occurs in approximately 2–4% of NSCLC cases, mostly in lung adenocarcinoma (LUAD) [47]. In NSCLC, HER2 oncogenic amplification occurs in approximately 3% of cases without EGFR-TKI treatment and accounts for approximately 10% of cases with EGFR-TKI resistance [48].
AXL activation
AXL is an RTK that regulates cell survival, proliferation, metastasis, and other cellular functions. Abnormalities in the AXL gene can generate acquired resistance to TKIs by activating relevant downstream signaling pathways. Osimertinib was found to trigger AXL activation by closing the negative feedback loop with SPRY4, thus triggering inherent osimertinib resistance [49].
Overexpression of HGF
Hepatocyte growth factor (HGF) is the ligand of the proto-oncogene c-Met; it can trigger MET activation through EGFR bypass signaling and induce lung cancer resistance to EGFR-TKIs. Yano et al. [50] found that high expression of HGF was related to the acquired and intrinsic drug resistance to EGFR-TKIs in patients with lung cancer. Tumor specimens from patients with acquired drug resistance showed high expression of HGF in the context of MET amplification and the T790M mutation.
Fibroblast growth factor receptor (FGFR) signaling
FGFR is a transmembrane RTK. Studies have shown that FGFR1 is amplified and fibroblast growth factor 2 (FGF2) mRNA levels are increased in patients with osimertinib resistance, suggesting that the FGFR2-FGFR1 autocrine loop may be related to drug resistance [51]. Patients with the T790M mutation have been reported to show disease progression after treatment with osimertinib and nilotinib. The FGFR3-TACC3 fusion was detected in ctDNA [52, 53]. These findings suggest that abnormalities in the FGFR signaling pathway may underlie the mechanism of acquired resistance to third-generation EGFR-TKIs.
Insulin-like growth factor receptor 1 (IGF1R)
IGF1R, a transmembrane heterotetrameric protein encoded by the gene located on chromosome 15q26.3, is involved in promoting the growth of tumor cells. Abnormal activation of IGF1R leads to EGFR-TKI resistance [54].
Aurora kinases (AURKs)
AURKs are an important category of enzymes within the serine/threonine kinase family consisting of three mammalian isoforms: Aurora kinase A (AURK A), AURK B, and AURK C [55, 56]. AURK A and AURK B are highly expressed in dividing cells and play important roles in mitotic progression. Mammalian AURK A and AURK B share approximately 71% similarity in the carboxy-terminal catalytic domain [57]. Aberrant expression of AURK A and AURK B is involved in a broad range of solid cancers and is associated with adverse prognosis and drug resistance [58, 59]. In addition, Tanaka et al. [60] reported that targeting AURK B can prevent and overcome resistance to EGFR inhibitors in lung cancer by enhancing BIM- and PUMA-mediated apoptosis.
Downstream signaling pathway activation
The activation of signaling pathways downstream of oncogenic receptors can regulate cell proliferation, cell cycle progression, and cell survival. Therefore, the direct regulation of downstream signaling pathway-related factors can lead to acquired resistance.
K-RAS mutation
An epidemiological meta-analysis found that K-RAS mutations are present in NSCLC patients, and all patients with K-RAS mutations were resistant to EGFR-TKIs [61]. K-RAS mutation is related to activation of the RAS-MAPK pathway. The common K-RAS mutations include G12S, G12D, G12A, Q61H, and A146T. Studies have found that inhibiting mutant K-RAS can reduce tumor growth and render NSCLC patients sensitive to EGFR inhibitors [62].
BRAF (v-RAF murine sarcoma viral oncogene homologue B1) mutation
BRAF is a serine/threonine protein kinase that plays a key role in the MAPK/ERK pathway, including in EGFR/RAS/RAF signal transduction. BRAF can regulate cell survival, proliferation, differentiation, and apoptosis, as well as tumor induction. Many BRAF mutations (G469A, V600E, and V599E) have been found in cancer, including lung cancer [63]. Ohashi et al. [64] reported that in patients with lung cancer, BRAF mutations can induce acquired resistance to EGFR-TKIs. Preclinical data showed that the BRAF V600E mutation has a strong association with resistance to the third-generation EGFR-TKI osimertinib in patients with T790M-mutated LUAD.
PI3K/AKT/mTOR
PIK3CA is a driver gene of LUAD. Mutation of PIK3CA can promote tumor cell invasion and increase the activity of downstream PI3Ks. Studies have shown that PIK3CA amplification or mutation (including E453K, E545K, and H1047R) may occur in patients with osimertinib resistance [52, 65]. Increased PI3K activity leads to the activation of various downstream kinases, thereby increasing PI3K/AKT/mTOR pathway activity in the absence of coupling to upstream EGFR phosphorylation.
STAT3 activation
STAT proteins, especially STAT3, are key downstream signal sensors of EGFR activation. In studies on NSCLC, Zhao et al. [66] discovered the clinical significance of JAK2/STAT3 in angiogenesis. Chaib et al. [67] found that osimertinib treatment activates not only STAT3 but also SrcYAP1 signaling, which may act downstream of IL-6 to promote disease progression.
Loss of PTEN
PTEN is a tumor suppressor gene that encodes a protein with lipid phosphatase activity and thus regulates cellular protein phosphatase activity. PTEN has dual antitumor effects and is a key component of many signaling pathways in the body. If mutation or deletion of the PTEN gene or downregulation of PTEN expression can reduce or eliminate its antitumor activity [68], loss of PTEN leads to hyperactivation of the PI3K/AKT signaling pathway and resistance to EGFR-TKIs, including osimertinib.
Hyperactivation of activated Cdc42-associated kinase 1 (ACK1)
Hyperphosphorylation of ACK1 and the subsequent activation of antiapoptotic signaling through the AKT pathway are associated with resistance to third-generation EGFR-TKIs [69].
c-Myc gene
The c-Myc gene is an important member of the MYC gene family. The c-Myc gene can induce cells to proliferate indefinitely and can promote cell division; these activities are related to the occurrence and development of various types of cancer. Studies have shown that c-Myc levels are substantially elevated in different EGFR-mutant NSCLC cell lines with acquired resistance to the third-generation EGFR-TKI osimertinib compared with the corresponding parental cell lines; moreover, these increased levels cannot be reduced by osimertinib. Consistently, c-Myc levels are elevated in the majority of EGFR-mutant NSCLC tissues from patients who relapsed on EGFR-TKI treatment compared with the corresponding baseline c-Myc levels prior to treatment [70]. These findings indicate that c-Myc mediates the therapeutic efficacy of third-generation EGFR-TKIs and the development of acquired resistance to these TKIs.
Other mechanisms
Epithelial–mesenchymal transition (EMT)
In EMT, cancer cells lose their epithelial properties through the loss of E-cadherin, leading to increased vimentin expression and transformation into a mesenchymal phenotype. A previous study found that osimertinib-resistant H1975 cells have EMT characteristics in the absence of other EGFR mutations [71]. EMT is a coordinated process involving multiple regulatory factors, such as EMT-induced transcription factors (EMT-TFs), noncoding RNAs (ncRNAs), and various extracellular signals. EMT-TFs play an important role in all stages of EMT; the most well-known EMT-TFs are members of the SNAIL, ZEB, and TWIST families. Many studies have shown that SLUG and SNAIL overexpression can induce drug resistance [72].
miRNAs and EMT
Long noncoding RNAs (lncRNAs) and microRNAs (miRNAs) play important roles in regulating EMT and TKI resistance. Although most miRNAs have been found to inhibit EMT, some have activity that promotes EMT, including miR-21 and miR-155 [73, 74]. Some miRNAs can promote TKI resistance by activating the PI3K/AKT/mTOR signaling pathway; for example, miR-21 and miR-23a can target PTEN and activate AKT, leading to resistance to EGFR-TKIs [75, 76].
Epigenetic alterations
Epigenetic modifications involved in cancer initiation and progression include changes in DNA methylation patterns and histone modifications. Epigenetic changes are common in the development and progression of lung cancer [77]. Studies have shown that epigenetic disorders can make cancer patients susceptible to acquired resistance to EGFR-TKIs [78].
Oncogene fusion
The AURA-3 and FLAURA trials showed that oncogene fusion might be one mechanism of osimertinib resistance; the identified fusions included transforming growth factor receptor (TGFR)-transforming acidic coiled-coil protein 3 (TACC3), neurotrophic receptor tyrosine kinase 1 (NTRK1)-thrombopoietin mimetic peptide 3 (TMP3), ERC1-RET, SPTBN1-ALK, coiled-coil domain-containing protein 6 (CCDC6)-RET, GOPC-ROS1, AGK-BRAF, NCOA4-RET, ESYT2-BRAF, and echinoderm microtubule-associated protein-like 4 (EML4)-ALK. Oncogene fusions can coexist with the EGFR C797S mutation, MET amplification, and BRAF mutation [79].
Cell cycle-related gene abnormalities
Recent studies have shown that changes in cell cycle-related genes, including the CDKN2A E27fs mutation, cyclin D (CCND) amplification, cyclin-dependent kinase 4/6 (CDK4/6) amplification, and cyclin E1 (CCNE1) amplification, can cause resistance to third-generation EGFR-TKIs [65].
Histologic and phenotypic transformation
Histopathological transformation to small cell lung cancer (SCLC) from NSCLC has been reported as a mechanism of acquired resistance to EGFR-TKIs in 3–15% of patients [80,81,82,83]. Transformed SCLC mainly occurs in Asian patients with adenocarcinoma harboring EGFR-TKI-sensitive mutations (such as the EGFR ex19del/T790M mutation) who are nonsmokers. The widely accepted hypothesis for this transformation posits that adenocarcinoma and SCLC originate from type II alveolar cells. RB1 and TP53 mutations might be involved in SCLC transformation but are not sufficient for the induction of complete transformation. Additional genomic alterations, including those that activate the PI3K/AKT family and downregulate NOTCH signaling and those affecting the MYC and SOX families, AKT pathway activation and other molecules, also participate in the transformation from EGFR-mutant NSCLC. However, the precise mechanisms in other cases are unclear [84]. In addition, squamous cell transformation was recently identified as a mechanism of acquired EGFR-TKI resistance that occurs in approximately 15% of patients who received osimertinib as both first- and second-line therapy. Similar to the case in SCLC transformation, the primary EGFR mutation is preserved in squamous cell transformation [85].
Immune escape
EGFR is expressed in different hematopoietic cell types, including macrophages, monocytes, and certain T-cell subsets. Therefore, it is likely that EGFR inhibitors can interfere with the function of these leukocytes. Immune checkpoint inhibitors (ICIs) have adverse effects and poor efficacy in patients with an EGFR mutation or a secondary T790M mutation, largely because of low tumor mutational burden and a noninflamed tumor microenvironment [86,87,88]. A previous study showed that secreted phosphoprotein 1 (SPP1) promotes macrophage M2 polarization and PD-L1 expression in LUAD, which may influence the response to immunotherapy. SPP1 levels might be a useful marker of immunosuppression in patients with an EGFR mutation and could provide therapeutic insight [207]. It is a promising target for the design of selective conformationally restricted drugs, with great potential in terms of affinity, efficacy, and selectivity.
DZ-SIM inhibitors
In addition, researchers found that a group of near-infrared heptamethine carbocyanine (DZ) fluorescent dyes, the prototype of which is heptamethylamine carbocyanine dye (IR-783) (104) (Fig. 14), have tumor-targeting activity through differentially expressed organic anion transport peptides on cancer cells [208]. This group of organic dyes can specifically deliver therapeutic payloads to tumor cells in the form of chemical conjugates. DZ-SIM was preliminarily synthesized; SIM specifically targets 3-hydroxy-3-methylglutaryl-CoA reductase (HMGCR) in the endoplasmic reticulum. After specific uptake by tumor cells, DZ-SIM was enriched in subcellular organelles (including mitochondria and lysosomes). NSCLC cells were killed by mitochondrial damage, which mainly led to cytochrome C release into the cytoplasm, thereby activating the caspase-3-dependent apoptosis cascade. DZ-SIM inhibited the formation of cancer cell colonies resistant to first-generation (H1650 and H1975) and third-generation EGFR-TKIs (PC9AR), and most IC50 values were lower than 10 μmol/L. DZ-SIM represents a promising new therapy to overcome drug resistance in patients with EGFR-mutant disease.
Selection of individualized combination therapy
For patients who experience SCLC transformation, chemotherapy after the development of osimertinib resistance is an option. Research has shown that patients with transformation to SCLC have higher response rates to etoposide, cisplatin, and paclitaxel. For patients with unclear resistance mechanisms, chemotherapy is still a treatment option. If the patient is asymptomatic or has symptomatic local progression, osimertinib can be combined with local treatment according to National Comprehensive Cancer Network (NCCN) guidelines. Carboplatin, paclitaxel, bevacizumab, and atezolizumab (anti-PD-L1 antibody) are also options for patients who experience systemic progression after osimertinib treatment [ As a crucial “controller” that is related to the inhibition of tumor cell proliferation, angiogenesis, invasion, metastasis, and apoptosis, EGFR actively participates in malignant disease progression. However, the intrinsic and acquired resistance in primary and recurrent cancer which is mediated by EGFR mutations after target treatment leads to difficult therapeutic. Understanding the complex resistance mechanisms of EGFR-TKIs and develo** potential strategies to combat it could be of potential interest for improving the individual therapeutic strategies for cancer.Conclusion
Availability of data and materials
The material supporting the conclusion of this review has been included within the article.
Abbreviations
- ACK1:
-
Activated Cdc42-associated kinase 1
- ADC:
-
Antibody drug conjugates
- AKR1B1:
-
Aldehyde–ketone reductase family 1 member B1
- A-loop:
-
Activation loop
- ATP:
-
Adenosine triphosphate
- AURK:
-
Aurora kinases
- AUTAC:
-
Autophagy-targeting chimera
- Bcl-2:
-
B cell lymphoma-2
- BRAF:
-
V-RAF murine sarcoma viral oncogene homolog B1
- CCND:
-
Cyclin D
- CCNE1:
-
Cyclin E1
- CDK4/6:
-
Cyclin-dependent kinase 4/6
- cIAP1:
-
Cellular inhibitor of apoptosis protein 1
- CRBN:
-
Cereblon
- DFG:
-
Asp-Phe-Gly
- DZ-SIM:
-
DZ-SIMvastatin
- EGF:
-
Epidermal growth factor
- EGFR:
-
Epidermal growth factor receptor
- EGFR-TKIs:
-
Epidermal growth factor receptor tyrosine kinase inhibitors
- EMT:
-
Epithelial–mesenchymal transformation
- EMT-TFs:
-
EMT-induced transcription factors
- ERC1:
-
Excision repair cross-complementation 1
- FGF:
-
Fibroblast growth factor
- FGFR:
-
Fibroblast growth factor receptor
- Fv:
-
Variable region fragment
- Grp94:
-
Glucose regulatory protein 94
- HB-EGF:
-
EGF-like growth factor
- HMGCR:
-
3-Hydroxy-3-methylglutaryl-CoA reductase
- HNK:
-
Honokiol
- ICIs:
-
Immune checkpoint inhibitors
- IGF1R:
-
Insulin-like growth factor receptor 1
- LUAD:
-
Lung adenocarcinoma
- MDM2:
-
Mouse double minute 2
- NCCN:
-
National Comprehensive Cancer Network
- NSCLC:
-
Non-small cell lung cancer
- NTRK1:
-
Neurotrophic tyrosine receptor kinase 1
- PARP:
-
Poly(ADP-ribose) polymerase
- PGAM1:
-
Phosphoglycerate mutase 1
- PIK3CA:
-
Phosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit
- PROTAC:
-
Proteolysis-targeting chimera
- PTEN:
-
Phosphatase and tensin homolog
- ROS:
-
Reactive oxygen species
- RTK:
-
Receptor tyrosine kinase
- SCLC:
-
Small cell lung cancer
- SLC7A11:
-
Solute carrier family 7 member 11
- SPP1:
-
Secreted phosphoprotein 1
- STAT:
-
Signal sensor and transcription activator
- TACC3:
-
Transforming acid helix protein 3
- TGF:
-
Transforming growth factor
- TMP3:
-
Thrombopoietin mimetic peptide 3
- VHL:
-
Von Hippel–Lindau
- WT:
-
Wild-type
- WT-EGFR:
-
Wild-type EGFR
References
Amelia T, Kartasasmita RE, Ohwada T, Tjahjono DH. Structural insight and development of EGFR tyrosine kinase inhibitors. Molecules. 2022;27:819.
da Cunha SG, Shepherd FA, Tsao MS. EGFR mutations and lung cancer. Annu Rev Pathol. 2011;6:49–69.
Campbell ID, Bork P. Epidermal growth factor-like modules. Curr Opin Struct Biol. 1993;3:385–92.
Roskoski R. The ERBB/HER family of protein-tyrosine kinases and cancer. Pharmacol Res. 2014;79:34–74.
Gazdar AF. Activating and resistance mutations of EGFR in non-small-cell lung cancer: role in clinical response to EGFR tyrosine kinase inhibitors. Oncogene. 2009;28(Suppl 1):S24–31.
Herbst RS, Langer CJ. Epidermal growth factor receptors as a target for cancer treatment: the emerging role of IMC-C225 in the treatment of lung and head and neck cancers. Semin Oncol. 2002;29:27–36.
Normanno N, Bianco C, De Luca A, Salomon DS. The role of EGF-related peptides in tumor growth. Front Biosci. 2001;6:D685–707.
Thomas R, Weihua Z. Rethink of EGFR in cancer with its kinase independent function on board. Front Oncol. 2019;9:800.
Sabbah DA, Hajjo R, Sweidan K. Review on epidermal growth factor receptor (EGFR) structure, signaling pathways, interactions, and recent updates of EGFR inhibitors. Curr Top Med Chem. 2020;20:815–34.
Cross DA, Ashton SE, Ghiorghiu S, Eberlein C, Nebhan CA, Spitzler PJ, et al. AZD9291, an irreversible EGFR TKI, overcomes T790M-mediated resistance to EGFR inhibitors in lung cancer. Cancer Discov. 2014;4:1046–61.
Ricordel C, Friboulet L, Facchinetti F, Soria JC. Molecular mechanisms of acquired resistance to third-generation EGFR-TKIs in EGFR T790M-mutant lung cancer. Ann Oncol. 2018;29:i28–37.
Roskoski R Jr. Classification of small molecule protein kinase inhibitors based upon the structures of their drug-enzyme complexes. Pharmacol Res. 2016;103:26–48.
Sequist LV, Yang JC, Yamamoto N, O’Byrne K, Hirsh V, Mok T, et al. Phase iii study of afatinib or cisplatin plus pemetrexed in patients with metastatic lung adenocarcinoma with EGFR mutations. J Clin Oncol. 2013;31:3327–34.
Wu YL, Zhou C, Hu CP, Feng J, Lu S, Huang Y, et al. Afatinib versus cisplatin plus gemcitabine for first-line treatment of Asian patients with advanced non-small-cell lung cancer harbouring EGFR mutations (LUX-Lung 6): an open-label, randomised phase 3 trial. Lancet Oncol. 2014;15:213–22.
Zhang H. Three generations of epidermal growth factor receptor tyrosine kinase inhibitors developed to revolutionize the therapy of lung cancer. Drug Des Devel Ther. 2016;10:3867–72.
Pao W, Chmielecki J. Rational, biologically based treatment of EGFR-mutant non-small-cell lung cancer. Nat Rev Cancer. 2010;10:760–74.
Jiang T, Zhou C. Clinical activity of the mutant-selective EGFR inhibitor AZD9291 in patients with EGFR inhibitor-resistant non-small cell lung cancer. Transl Lung Cancer Res. 2014;3:370–2.
Yan XE, Zhu SJ, Liang L, Zhao P, Choi HG, Yun CH. Structural basis of mutant-selectivity and drug-resistance related to CO-1686. Oncotarget. 2017;8:53508–17.
Leonetti A, Sharma S, Minari R, Perego P, Giovannetti E, Tiseo M. Resistance mechanisms to osimertinib in EGFR-mutated non-small cell lung cancer. Br J Cancer. 2019;121:725–37.
Hata AN, Niederst MJ, Archibald HL, Gomez-Caraballo M, Siddiqui FM, Mulvey HE, et al. Tumor cells can follow distinct evolutionary paths to become resistant to epidermal growth factor receptor inhibition. Nat Med. 2016;22:262–9.
Guibert N, Barlesi F, Descourt R, Lena H, Besse B, Beau-Faller M, et al. Characteristics and outcomes of patients with lung cancer harboring multiple molecular alterations: results from the IFCT study biomarkers france. J Thorac Oncol. 2017;12:963–73.
Li X, Wang S, Li B, Wang Z, Shang S, Shao Y, et al. Bim deletion polymorphism confers resistance to osimertinib in EGFR T790M lung cancer: a case report and literature review. Target Oncol. 2018;13:517–23.
Eck MJ, Yun CH. Structural and mechanistic underpinnings of the differential drug sensitivity of EGFR mutations in non-small cell lung cancer. Biochim Biophys Acta. 2010;1804:559–66.
Westover D, Zugazagoitia J, Cho BC, Lovly CM, Paz-Ares L. Mechanisms of acquired resistance to first- and second-generation EGFR tyrosine kinase inhibitors. Ann Oncol. 2018;29:i10–9.
Zhou W, Ercan D, Chen L, Yun CH, Li D, Capelletti M, et al. Novel mutant-selective EGFR kinase inhibitors against EGFR T790M. Nature. 2009;462:1070–4.
Zhang Q, Zhang XC, Yang JJ, Yang ZF, Bai Y, Su J, et al. EGFR L792H and G796R: two novel mutations mediating resistance to the third-generation EGFR tyrosine kinase inhibitor osimertinib. J Thorac Oncol. 2018;13:1415–21.
Zheng D, Hu M, Bai Y, Zhu X, Lu X, Wu C, et al. EGFR G796D mutation mediates resistance to osimertinib. Oncotarget. 2017;8:49671–9.
Ou SI, Cui J, Schrock AB, Goldberg ME, Zhu VW, Albacker L, et al. Emergence of novel and dominant acquired EGFR solvent-front mutations at Gly796 (G796S/R) together with C797S/R and L792F/H mutations in one EGFR (L858R/T790M) NSCLC patient who progressed on osimertinib. Lung Cancer. 2017;108:228–31.
Castellano GM, Aisner J, Burley SK, Vallat B, Yu HA, Pine SR, et al. A novel acquired exon 20 EGFR M766Q mutation in lung adenocarcinoma mediates osimertinib resistance but is sensitive to neratinib and poziotinib. J Thorac Oncol. 2019;14:1982–8.
Liu J, ** B, Su H, Qu X, Liu Y. Afatinib helped overcome subsequent resistance to osimertinib in a patient with NSCLC having leptomeningeal metastasis baring acquired EGFR L718Q mutation: a case report. BMC Cancer. 2019;19:702.
Bersanelli M, Minari R, Bordi P, Gnetti L, Bozzetti C, Squadrilli A, et al. L718Q mutation as new mechanism of acquired resistance to AZD9291 in EGFR -mutated NSCLC. J Thorac Oncol. 2016;11:e121–3.
Callegari D, Ranaghan KE, Woods CJ, Minari R, Tiseo M, Mor M, et al. L718Q mutant EGFR escapes covalent inhibition by stabilizing a non-reactive conformation of the lung cancer drug osimertinib. Chem Sci. 2018;9:2740–9.
Yang Z, Yang J, Chen Y, Shao YW, Wang X. Acquired EGFR L718V mutation as the mechanism for osimertinib resistance in a T790M-negative non-small-cell lung cancer patient. Target Oncol. 2019;14:369–74.
Fassunke J, Muller F, Keul M, Michels S, Dammert MA, Schmitt A, et al. Overcoming EGFR (G724S)-mediated osimertinib resistance through unique binding characteristics of second-generation EGFR inhibitors. Nat Commun. 2018;9:4655.
Tu HY, Ke EE, Yang JJ, Sun YL, Yan HH, Zheng MY, et al. A comprehensive review of uncommon EGFR mutations in patients with non-small cell lung cancer. Lung Cancer. 2017;114:96–102.
Xu J, ** B, Chu T, Dong X, Yang H, Zhang Y, et al. EGFR tyrosine kinase inhibitor (TKI) in patients with advanced non-small cell lung cancer (NSCLC) harboring uncommon EGFR mutations: a real-world study in china. Lung Cancer. 2016;96:87–92.
Shen YC, Tseng GC, Tu CY, Chen WC, Liao WC, Chen WC, et al. Comparing the effects of afatinib with gefitinib or erlotinib in patients with advanced-stage lung adenocarcinoma harboring non-classical epidermal growth factor receptor mutations. Lung Cancer. 2017;110:56–62.
Ercan D, Choi HG, Yun CH, Capelletti M, **e T, Eck MJ, et al. EGFR mutations and resistance to irreversible pyrimidine-based EGFR inhibitors. Clin Cancer Res. 2015;21:3913–23.
Piotrowska Z, Isozaki H, Lennerz JK, Gainor JF, Lennes IT, Zhu VW, et al. Landscape of acquired resistance to osimertinib in EGFR -mutant NSCLC and clinical validation of combined EGFR and RET inhibition with osimertinib and BLU-667 for acquired RET fusion. Cancer Discov. 2018;8:1529–39.
Piotrowska Z, Niederst MJ, Karlovich CA, Wakelee HA, Neal JW, Mino-Kenudson M, et al. Heterogeneity underlies the emergence of EGFRT790 wild-type clones following treatment of T790M-positive cancers with a third-generation EGFR inhibitor. Cancer Discov. 2015;5:713–22.
Nukaga S, Yasuda H, Tsuchihara K, Hamamoto J, Masuzawa K, Kawada I, et al. Amplification of EGFR wild-type alleles in non-small cell lung cancer cells confers acquired resistance to mutation-selective EGFR tyrosine kinase inhibitors. Cancer Res. 2017;77:2078–89.
Huang C, Zou Q, Liu H, Qiu B, Li Q, Lin Y, et al. Management of non-small cell lung cancer patients with MET exon 14 skip** mutations. Curr Treat Opt Oncol. 2020;21:33.
Mueller KL, Madden JM, Zoratti GL, Kuperwasser C, List K, Boerner JL. Fibroblast-secreted hepatocyte growth factor mediates epidermal growth factor receptor tyrosine kinase inhibitor resistance in triple-negative breast cancers through paracrine activation of MET. Breast Cancer Res. 2012;14:R104.
Hsu CC, Liao BC, Liao WY, Markovets A, Stetson D, Thress K, et al. Exon 16-skip** HER2 as a novel mechanism of osimertinib resistance in EGFR L858R/T790M-positive non-small cell lung cancer. J Thorac Oncol. 2020;15:50–61.
Ou S-HI, Madison R, Robichaux JP, Ross JS, Miller VA, Ali SM, et al. Characterization of 648 non-small cell lung cancer (NSCLC) cases with 28 unique HER2 exon 20 insertions. J Clin Oncol. 2019;37:9063–63.
Gao G, Li X, Wang Q, Zhang Y, Chen J, Shu Y, et al. Single-arm, phase ii study of pyrotinib in advanced non-small cell lung cancer (NSCLC) patients with HER2 exon 20 mutation. J Clin Oncol. 2019;37:9089–189.
Wu SG, Shih JY. Management of acquired resistance to EGFR TKI-targeted therapy in advanced non-small cell lung cancer. Mol Cancer. 2018;17:38.
Zhu J, Yang Q, Xu W. Iterative upgrading of small molecular tyrosine kinase inhibitors for EGFR mutation in NSCLC: necessity and perspective. Pharmaceutics. 2021;13:1500.
Taniguchi H, Yamada T, Wang R, Tanimura K, Adachi Y, Nishiyama A, et al. AXL confers intrinsic resistance to osimertinib and advances the emergence of tolerant cells. Nat Commun. 2019;10:259.
Yano S, Yamada T, Takeuchi S, Tachibana K, Minami Y, Yatabe Y, et al. Hepatocyte growth factor expression in EGFR mutant lung cancer with intrinsic and acquired resistance to tyrosine kinase inhibitors in a Japanese cohort. J Thorac Oncol. 2011;6:2011–7.
Kim TM, Song A, Kim DW, Kim S, Ahn YO, Keam B, et al. Mechanisms of acquired resistance to AZD9291 a mutation-selective, irreversible EGFR inhibitor. J Thorac Oncol. 2015;10:1736–44.
Papadimitrakopoulou VA, Wu YL, Han JY, Ahn MJ, Ramalingam SS, John T, et al. Analysis of resistance mechanisms to osimertinib in patients with EGFR T790M advanced NSCLC from the AURA3 study. Ann Oncol. 2018;29:741–841.
Tanaka H, Sakagami H, Kaneko N, Konagai S, Yamamoto H, Matsuya T, et al. Mutant-selective irreversible EGFR inhibitor, naquotinib, inhibits tumor growth in NSCLC models with EGFR-activating mutations, T790M mutation, and AXL overexpression. Mol Cancer Ther. 2019;18:1366–73.
Park JH, Choi YJ, Kim SY, Lee JE, Sung KJ, Park S, et al. Activation of the IGF1R pathway potentially mediates acquired resistance to mutant-selective 3rd-generation EGF receptor tyrosine kinase inhibitors in advanced non-small cell lung cancer. Oncotarget. 2016;7:22005–15.
Carmena M, Earnshaw WC. The cellular geography of aurora kinases. Nat Rev Mol Cell Biol. 2003;4:842–54.
Cheetham GMT, Knegtel RMA, Coll JT, Renwick SB, Swenson L, Weber P, et al. Crystal structure of aurora-2, an oncogenic serine/threonine kinase*. J Biol Chem. 2002;277:42419–22.
Pradhan T, Gupta O, Singh G, Monga V. Aurora kinase inhibitors as potential anticancer agents: recent advances. Eur J Med Chem. 2021;221:113495.
Falchook GS, Bastida CC, Kurzrock R. Aurora kinase inhibitors in oncology clinical trials: current state of the progress. Semin Oncol. 2015;42:832–48.
Hu L, Fan M, Shi S, Song X, Wang F, He H, et al. Dual target inhibitors based on EGFR: promising anticancer agents for the treatment of cancers (2017-). Eur J Med Chem. 2022;227:113963.
Tanaka K, Yu HA, Yang S, Han S, Selcuklu SD, Kim K, et al. Targeting aurora b kinase prevents and overcomes resistance to EGFR inhibitors in lung cancer by enhancing BIM- and PUMA-mediated apoptosis. Cancer Cell. 2021;39(1245–61):e6.
Mao C, Qiu LX, Liao RY, Du FB, Ding H, Yang WC, et al. KRAS mutations and resistance to EGFR-TKIs treatment in patients with non-small cell lung cancer: a meta-analysis of 22 studies. Lung Cancer. 2010;69:272–8.
Sunaga N, Shames DS, Girard L, Peyton M, Larsen JE, Imai H, et al. Knockdown of oncogenic KRAS in non-small cell lung cancers suppresses tumor growth and sensitizes tumor cells to targeted therapy. Mol Cancer Ther. 2011;10:336–46.
Leonetti A, Facchinetti F, Rossi G, Minari R, Conti A, Friboulet L, et al. Braf in non-small cell lung cancer (NSCLC): pickaxing another brick in the wall. Cancer Treat Rev. 2018;66:82–94.
Ho CC, Liao WY, Lin CA, Shih JY, Yu CJ, Yang JC. Acquired BRAF V600E mutation as resistant mechanism after treatment with osimertinib. J Thorac Oncol. 2017;12:567–72.
Ramalingam SS, Cheng Y, Zhou C, Ohe Y, Imamura F, Cho BC, et al. Mechanisms of acquired resistance to first-line osimertinib: preliminary data from the phase iii flaura study. Ann Oncol. 2018;29:viii740.
Zhao M, Gao FH, Wang JY, Liu F, Yuan HH, Zhang WY, et al. JAK2/STAT3 signaling pathway activation mediates tumor angiogenesis by upregulation of VEGF and bFGF in non-small-cell lung cancer. Lung Cancer. 2011;73:366–74.
Chaib I, Karachaliou N, Pilotto S, Codony Servat J, Cai X, Li X, et al. Co-activation of STAT3 and YES-associated protein 1 (YAP1) pathway in EGFR-mutant NSCLC. J Natl Cancer Inst. 2017;109.
Soria J-C, Lee H-Y, Lee JI, Wang L, Issa J-P, Kemp BL, et al. Lack of PTEN expression in non-small cell lung cancer could be related to promoter methylation. Clin Cancer Res. 2002;8:1178–84.
Zhang T, Qu R, Chan S, Lai M, Tong L, Feng F, et al. Discovery of a novel third-generation EGFR inhibitor and identification of a potential combination strategy to overcome resistance. Mol Cancer. 2020;19:90.
Zhu L, Chen Z, Zang H, Fan S, Gu J, Zhang G, et al. Targeting c-Myc to overcome acquired resistance of EGFR mutant NSCLC cells to the third-generation EGFR tyrosine kinase inhibitor, osimertinib. Cancer Res. 2021;81:4822–34.
Weng CH, Chen LY, Lin YC, Shih JY, Lin YC, Tseng RY, et al. Epithelial–mesenchymal transition (emtEMT) beyond EGFR mutations per se is a common mechanism for acquired resistance to EGFR TKI. Oncogene. 2019;38:455–68.
Brabletz T, Kalluri R, Nieto MA, Weinberg RA. EMT in cancer. Nat Rev Cancer. 2018;18:128–34.
Kong W, Yang H, He L, Zhao JJ, Coppola D, Dalton WS, et al. MicroRNA-155 is regulated by the transforming growth factor beta/Smad pathway and contributes to epithelial cell plasticity by targeting RhoA. Mol Cell Biol. 2008;28:6773–84.
Liu CH, Huang Q, ** ZY, Zhu CL, Liu Z, Wang C. miR-21 and KLF4 jointly augment epithelialmesenchymal transition via the Akt/ERK1/2 pathway. Int J Oncol. 2017;50:1109–15.
Han Z, Zhou X, Li S, Qin Y, Chen Y, Liu H. Inhibition of miR-23a increases the sensitivity of lung cancer stem cells to erlotinib through PTEN/PI3K/Akt pathway. Oncol Rep. 2017;38:3064–70.
Shen H, Zhu F, Liu J, Xu T, Pei D, Wang R, et al. Alteration in Mir-21/PTEN expression modulates gefitinib resistance in non-small cell lung cancer. PLoS ONE. 2014;9:e103305.
Fardi M, Solali S, Farshdousti HM. Epigenetic mechanisms as a new approach in cancer treatment: an updated review. Genes Dis. 2018;5:304–11.
Del Re M, Arrigoni E, Restante G, Passaro A, Rofi E, Crucitta S, et al. Concise review: resistance to tyrosine kinase inhibitors in non-small cell lung cancer: the role of cancer stem cells. Stem Cells. 2018;36:633–40.
Papadimitrakopoulou VA, Wu YL, Han JY, Ahn MJ, Ramalingam SS, John T, et al. Analysis of resistance mechanisms to osimertinib in patients with EGFR T790M advanced NSCLC from the AURA3 study. Ann Oncol. 2018;29:viii741.
Dorantes-Heredia R, Ruiz-Morales JM, Cano-Garcia F. Histopathological transformation to small-cell lung carcinoma in non-small cell lung carcinoma tumors. Transl Lung Cancer Res. 2016;5:401–12.
Sequist LV, Waltman BA, Dias-Santagata D, Digumarthy S, Turke AB, Fidias P, et al. Genotypic and histological evolution of lung cancers acquiring resistance to EGFR inhibitors. Sci Transl Med. 2011;3:75ra26.
Oser MG, Niederst MJ, Sequist LV, Engelman JA. Transformation from non-small-cell lung cancer to small-cell lung cancer: Molecular drivers and cells of origin. Lancet Oncol. 2015;16:e165-172.
Norkowski E, Ghigna MR, Lacroix L, Le Chevalier T, Fadel E, Dartevelle P, et al. Small-cell carcinoma in the setting of pulmonary adenocarcinoma: new insights in the era of molecular pathology. J Thorac Oncol. 2013;8:1265–71.
Yin X, Li Y, Wang H, Jia T, Wang E, Luo Y, et al. Small cell lung cancer transformation: from pathogenesis to treatment. Semin Cancer Biol. 2022. https://doi.org/10.1016/j.semcancer.2022.03.006.
Schoenfeld AJ, Chan JM, Kubota D, Sato H, Rizvi H, Daneshbod Y, et al. Tumor analyses reveal squamous transformation and off-target alterations as early resistance mechanisms to first-line osimertinib in EGFR-mutant lung cancer. Clin Cancer Res. 2020;26:2654–63.
Garassino MC, Cho BC, Kim JH, Mazieres J, Vansteenkiste J, Lena H, et al. Durvalumab as third-line or later treatment for advanced non-small-cell lung cancer (ATLANTIC): an open-label, single-arm, phase 2 study. Lancet Oncol. 2018;19:521–36.
Gainor JF, Shaw AT, Sequist LV, Fu X, Azzoli CG, Piotrowska Z, et al. EGFR mutations and ALK rearrangements are associated with low response rates to PD-1 pathway blockade in non-small cell lung cancer: a retrospective analysis. Clin Cancer Res. 2016;22:4585–93.
Haratani K, Hayashi H, Tanaka T, Kaneda H, Togashi Y, Sakai K, et al. Tumor immune microenvironment and nivolumab efficacy in EGFR mutation-positive non-small-cell lung cancer based on T790M status after disease progression during EGFR-TKI treatment. Ann Oncol. 2017;28:1532–9.
Zheng Y, Hao S, **ang C, Han Y, Shang Y, Zhen Q, et al. The correlation between SPP1 and immune escape of EGFR mutant lung adenocarcinoma was explored by bioinformatics analysis. Front Oncol. 2021;11:592854.
Peng S, Wang R, Zhang X, Ma Y, Zhong L, Li K, et al. EGFR-TKI resistance promotes immune escape in lung cancer via increased PD-L1 expression. Mol Cancer. 2019;18:165.
Yu HA, Tian SK, Drilon AE, Borsu L, Riely GJ, Arcila ME, et al. Acquired resistance of EGFR-mutant lung cancer to a T790M-specific EGFR inhibitor: emergence of a third mutation (C797S) in the EGFR tyrosine kinase domain. JAMA Oncol. 2015;1:982–4.
Wang Z, Yang JJ, Huang J, Ye JY, Zhang XC, Tu HY, et al. Lung adenocarcinoma harboring EGFR T790M and in trans C797S responds to combination therapy of first- and third-generation EGFR TKIs and shifts allelic configuration at resistance. J Thorac Oncol. 2017;12:1723–7.
Niederst MJ, Hu H, Mulvey HE, Lockerman EL, Garcia AR, Piotrowska Z, et al. The allelic context of the C797S mutation acquired upon treatment with third-generation EGFR inhibitors impacts sensitivity to subsequent treatment strategies. Clin Cancer Res. 2015;21:3924–33.
Tsai CJ, Nussinov R. Emerging allosteric mechanism of EGFR activation in physiological and pathological contexts. Biophys J. 2019;117:5–13.
Engel J, Richters A, Getlik M, Tomassi S, Keul M, Termathe M, et al. Targeting drug resistance in EGFR with covalent inhibitors: a structure-based design approach. J Med Chem. 2015;58:6844–63.
Jia Y, Yun CH, Park E, Ercan D, Manuia M, Juarez J, et al. Overcoming EGFR(T790M) and EGFR(C797S) resistance with mutant-selective allosteric inhibitors. Nature. 2016;534:129–32.
Lee S, Kim J, Duggirala KB, Go A, Shin I, Cho BC, et al. Allosteric inhibitor TREA-0236 containing non-hydrolysable quinazoline-4-one for EGFR T790M/C797S mutants inhibition. Bull Korean Chem Soc. 2018;39:895–8.
To C, Jang J, Chen T, Park E, Mushajiang M, De Clercq DJH, et al. Single and dual targeting of mutant EGFR with an allosteric inhibitor. Cancer Discov. 2019;9:926–43.
Maity S, Pai KSR, Nayak Y. Advances in targeting EGFR allosteric site as anti-NSCLC therapy to overcome the drug resistance. Pharmacol Rep. 2020;72:799–813.
De Clercq DJH, Heppner DE, To C, Jang J, Park E, Yun C-H, et al. Discovery and optimization of dibenzodiazepinones as allosteric mutant-selective EGFR inhibitors. ACS Med Chem Lett. 2019;10:1549–53.
Duplessis M, Goergler A, Jaeschke G, Kocer B, Kuhn B, Lazarski K, et al. COMPOUNDS. Publication number: 20210079005, March 18, 2021.
Lu X, Zhang T, Zhu SJ, Xun Q, Tong L, Hu X, et al. Discovery of JND3229 as a new EGFR(C797S) mutant inhibitor with in vivo monodrug efficacy. ACS Med Chem Lett. 2018;9:1123–7.
Engel J, Becker C, Lategahn J, Keul M, Ketzer J, Muhlenberg T, et al. Insight into the inhibition of drug-resistant mutants of the receptor tyrosine kinase EGFR. Angew Chem Int Ed Engl. 2016;55:10909–12.
Gunther M, Lategahn J, Juchum M, Doring E, Keul M, Engel J, et al. Trisubstituted pyridinylimidazoles as potent inhibitors of the clinically resistant L858R/T790M/C797S EGFR mutant: targeting of both hydrophobic regions and the phosphate binding site. J Med Chem. 2017;60:5613–37.
Park H, Jung HY, Mah S, Hong S. Discovery of EGF receptor inhibitors that are selective for the D746–750/T790M/C797S mutant through structure-based de novo design. Angew Chem Int Ed Engl. 2017;56:7634–8.
Zhang M, Wang Y, Wang J, Liu Z, Shi J, Li M, et al. Design, synthesis and biological evaluation of the quinazoline derivatives as L858R/T790M/C797S triple mutant epidermal growth factor receptor tyrosine kinase inhibitors. Chem Pharm Bull (Tokyo). 2020;68:971–80.
Shen J, Zhang T, Zhu SJ, Sun M, Tong L, Lai M, et al. Structure-based design of 5-methylpyrimidopyridone derivatives as new wild-type sparing inhibitors of the epidermal growth factor receptor triple mutant (EGFR(L858R/T790M/C797S). J Med Chem. 2019;62:7302–8.
Zhang H, Wang J, Shen Y, Wang HY, Duan WM, Zhao HY, et al. Discovery of 2,4,6-trisubstitued pyrido[3,4-d]pyrimidine derivatives as new EGFR-TKIs. Eur J Med Chem. 2018;148:221–37.
Hei YY, Shen Y, Wang J, Zhang H, Zhao HY, **n M, et al. Synthesis and evaluation of 2,9-disubstituted 8-phenylthio/phenylsulfinyl-9H-purine as new EGFR inhibitors. Bioorg Med Chem. 2018;26:2173–85.
Lei H, Fan S, Zhang H, Liu YJ, Hei YY, Zhang JJ, et al. Discovery of novel 9-heterocyclyl substituted 9H-purines as L858R/T790M/C797S mutant EGFR tyrosine kinase inhibitors. Eur J Med Chem. 2020;186:111888.
Lategahn J, Keul M, Klovekorn P, Tumbrink HL, Niggenaber J, Muller MP, et al. Inhibition of osimertinib-resistant epidermal growth factor receptor EGFR-T790M/C797S. Chem Sci. 2019;10:10789–801.
Hu X, Xun Q, Zhang T, Zhu S-J, Li Q, Tong L, et al. 2-Oxo-3,4-dihydropyrimido[4,5-d] pyrimidines as new reversible inhibitors of EGFR C797S (Cys797 to Ser797) mutant. Chin Chem Lett. 2020;31:1281–7.
Su Z, Yang T, Wang J, Lai M, Tong L, Wumaier G, et al. Design, synthesis and biological evaluation of potent EGFR kinase inhibitors against 19D/T790M/C797S mutation. Bioorg Med Chem Lett. 2020;30:127327.
Lee Kwangho, SHIN Inji, CHOI Gildon, CHAE Chong Hak, Choe Hyeon Jeong, JUNG Myoung Eun, et al. N2,N4-diphenylpyrimidine-2,4-diamine derivative, method for preparing same, and pharmaceutical composition containing same as active ingredient for prevention or treatment of cancer. WO2018230934, 2018.
Wu L, Liu X, Ding CZ, Chen S, Hu L, Zhao L, et al. Spiro-aryl-phosphorus-oxygen compound as fourth generation of EGFR kinase inhibitor. WO 2018108064 A1, 2016.
Iwao M, Fukuda T, Ishibashi F, Uehara Y, Nishiya N, Oku Y, et al. Fourth-generation EGFR tyrosine kinase inhibitor. CN 110461850 A, 2019.
Boese D, Dahmann G, Engelhardt H, Petronczki M, Scharn D. New benzimidazole compounds and derivatives as EGFR inhibitors. WO 2019162323 A1, 2019.
Ding K, Ding J, Shen J, Geng M, Lu X, **e H, et al. Pyrimidopyridone or pyridopyridone compound and use thereof. WO 2019015593 A1, 2019.
Ferlenghi F, Scalvini L, Vacondio F, Castelli R, Bozza N, Marseglia G, et al. A sulfonyl fluoride derivative inhibits EGFR(L858R/T790M/C797S) by covalent modification of the catalytic lysine. Eur J Med Chem. 2021;225:113786.
Morabito A, Piccirillo MC, Falasconi F, De Feo G, Del Giudice A, Bryce J, et al. Vandetanib (ZD6474), a dual inhibitor of vascular endothelial growth factor receptor (vEGFR) and epidermal growth factor receptor (EGFR) tyrosine kinases: Current status and future directions. Oncologist. 2009;14:378–90.
Li Q, Zhang T, Li S, Tong L, Li J, Su Z, et al. Discovery of potent and noncovalent reversible EGFR kinase inhibitors of EGFR(L858R/T790M/C797S). ACS Med Chem Lett. 2019;10:869–73.
Wittlinger F, Heppner DE, To C, Gunther M, Shin BH, Rana JK, et al. Design of a “two-in-one” mutant-selective epidermal growth factor receptor inhibitor that spans the orthosteric and allosteric sites. J Med Chem. 2022;65:1370–83.
Yu HA, Arcila ME, Rekhtman N, Sima CS, Zakowski MF, Pao W, et al. Analysis of tumor specimens at the time of acquired resistance to EGFR-TKI therapy in 155 patients with EGFR-mutant lung cancers. Clin Cancer Res. 2013;19:2240–7.
Noble ME, Endicott JA, Johnson LN. Protein kinase inhibitors: Insights into drug design from structure. Science. 2004;303:1800–5.
Bondeson DP, Mares A, Smith IE, Ko E, Campos S, Miah AH, et al. Catalytic in vivo protein knockdown by small-molecule protacs. Nat Chem Biol. 2015;11:611–7.
Winter GE, Buckley DL, Paulk J, Roberts JM, Souza A, Dhe-Paganon S, et al. Drug development. Phthalimide conjugation as a strategy for in vivo target protein degradation. Science. 2015;348:1376–81.
Scheepstra M, Hekking KFW, van Hijfte L, Folmer RHA. Bivalent ligands for protein degradation in drug discovery. Comput Struct Biotechnol J. 2019;17:160–76.
An S, Fu L. Small-molecule protacs: an emerging and promising approach for the development of targeted therapy drugs. EBioMedicine. 2018;36:553–62.
Chamberlain PP, Hamann LG. Development of targeted protein degradation therapeutics. Nat Chem Biol. 2019;15:937–44.
Churcher I. Protac-induced protein degradation in drug discovery: breaking the rules or just making new ones? J Med Chem. 2018;61:444–52.
Cromm PM, Crews CM. Targeted protein degradation: from chemical biology to drug discovery. Cell Chem Biol. 2017;24:1181–90.
Toure M, Crews CM. Small-molecule protacs: new approaches to protein degradation. Angew Chem Int Ed Engl. 2016;55:1966–73.
Jang J, To C, De Clercq DJH, Park E, Ponthier CM, Shin BH, et al. Mutant-selective allosteric EGFR degraders are effective against a broad range of drug-resistant mutations. Angew Chem Int Ed Engl. 2020;59:14481–9.
Zhao HY, Yang XY, Lei H, ** XX, Lu SM, Zhang JJ, et al. Discovery of potent small molecule protacs targeting mutant EGFR. Eur J Med Chem. 2020;208:112781.
Qu X, Liu H, Song X, Sun N, Zhong H, Qiu X, et al. Effective degradation of EGFRL858R+T790M mutant proteins by CRBN-based PROTAC s through both proteosome and autophagy/lysosome degradation systems. Eur J Med Chem. 2021;218:113328.
Zheng M, Huo J, Gu X, Wang Y, Wu C, Zhang Q, et al. Rational design and synthesis of novel dual PROTACS for simultaneous degradation of EGFR and PARP. J Med Chem. 2021;64:7839–52.
Kim JH, Nam B, Choi YJ, Kim SY, Lee JE, Sung KJ, et al. Enhanced glycolysis supports cell survival in EGFR-mutant lung adenocarcinoma by inhibiting autophagy-mediated EGFR degradation. Cancer Res. 2018;78:4482–96.
Takahashi D, Arimoto H. Targeting selective autophagy by AUTAC degraders. Autophagy. 2020;16:765–6.
Takahashi D, Moriyama J, Nakamura T, Miki E, Takahashi E, Sato A, et al. Autacs: Cargo-specific degraders using selective autophagy. Mol Cell. 2019;76(797–810):e10.
Ramalingam SS, Vansteenkiste J, Planchard D, Cho BC, Gray JE, Ohe Y, et al. Overall survival with osimertinib in untreated, EGFR-mutated advanced NSCLC. N Engl J Med. 2020;382:41–50.
Chau CH, Steeg PS, Figg WD. Antibody-drug conjugates for cancer. Lancet. 2019;394:793–804.
He K, Xu J, Liang J, Jiang J, Tang M, Ye X, et al. Discovery of a novel EGFR-targeting antibody-drug conjugate, SHR-A1307, for the treatment of solid tumors resistant or refractory to anti-EGFR therapies. Mol Cancer Ther. 2019;18:1104–14.
Xu R-h, Qiu M-Z, Zhang Y, Wei X-L, Hu C. First-in-human dose-escalation study of anti-EGFR adc MRG003 in patients with relapsed/refractory solid tumors. J Clin Oncol. 2020;38:3550–50.
Li Z, Wang M, Yao X, Luo W, Qu Y, Yu D, et al. Development of a novel EGFR-targeting antibody-drug conjugate for pancreatic cancer therapy. Target Oncol. 2019;14:93–105.
Shi P, Oh YT, Zhang G, Yao W, Yue P, Li Y, et al. Met gene amplification and protein hyperactivation is a mechanism of resistance to both first and third generation EGFR inhibitors in lung cancer treatment. Cancer Lett. 2016;380:494–504.
Giroux-Leprieur E, Dumenil C, Chinet T. Combination of crizotinib and osimertinib or erlotinib might overcome MET-mediated resistance to EGFR tyrosine kinase inhibitor in EGFR-mutated adenocarcinoma. J Thorac Oncol. 2018;13:e232–4.
Kang J, Chen HJ, Wang Z, Liu J, Li B, Zhang T, et al. Osimertinib and cabozantinib combinatorial therapy in an EGFR-mutant lung adenocarcinoma patient with multiple MET secondary-site mutations after resistance to crizotinib. J Thorac Oncol. 2018;13:e49–53.
Fu**o T, Suda K, Mitsudomi T. Emerging MET tyrosine kinase inhibitors for the treatment of non-small cell lung cancer. Expert Opin Emerg Drugs. 2020;25:229–49.
Quintanal-Villalonga A, Molina-Pinelo S, Cirauqui C, Ojeda-Marquez L, Marrugal A, Suarez R, et al. FGFR1 cooperates with EGFR in lung cancer oncogenesis, and their combined inhibition shows improved efficacy. J Thorac Oncol. 2019;14:641–55.
Shaw AT, Felip E, Bauer TM, Besse B, Navarro A, Postel-Vinay S, et al. Lorlatinib in non-small-cell lung cancer with ALK or ROS1 rearrangement: an international, multicentre, open-label, single-arm first-in-man phase 1 trial. Lancet Oncol. 2017;18:1590–9.
Uchibori K, Inase N, Araki M, Kamada M, Sato S, Okuno Y, et al. Brigatinib combined with anti-EGFR antibody overcomes osimertinib resistance in EGFR-mutated non-small-cell lung cancer. Nat Commun. 2017;8:14768.
Liu S, Li S, Hai J, Wang X, Chen T, Quinn MM, et al. Targeting HER2 aberrations in non-small cell lung cancer with osimertinib. Clin Cancer Res. 2018;24:2594–604.
La Monica S, Cretella D, Bonelli M, Fumarola C, Cavazzoni A, Digiacomo G, et al. Trastuzumab emtansine delays and overcomes resistance to the third-generation EGFR-TKI osimertinib in NSCLC EGFR mutated cell lines. J Exp Clin Cancer Res. 2017;36:174.
Jani JP, Arcari J, Bernardo V, Bhattacharya SK, Briere D, Cohen BD, et al. PF-03814735, an orally bioavailable small molecule aurora kinase inhibitor for cancer therapy. Mol Cancer Ther. 2010;9:883–94.
Kim C, Giaccone G. MEK inhibitors under development for treatment of non-small-cell lung cancer. Expert Opin Investig Drugs. 2018;27:17–30.
Ortiz-Cuaran S, Scheffler M, Plenker D, Dahmen L, Scheel AH, Fernandez-Cuesta L, et al. Heterogeneous mechanisms of primary and acquired resistance to third-generation EGFR inhibitors. Clin Cancer Res. 2016;22:4837–47.
Della Corte CM, Ciaramella V, Cardone C, La Monica S, Alfieri R, Petronini PG, et al. Antitumor efficacy of dual blockade of EGFR signaling by osimertinib in combination with selumetinib or cetuximab in activated EGFR human NCLC tumor models. J Thorac Oncol. 2018;13:810–20.
Jacobsen K, Bertran-Alamillo J, Molina MA, Teixido C, Karachaliou N, Pedersen MH, et al. Convergent Akt activation drives acquired EGFR inhibitor resistance in lung cancer. Nat Commun. 2017;8:410.
Namba K, Shien K, Takahashi Y, Torigoe H, Sato H, Yoshioka T, et al. Activation of AXL as a preclinical acquired resistance mechanism against osimertinib treatment in EGFR-mutant non-small cell lung cancer cells. Mol Cancer Res. 2019;17:499–507.
Jimbo T, Hatanaka M, Komatsu T, Taira T, Kumazawa K, Maeda N, et al. DS-1205b, a novel selective inhibitor of AXL kinase, blocks resistance to EGFR-tyrosine kinase inhibitors in a non-small cell lung cancer xenograft model. Oncotarget. 2019;10:5152–67.
Kim D, Bach DH, Fan YH, Luu TT, Hong JY, Park HJ, et al. AXL degradation in combination with EGFR-TKI can delay and overcome acquired resistance in human non-small cell lung cancer cells. Cell Death Dis. 2019;10:361.
Liu YN, Tsai MF, Wu SG, Chang TH, Tsai TH, Gow CH, et al. Acquired resistance to EGFR tyrosine kinase inhibitors is mediated by the reactivation of STC2/JUN/AXL signaling in lung cancer. Int J Cancer. 2019;145:1609–24.
Gu J, Qian L, Zhang G, Mahajan NP, Owonikoko TK, Ramalingam SS, et al. Inhibition of ACK1 delays and overcomes acquired resistance of EGFR mutant NSCLC cells to the third generation EGFR inhibitor, osimertinib. Lung Cancer. 2020;150:26–35.
Lawrence HR, Mahajan K, Luo Y, Zhang D, Tindall N, Huseyin M, et al. Development of novel ACK1/TNK2 inhibitors using a fragment-based approach. J Med Chem. 2015;58:2746–63.
Sequist LV, Lynch TJ. EGFR tyrosine kinase inhibitors in lung cancer: an evolving story. Annu Rev Med. 2008;59:429–42.
Kummar S, Chen HX, Wright J, Holbeck S, Millin MD, Tomaszewski J, et al. Utilizing targeted cancer therapeutic agents in combination: novel approaches and urgent requirements. Nat Rev Drug Discov. 2010;9:843–56.
Anighoro A, Bajorath J, Rastelli G. Polypharmacology: challenges and opportunities in drug discovery. J Med Chem. 2014;57:7874–87.
Chen G, Bao Y, Weng Q, Zhao Y, Lu X, Fu L, et al. Compound 15c, a novel dual inhibitor of EGFR(L858R/T790M) and FGFR1, efficiently overcomes epidermal growth factor receptor-tyrosine kinase inhibitor resistance of non-small-cell lung cancers. Front Pharmacol. 2019;10:1533.
Cui Z, Chen S, Wang Y, Gao C, Chen Y, Tan C, et al. Design, synthesis and evaluation of azaacridine derivatives as dual-target EGFR and Src kinase inhibitors for antitumor treatment. Eur J Med Chem. 2017;136:372–81.
Mansour TS, Pallepati RR, Basetti V. Potent dual EGFR/HER4 tyrosine kinase inhibitors containing novel (1,2-dithiolan-4-yl)acetamides. Bioorg Med Chem Lett. 2020;30:127288.
El-Sayed NA, Nour MS, Salem MA, Arafa RK. New oxadiazoles with selective- COX-2 and EGFR dual inhibitory activity: design, synthesis, cytotoxicity evaluation and in silico studies. Eur J Med Chem. 2019;183:111693.
Abdelatef SA, El-Saadi MT, Amin NH, Abdelazeem AH, Omar HA, Abdellatif KRA. Design, synthesis and anticancer evaluation of novel spirobenzo[h]chromene and spirochromane derivatives with dual EGFR and B-RAF inhibitory activities. Eur J Med Chem. 2018;150:567–78.
Jang J, Son JB, To C, Bahcall M, Kim SY, Kang SY, et al. Discovery of a potent dual ALK and EGFR T790M inhibitor. Eur J Med Chem. 2017;136:497–510.
Chen Y, Wu J, Wang A, Qi Z, Jiang T, Chen C, et al. Discovery of n-(5-((5-chloro-4-((2-(isopropylsulfonyl)phenyl)amino)pyrimidin-2-yl)amino)-4-met hoxy-2-(4-methyl-1,4-diazepan-1-yl)phenyl)acrylamide (chmfl-alk/EGFR-050) as a potent ALK/EGFR dual kinase inhibitor capable of overcoming a variety of ALK/EGFR associated drug resistant mutants in NSCLC. Eur J Med Chem. 2017;139:674–97.
**g T, Miao X, Jiang F, Guo M, **ng L, Zhang J, et al. Discovery and optimization of tetrahydropyrido[4,3-d]pyrimidine derivatives as novel ATX and EGFR dual inhibitors. Bioorg Med Chem. 2018;26:1784–96.
Kurup S, McAllister B, Liskova P, Mistry T, Fanizza A, Stanford D, et al. Design, synthesis and biological activity of n(4)-phenylsubstituted-7h-pyrrolo[2,3-d]pyrimidin-4-amines as dual inhibitors of aurora kinase a and epidermal growth factor receptor kinase. J Enzyme Inhib Med Chem. 2018;33:74–84.
Gadekar PK, Urunkar G, Roychowdhury A, Sharma R, Bose J, Khanna S, et al. Design, synthesis and biological evaluation of 2,3-dihydroimidazo[2,1-b]thiazoles as dual EGFR and IGF1R inhibitors. Bioorg Chem. 2021;115:105151.
Romagnoli R, Prencipe F, Oliva P, Baraldi S, Baraldi PG, Schiaffino Ortega S, et al. Design, synthesis, and biological evaluation of 6-substituted thieno[3,2-d]pyrimidine analogues as dual epidermal growth factor receptor kinase and microtubule inhibitors. J Med Chem. 2019;62:1274–90.
Alswah M, Bayoumi AH, Elgamal K, Elmorsy A, Ihmaid S, Ahmed HEA. Design, synthesis and cytotoxic evaluation of novel chalcone derivatives bearing triazolo[4,3-a]-quinoxaline moieties as potent anticancer agents with dual EGFR kinase and tubulin polymerization inhibitory effects. Molecules. 2017;23:48.
Khan I, Garikapati KR, Setti A, Shaik AB, Kanth Makani VK, Shareef MA, et al. Design, synthesis, in silico pharmacokinetics prediction and biological evaluation of 1,4-dihydroindeno[1,2-c]pyrazole chalcone as EGFR/AKT pathway inhibitors. Eur J Med Chem. 2019;163:636–48.
Dong H, Yin H, Zhao C, Cao J, Xu W, Zhang Y. Design, synthesis and biological evaluation of novel osimertinib-based HDAC and EGFR dual inhibitors. Molecules. 2019;24:2407.
Fischer T, Najjar A, Totzke F, Schachtele C, Sippl W, Ritter C, et al. Discovery of novel dual inhibitors of receptor tyrosine kinases EGFR and PDGFR-β related to anticancer drug resistance. J Enzyme Inhib Med Chem. 2018;33:1–8.
Hamed MM, Darwish SS, Herrmann J, Abadi AH, Engel M. First bispecific inhibitors of the epidermal growth factor receptor kinase and the NF-κB activity as novel anticancer agents. J Med Chem. 2017;60:2853–68.
Dokla EME, Fang CS, Abouzid KAM, Chen CS. 1,2,4-oxadiazole derivatives targeting EGFR and c-Met degradation in TKI resistant NSCLC. Eur J Med Chem. 2019;182:111607.
Singh PK, Silakari O. Molecular dynamics guided development of indole based dual inhibitors of EGFR (T790M) and c-Met. Bioorg Chem. 2018;79:163–70.
Fischer T, Kruger T, Najjar A, Totzke F, Schachtele C, Sippl W, et al. Discovery of novel substituted benzo-anellated 4-benzylamino pyrrolopyrimidines as dual EGFR and vEGFR2 inhibitors. Bioorg Med Chem Lett. 2017;27:2708–12.
Zhang HQ, Gong FH, Ye JQ, Zhang C, Yue XH, Li CG, et al. Design and discovery of 4-anilinoquinazoline-urea derivatives as dual TK inhibitors of EGFR and vEGFR-2. Eur J Med Chem. 2017;125:245–54.
Wei H, Duan Y, Gou W, Cui J, Ning H, Li D, et al. Design, synthesis and biological evaluation of novel 4-anilinoquinazoline derivatives as hypoxia-selective EGFR and vEGFR-2 dual inhibitors. Eur J Med Chem. 2019;181:111552.
Sun S, Zhang J, Wang N, Kong X, Fu F, Wang H, et al. Design and discovery of quinazoline- and thiourea-containing sorafenib analogs as EGFR and vEGFR-2 dual TK inhibitors. Molecules. 2017;23:24.
Das D, **e L, Wang J, Xu X, Zhang Z, Shi J, et al. Discovery of new quinazoline derivatives as irreversible dual EGFR/HER2 inhibitors and their anticancer activities: part 1. Bioorg Med Chem Lett. 2019;29:591–6.
Maher M, Kassab AE, Zaher AF, Mahmoud Z. Novel pyrazolo[3,4-d]pyrimidines: design, synthesis, anticancer activity, dual EGFR/ErbB2 receptor tyrosine kinases inhibitory activity, effects on cell cycle profile and caspase-3-mediated apoptosis. J Enzyme Inhib Med Chem. 2019;34:532–46.
Zou M, Li J, ** B, Wang M, Chen H, Zhang Z, et al. Design, synthesis and anticancer evaluation of new 4-anilinoquinoline-3-carbonitrile derivatives as dual EGFR/HER2 inhibitors and apoptosis inducers. Bioorg Chem. 2021;114:105200.
Alsaid MS, Al-Mishari AA, Soliman AM, Ragab FA, Ghorab MM. Discovery of benzo[g]quinazolin benzenesulfonamide derivatives as dual EGFR/HER2 inhibitors. Eur J Med Chem. 2017;141:84–91.
Ghorab MM, Alsaid MS, Soliman AM. Dual EGFR/HER2 inhibitors and apoptosis inducers: new benzo[g]quinazoline derivatives bearing benzenesulfonamide as anticancer and radiosensitizers. Bioorg Chem. 2018;80:611–20.
Soliman AM, Alqahtani AS, Ghorab M. Novel sulphonamide benzoquinazolinones as dual EGFR/HER2 inhibitors, apoptosis inducers and radiosensitizers. J Enzyme Inhib Med Chem. 2019;34:1030–40.
Liu X, Du Q, Tian C, Tang M, Jiang Y, Wang Y, et al. Discovery of cape derivatives as dual EGFR and CSK inhibitors with anticancer activity in a murine model of hepatocellular carcinoma. Bioorg Chem. 2021;107:104536.
Zhang B, Liu Z, **a S, Liu Q, Gou S. Design, synthesis and biological evaluation of sulfamoylphenyl-quinazoline derivatives as potential EGFR/CAIX dual inhibitors. Eur J Med Chem. 2021;216:113300.
Zang H, Qian G, Arbiser J, Owonikoko TK, Ramalingam SS, Fan S, et al. Overcoming acquired resistance of EGFR-mutant NSCLC cells to the third generation EGFR inhibitor, osimertinib, with the natural product honokiol. Mol Oncol. 2020;14:882–95.
Cao F, Gong YB, Kang XH, Lu ZH, Wang Y, Zhao KL, et al. Degradation of MCL-1 by bufalin reverses acquired resistance to osimertinib in EGFR-mutant lung cancer. Toxicol Appl Pharmacol. 2019;379:114662.
Sun P, Qu Y, Wang Y, Wang J, Wang X, Sheng J. Wighteone exhibits an antitumor effect against EGFR L858R/T790M mutation non-small cell lung cancer. J Cancer. 2021;12:3900–8.
Niu M, Xu J, Liu Y, Li Y, He T, Ding L, et al. FBXL 2 counteracts Grp94 to destabilize EGFR and inhibit EGFR-driven NSCLC growth. Nat Commun. 2021;12:5919.
Zhang KR, Zhang YF, Lei HM, Tang YB, Ma CS, Lv QM, et al. Targeting AKR1B1 inhibits glutathione de novo synthesis to overcome acquired resistance to EGFR-targeted therapy in lung cancer. Sci Transl Med. 2021;13:eabg6428.
Hitosugi T, Zhou L, Elf S, Fan J, Kang HB, Seo JH, et al. Phosphoglycerate mutase 1 coordinates glycolysis and biosynthesis to promote tumor growth. Cancer Cell. 2012;22:585–600.
Liang Q, Gu WM, Huang K, Luo MY, Zou JH, Zhuang GL, et al. HKB99, an allosteric inhibitor of phosphoglycerate mutase 1, suppresses invasive pseudopodia formation and upregulates plasminogen activator inhibitor-2 in erlotinib-resistant non-small cell lung cancer cells. Acta Pharmacol Sin. 2021;42:115–9.
Huang K, Liang Q, Zhou Y, Jiang LL, Gu WM, Luo MY, et al. A novel allosteric inhibitor of phosphoglycerate mutase 1 suppresses growth and metastasis of non-small-cell lung cancer. Cell Metab. 2021;33:223.
Huang K, Liang Q, Zhou Y, Jiang L-l, Gu W-m, Luo M-y, et al. A novel allosteric inhibitor of phosphoglycerate mutase 1 suppresses growth and metastasis of non-small-cell lung cancer. Cell Metab. 2019;30:1107-19.e8.
Qiu Y, Yin X, Li X, Wang Y, Fu Q, Huang R, et al. Untangling dual-targeting therapeutic mechanism of epidermal growth factor receptor (EGFR) based on reversed allosteric communication. Pharmaceutics. 2021;13:747.
Yin L, Zhang Y, Yin L, Ou Y, Lewis MS, Wang R, et al. Novel mitochondria-based targeting restores responsiveness in therapeutically resistant human lung cancer cells. Mol Cancer Ther. 2021;20(12):2527–38.
He J, Huang Z, Han L, Gong Y, **e C. Mechanisms and management of 3rd-generation EGFR-TKI resistance in advanced non-small cell lung cancer (Review). Int J Oncol. 2021;59:90.
Planchard D, Feng PH, Karaseva N, Kim SW, Kim TM, Lee CK, et al. Osimertinib plus platinum–pemetrexed in newly diagnosed epidermal growth factor receptor mutation-positive advanced/metastatic non-small-cell lung cancer: safety run-in results from the FLAURA2 study. ESMO Open. 2020;6:100271.
Yi M, Zheng X, Niu M, Zhu S, Ge H, Wu K. Combination strategies with PD-1/PD-L1 blockade: current advances and future directions. Mol Cancer. 2022;21:28.
Gandhi L, Rodriguez-Abreu D, Gadgeel S, Esteban E, Felip E, De Angelis F, et al. Pembrolizumab plus chemotherapy in metastatic non-small-cell lung cancer. N Engl J Med. 2018;378:2078–92.
Paz-Ares L, Luft A, Vicente D, Tafreshi A, Gumus M, Mazieres J, et al. Pembrolizumab plus chemotherapy for squamous non-small-cell lung cancer. N Engl J Med. 2018;379:2040–51.
Zhou C, Wu L, Fan Y, Wang Z, Liu L, Chen G, et al. Sintilimab plus platinum and gemcitabine as first-line treatment for advanced or metastatic squamous nsclc: results from a randomized, double-blind, phase 3 trial (ORIENT-12). J Thorac Oncol. 2021;16:1501–11.
Socinski MA, Jotte RM, Cappuzzo F, Orlandi F, Stroyakovskiy D, Nogami N, et al. Atezolizumab for first-line treatment of metastatic nonsquamous NSCLC. N Engl J Med. 2018;378:2288–301.
Jabbour SK, Berman AT, Decker RH, Lin Y, Feigenberg SJ, Gettinger SN, et al. Phase 1 trial of pembrolizumab administered concurrently with chemoradiotherapy for locally advanced non-small cell lung cancer: a nonrandomized controlled trial. JAMA Oncol. 2020;6:848–55.
Liu D, Gong J, Liu T, Li K, Yin X, Liu Y, et al. Phase 1 study of SHR-1701, a bifunctional fusion protein targeting PD-L1 and TGF-β, in patients with advanced solid tumors. J Clin Oncol. 2021;39:2503–2503.
Sequist LV, Waltman BA, Dias-Santagata D, Digumarthy S, Turke AB, Fidias P, et al. Genotypic and histological evolution of lung cancers acquiring resistance to EGFR inhibitors. Sci Trans Med. 2011;3:75ra26.
Wu L, Ke L, Zhang Z, Yu J, Meng X. Development of EGFR TKIs and options to manage resistance of third-generation EGFR TKI osimertinib: conventional ways and immune checkpoint inhibitors. Front Oncol. 2020;10:602762–862.
Acknowledgements
Not applicable.
Funding
This work was supported by grants from the National Natural Science Foundation of China (Grants 82037718, 81922064, 22177083), the Fundamental Research Funds for the Central Universities (SCU2022D025), 1.3.5 project for disciplines of excellent, West China Hospital, Sichuan University (ZYJC18001), the Sichuan Science and Technology Program (grant number 2022NSFSC1290, 2019YFS0003), and West China Nursing Discipline Development Special Fund Project, Sichuan University (Grant HXHL21011).
Author information
Authors and Affiliations
Contributions
WL, YW, and LO conceived the project, supervised the project, and revised the manuscript. KS, GW, and JP summed up the literature, drafted the manuscript, and drew the figures. JP and JZ collected and organized the inhibitors. JW and GW proofread the structures and figures. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This is not applicable for this review.
Consent for publication
This is not applicable for this review.
Competing interests
The authors declare no competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Shi, K., Wang, G., Pei, J. et al. Emerging strategies to overcome resistance to third-generation EGFR inhibitors. J Hematol Oncol 15, 94 (2022). https://doi.org/10.1186/s13045-022-01311-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13045-022-01311-6