Abstract
Background
Computer-aided diagnosis of skin lesions is a growing area of research, but its application to nonmelanoma skin cancer (NMSC) is relatively under-studied. The purpose of this review is to synthesize the research that has been conducted on automated detection of NMSC using digital images and to assess the quality of evidence for the diagnostic accuracy of these technologies.
Methods
Eight databases (PubMed, Google Scholar, Embase, IEEE Xplore, Web of Science, SpringerLink, ScienceDirect, and the ACM Digital Library) were searched to identify diagnostic studies of NMSC using image-based machine learning models. Two reviewers independently screened eligible articles. The level of evidence of each study was evaluated using a five tier rating system, and the applicability and risk of bias of each study was assessed using the Quality Assessment of Diagnostic Accuracy Studies tool.
Results
Thirty-nine studies were reviewed. Twenty-four models were designed to detect basal cell carcinoma, two were designed to detect squamous cell carcinoma, and thirteen were designed to detect both. All studies were conducted in silico. The overall diagnostic accuracy of the classifiers, defined as concordance with histopathologic diagnosis, was high, with reported accuracies ranging from 72 to 100% and areas under the receiver operating characteristic curve ranging from 0.832 to 1. Most studies had substantial methodological limitations, but several were robustly designed and presented a high level of evidence.
Conclusion
Most studies of image-based NMSC classifiers report performance greater than or equal to the reported diagnostic accuracy of the average dermatologist, but relatively few studies have presented a high level of evidence. Clinical studies are needed to assess whether these technologies can feasibly be implemented as a real-time aid for clinical diagnosis of NMSC.
Similar content being viewed by others
Background
Nonmelanoma skin cancer (NMSC) is by far the most common malignancy in humans, with an estimated 3,300,000 annual cases in the United States alone [1]. Over 95% of NMSC cases are basal cell carcinoma (BCC) and cutaneous squamous cell carcinoma (CSCC) [2], both of which may be readily identified through visual inspection by a skilled dermatologist. However, multiple benign lesions can mimic these cancers, resulting in unnecessary morbidity through invasive biopsies and treatments. For example, the SCREEN study, which included 15,983 biopsies performed in 360,288 adults for suspected skin cancer, found that approximately five biopsies had to be performed to detect one malignant skin lesion of any type [3].
The use of artificial intelligence (AI) as a diagnostic aid is a growing trend in dermatology. These systems generally utilize some form of machine learning (ML), which is a subset of AI involving methods that enable machines to make predictions based on their prior data and experiences. In contrast to conventional models that are explicitly programmed to handle a static set of cases, ML models can derive their own generalizations based on a training set and perform accurately in novel scenarios.
Automated classification of NMSC has been achieved through a variety of modalities, such as Raman spectroscopy, optical coherence tomography, and electrical impedance [4,5,6]. However, the simplest modality is digital photography, often enhanced by a dermatoscope. Given the near ubiquitous use of digital cameras and dermatoscopes in dermatologic practice, digital image-based ML models have the greatest potential for clinical implementation and are thus the focus of this review.
Previous reviews of artificial intelligence and skin cancer have focused on melanoma [7,8,9]. To our knowledge, the present study represents the first systematic review of automated detection of NMSC using digital image analysis. The objectives of this study are to identify which digital image-based ML models have been used to diagnose BCC and CSCC and to assess the evidence for their diagnostic accuracy.
Methods
The review was registered in the PROSPERO international prospective register of systematic reviews (Record number: CRD42017060981) and follows the guidelines of the PRISMA Statement. The PRISMA checklist is included in Additional file 1.
Search strategy
Articles were identified from searches of PubMed, Google Scholar, Embase, IEEE Xplore, SpringerLink, ScienceDirect, Web of Science, and the ACM Digital Library using Boolean operators with no search restrictions. Syntactic modifications were made to accommodate the parameters of the databases while preserving the logic of the search string. The following search string was used:
(Association rule OR Automat* detection OR Classification OR Classifier OR Computer-aided OR Computer-assisted OR Computer vision OR Cluster OR Bayes* OR Deep learning OR Decision tree OR Ensemble OR (Feature AND (extraction OR selection)) OR Genetic algorithm OR Inductive logic OR KNN OR K-means OR Machine learning OR Neural network OR Pattern recognition OR Regression OR Random forest OR Support vector) AND (Basal cell carcinoma OR Squamous cell carcinoma) AND (Skin OR Cutaneous OR Dermatolog*) AND (Dermatoscop* OR Dermoscop* OR Image OR Photograph* OR Picture)
Machine learning terms were taken from textbooks on machine learning and represent the most commonly used models [10, 11]. Note that the search string contained terms to exclude studies of noncutaneous cancers.
Study selection
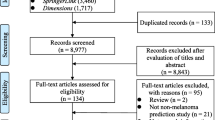
Two investigators extracted data, and results were cross-validated at each step of the selection protocol. Studies were included according to the following selection criteria: (i) classification of NMSC versus benign lesion, (ii) machine learning method, (iii) digital image modality, and (iv) publication in English. Several studies met these criteria but were excluded because they involved classification of both melanoma and NMSC but did not report NMSC-specific performance metrics. We have reported only the NMSC-specific results in studies that classified both melanoma and NMSC. Furthermore, while some studies tested multiple models, we have reported only the model that achieved the highest NMSC-specific performance in each study. References cited in the studies identified from the literature databases served as an additional source of included articles. The selection protocol has been illustrated in the PRISMA flow diagram in Fig. 1.
Quality assessment
The overall quality of each included study was rated according to a modified version of the Levels of Evidence from The Rational Clinical Examination, shown in Table 1 [12]. The original rating scheme specifies the highest level of evidence as blinded, independent studies that compare the diagnostic tool in question against a criterion standard in a large, consecutive sample of patients suspected of having the target condition. Given that all of the included studies were conducted in silico, the interpretation of this definition was modified as follows: (i) blinding was equated to no overlap of images between training and test sets, (ii) independence was equated to avoiding the selective use of images containing features of interest in the test set, and (iii) test sets were considered consecutive if they were obtained from a clinic or set of clinics in which all lesions for which there was any suspicion of malignancy were included. The reporting quality, risk of bias, and applicability of each study was further assessed using the Quality Assessment of Diagnostic Accuracy Studies (2nd edition, QUADAS-2) tool [13].
Results
The database searches returned 8657 total results, of which 2285 were found to be unique after de-duplication. The titles and abstracts of the unique studies were reviewed, and 2211 articles were deemed irrelevant and excluded. Manual review of the references cited within the remaining 74 studies identified seven additional studies of potential relevance, for a total of 81 studies, which were read in their entirety for assessment of eligibility. Of these 81 studies, 42 were excluded due to disqualifying methodologies or insufficient reporting of results. Thus, a total of 39 studies were ultimately included in the review. The characteristics of the included studies are shown in Table 2.
[Table 2. Overview of literature search].
Skin lesion databases
Twenty exclusively on NMSC, whereas the other 17 studies also included classification of melanoma [14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30]. The size of NMSC test sets ranged from as few as ten lesions [22] to as many as 710 [23]. All studies acquired their images either directly from clinics or from publicly available datasets >composed of clinically-obtained and annotated images, with the exception of nine studies that used images of unverifiable origin from online repositories [14, 19, 21, 22, 24,25,26, 31, 32]. Among the studies using clinically-obtained image sets, all NMSC images represented biopsy-proven lesions, and seven studies also used exclusively biopsy-proven benign lesions for their competitive sets [15,16,17, 23, 28, 30, 33].
Eight studies used test sets comprising all lesions examined in a clinic or set of clinics during a specific time frame for which there was any suspicion of malignancy, thus constituting a consecutive sample [15, 17, 30, 34,35,36,37,38]. Two studies, while they did use a set of clinically-obtained images spanning a broad variety of benign lesions suggestive of a consecutive sample, did not explicitly report that this set represented all lesions of suspected malignancy seen in those clinics [39, 40]. The rest of the studies used sets of NMSC lesions and benign mimics chosen by the experimenters. Among the studies using non-consecutive sets, three used actinic keratoses (AKs) as their benign mimic [19, 24, 25], two used seborrheic keratoses (SKs) [16, 41], one used nevi [22], four used SKs and nevi [20, 28, 33, 42], three used AKs, SKs, and nevi [14, 31, 43], one used AKs, SKs, nevi, lentigines, and poromas [18], two used AKs, SKs, nevi, dermatofibromas, and vascular lesions [27, 29], one used AKs, SKs, nevi, dermatofibromas, lentigines, warts, and vascular lesions [23], two used SKs, nevi, and psoriasis [44, 45], one used SKs, nevi, psoriasis, eczema, and seborrheic dermatitis [ Artificial intelligence Actinic keratosis Artificial neural network Area under receiver operating characteristic Basal cell carcinoma Cutaneous squamous cell carcinoma K-nearest neighbors Machine learning Malignant melanoma Multiclass support vector machine Nonmelanoma skin cancer Preferred Reporting Items for Systematic Reviews and Meta-Analyses Quality Assessment of Diagnostic Accuracy Studies, 2nd edition Receiver operating characteristic Seborrheic keratosis Rogers HW, Weinstock MA, Feldman SR, Coldiron BM. Incidence estimate of nonmelanoma skin cancer (keratinocyte carcinomas) in the US population. JAMA Dermatol. 2015. https://doi.org/10.1001/jamadermatol.2015.1187. Gillard M, Wang TS, Johnson TM. Nonmelanoma cutaneous malignancies. In: Chang AE, Ganz PA, Hayes DF, Kinsella T, Pass HI, Schiller JH, et al., editors. Oncology. New York: Springer; 2006. p. 1102–18. Breitbart EW, Waldmann A, Nolte S, Capellaro M, Greinert R, Volkmer B, et al. Systematic skin cancer screening in northern Germany. J Am Acad Dermatol. 2012;66:201–11. Sigurdsson S, Philipsen PA, Hansen LK, Larsen J, Gniadecka M, Wulf HC. Detection of skin cancer by classification of Raman spectra. IEEE Trans Biomed Eng. 2004. https://doi.org/10.1109/TBME.2004.831538. Jørgensen TM, Tycho A, Mogensen M, Bjerring P, Jemec GB. Machine-learning classification of non-melanoma skin cancers from image features obtained by optical coherence tomography. Skin Res Technol. 2008. https://doi.org/10.1111/j.1600-0846.2008.00304.x. Dua R, Beetner DG, Stoecker WV, Wunsch DC 2nd. Detection of basal cell carcinoma using electrical impedance and neural networks. IEEE Trans Biomed Eng. 2004. https://doi.org/10.1109/TBME.2003.820387. Dreiseitl S, Ohno-Machado L, Kittler H, Vinterbo S, Billhardt H, Binder M. A comparison of machine learning methods for the diagnosis of pigmented skin lesions. J Biomed Inform. 2001;34:28–36. Rajpara SM, Botello AP, Townend J, Ormerod AD. Systematic review of dermoscopy and digital dermoscopy/artificial intelligence for the diagnosis of melanoma. Br J Dermatol. 2009. https://doi.org/10.1111/j.1365-2133.2009.09093.x. Korotkov K, Garcia R. Computerized analysis of pigmented skin lesions: a review. Artif Intell Med. 2012. https://doi.org/10.1016/j.artmed.2012.08.002. Kelleher JD, Mac Namee B, D’Arcy A. Fundamentals of machine learning for predictive data analytics: algorithms, worked examples, and case studies. 1st ed. Cambridge: MIT Press; 2015. Bishop C. Pattern Recognition and Machine learning. 1st ed. New York: Springer; 2006. Simel D, Rennie D. Primer on precision and accuracy. In: Simel D, Rennie D, Keitz S, editors. The rational clinical examination: evidence-based clinical diagnosis. New York: McGraw-Hill Medical; 2008. p. 15. Whiting PF, Rutjes AWS, Westwood ME, Mallett S, Deeks JJ, Reitsma JB, et al. QUADAS-2 group. QUADAS-2: a revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med. 2011. https://doi.org/10.7326/0003-4819-155-8-201110180-00009. Abbas Q. Computer-aided decision support system for classification of pigmented skin lesions. Int J Comput Sci Network Security. 2016;16:9–15. Chang WY, Huang A, Yang CY, Lee CH, Chen YC, Wu TY, et al. Computer-aided diagnosis of skin lesions using conventional digital photography: a reliability and feasibility study. PLoS One. 2013. https://doi.org/10.1371/journal.pone.0076212. Esteva A, Kuprel B, Novoa RA, Ko J, Swetter SM, Blau HM, et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature. 2017. https://doi.org/10.1038/nature21056. Ferris LK, Harkes JA, Gilbert B, Winger DG, Golubets K, Akilov O, et al. Computer-aided classification of melanocytic lesions using dermoscopic images. J Am Acad Dermatol. 2015. https://doi.org/10.1016/j.jaad.2015.07.028. Fujisawa Y, Otomo Y, Ogata Y, Nakamura Y, Fujita R, Ishitsuka Y, et al. Deep-learning-based, computer-aided classifier developed with a small dataset of clinical images surpasses board-certified dermatologists in skin tumour diagnosis. Br J Dermatol. 2018. https://doi.org/10.1111/bjd.16924. Maurya R, Singh SK, Maurya AK, Kumar A. GLCM and multi class support vector machine based automated skin cancer classification. Int Conf Comput Sustain Global Dev. 2014. https://doi.org/10.1109/IndiaCom.2014.6828177. Shimizu K, Iyatomi H, Celebi ME, Norton KA, Tanaka M. Four-class classification of skin lesions with task decomposition strategy. IEEE Trans Biomed Eng. 2015. https://doi.org/10.1109/TBME.2014.2348323. Sumithra R, Suhil M, Guru DS. Segmentation and classification of skin lesions for disease diagnosis. Procedia Computer Science. 2015. https://doi.org/10.1016/j.procs.2015.03.090. Wahba MA, Ashour AS, Napoleon SA, Abd Elnaby MM, Guo Y. Combined empirical mode decomposition and texture features for skin lesion classification using quadratic support vector machine. Health Inf Sci Syst. 2017. https://doi.org/10.1007/s13755-017-0033-x. Han SS, Kim MS, Lim W, Park GH, Park I, Chang SE. Classification of the clinical images for benign and malignant cutaneous tumors using a deep learning algorithm. J Invest Dermatol. 2018. https://doi.org/10.1016/j.jid.2018.01.028. Choudhury D, Naug A, Ghosh S. Texture and color feature based WLS framework aided skin cancer classification using MSVM and ELM. Annual IEEE India Conference. 2015. https://doi.org/10.1109/INDICON.2015.7443780. Dorj UO, Lee KK, Choi JY, Lee M. The skin cancer classification using deep convolutional neural network. Multimed Tools Appl. 2018. https://doi.org/10.1007/s11042-018-5714-1. Shoieb DA, Youssef SM, Aly WM. Computer-aided model for skin diagnosis using deep learning. Int J Image Graph. https://doi.org/10.18178/joig.4.2.122-129. Upadhyay PK, Chandra S. Construction of adaptive pulse coupled neural network for abnormality detection in medical images. Appl Artif Intell. 2018. https://doi.org/10.1080/08839514.2018.1481818. Yap J, Yolland W, Tschandl P. Multimodal skin lesion classification using deep learning. Exp Dermatol. 2018. https://doi.org/10.1111/exd.13777. Lee YC, Jung SH, Won HH. WonDerM: Skin lesion classification with fine-tuned neural networks. https://arxiv.org/abs/1808.03426 (2018). Accessed 14 Oct 2018. Møllersen K, Kirchesch H, Zortea M, Schopf TR, Hindberg K, Godtliebsen F. Computer-aided decision support for melanoma detection applied on melanocytic and nonmelanocytic skin lesions: a comparison of two systems based on automatic analysis of Dermoscopic images. Biomed Res Int. 2015. https://doi.org/10.1155/2015/579282. I I. Categorization of non-melanoma skin lesion diseases using support vector machine and its variants. Int J Med Imaging. 2015. https://doi.org/10.11648/j.ijmi.20150302.15. Wahab NA, Wahed MA, Mohamed ASA. Texture features neural classifier of some skin diseases. IEEE Midwest Symposium on Circuits and Systems. 2003. https://doi.org/10.1109/MWSCAS.2003.1562298. Chuang SH, Sun X, Chang WY, Chen GS, Huang A, Li J, et al. BCC skin cancer diagnosis based on texture analysis techniques. SPIE Proc. 2011. https://doi.org/10.1117/12.878124. Cheng B, Erdos D, Stanley RJ, Stoecker WV, Calcara DA, Gómez DD. Automatic detection of basal cell carcinoma using telangiectasia analysis in dermoscopy skin lesion images. Skin Res Technol. 2011. https://doi.org/10.1111/j.1600-0846.2010.00494.x. Stoecker WV, Gupta K, Shrestha B. Detection of basal cell carcinoma using color and histogram measures of semitranslucent areas. Skin Res Technol. 2009. https://doi.org/10.1111/j.1600-0846.2009.00354.x. Cheng B, Stanley RJ, Stoecker WV, Hinton K. Automatic telangiectasia analysis in dermoscopy images using adaptive critic design. Skin Res Technol. 2012. https://doi.org/10.1111/j.1600-0846.2011.00584.x. Cheng B, Stanley RJ, Stoecker WV, Osterwise CT, Stricklin SM, Hinton KA, et al. Automatic dirt trail analysis in dermoscopy images. Skin Res Technol. 2013. https://doi.org/10.1111/j.1600-0846.2011.00602.x. Kefel S, Kefel SP, LeAnder RW, Kaur R, Kasmi R, Mishra NK, et al. Adaptable texture-based segmentation by variance and intensity for automatic detection of semitranslucent and pink blush areas in basal cell carcinoma. Skin Res Technol. 2016. https://doi.org/10.1111/srt.12281. Mishra NK, Kaur R, Kasmi R, Kefel S, Guvenc P, Cole JG, et al. Automatic separation of basal cell carcinoma from benign lesions in dermoscopy images with border thresholding techniques. In: Proceedings of the 12th International Joint Conference on Computer Vision, Imaging and Computer Graphics Theory and Applications; 2017. https://doi.org/10.5220/0006173601150123. Cheng B, Stanley RJ, Stoecker WV, Stricklin SM, Hinton KA, Nguyen TK, et al. Analysis of clinical and dermoscopic features for basal cell carcinoma neural network classification. Skin Res Technol. 2013. https://doi.org/10.1111/j.1600-0846.2012.00630.x. Shakya NM, LeAnder RW, Hinton KA, Stricklin SM, Rader RK, Hagerty J, et al. Discrimination of squamous cell carcinoma in situ from seborrheic keratosis by color analysis techniques requires information from scale, scale-crust and surrounding areas in dermoscopy images. Comput Biol Med. 2012. https://doi.org/10.1016/j.compbiomed.2012.09.010. Wahba MA, Ashour AS, Guo Y, Napoleon SA, Elnaby MM. A novel cumulative level difference mean based GLDM and modified ABCD features ranked using eigenvector centrality approach for four skin lesion types classification. Comput Methods Prog Biomed. 2018. https://doi.org/10.1016/j.cmpb.2018.08.009. Ballerini L, Fisher RB, Aldridge B, Rees J. Non-melanoma skin lesion classification using colour image data in a hierarchical K-NN classifier. IEEE Int Symp Biomed Imaging. 2012. https://doi.org/10.1109/ISBI.2012.6235558.17. Zhang X, Wang S, Liu J, Tao C. Computer-aided diagnosis of four common cutaneous diseases using deep learning algorithm. IEEE Int Conf Bioinform Biomed. 2017. https://doi.org/10.1109/BIBM.2017.8217850. Zhang X, Wang S, Liu J, Tao C. Towards improving diagnosis of skin diseases by combining deep neural network and human knowledge. BMC Med Inform Decis Mak. 2018. https://doi.org/10.1186/s12911-018-0631-9. Zhou H, **e F, Jiang Z, Liu J, Wang S, Zhu C. Multi-classification of skin diseases for dermoscopy images using deep learning. IEEE Int Conf Imaging Syst Techniques. 2017. https://doi.org/10.1109/IST.2017.8261543. Guvenc P, LeAnder RW, Kefel S, Stoecker WV, Rader RK, Hinton KA, et al. Sector expansion and elliptical modeling of blue-gray ovoids for basal cell carcinoma discrimination in dermoscopy images. Skin Res Technol. 2013. https://doi.org/10.1111/srt.12006. Kharazmi P, Lui H, Wang ZJ, Lee TK. Automatic detection of basal cell carcinoma using vascular-extracted features from dermoscopy images. IEEE Can Conf Electric Comput Eng. 2016. https://doi.org/10.1109/CCECE.2016.7726666. Kefel S, Guvenc P, LeAnder R, Stricklin SM, Stoecker WV. Discrimination of basal cell carcinoma from benign lesions based on extraction of ulcer features in polarized-light dermoscopy images. Skin Res Technol. 2012. https://doi.org/10.1111/j.1600-0846.2011.00595.x. Kharazmi P, AlJasser MI, Lui H, Wang ZJ, Lee TK. Automated detection and segmentation of vascular structures of skin lesions seen in Dermoscopy, with an application to basal cell carcinoma classification. IEEE J Biomed Health Inform. 2016. https://doi.org/10.1109/JBHI.2016.2637342. Kharazmi P, Kalia S, Lui H, Wang ZJ, Lee TK. A feature fusion system for basal cell carcinoma detection through data-driven feature learning and patient profile. Skin Res Technol. 2017. https://doi.org/10.1111/srt.12422. Kharazmi P, Kalia S, Lui H, Wang ZJ, Lee TK. Computer-aided detection of basal cell carcinoma through blood content analysis in dermoscopy images. SPIE Proc. 2018. https://doi.org/10.1117/12.2293353. Marks R, Rennie G, Selwood TS. Malignant transformation of solar keratoses to squamous cell carcinoma. Lancet. 1988;(8589):795–7. Presser SE, Taylor JR. Clinical diagnostic accuracy of basal cell carcinoma. J Am Acad Dermatol. 1987;16:988–90. Heal CF, Raasch BA, Buettner PG, Weedon D. Accuracy of clinical diagnosis of skin lesions. Br J Dermatol. 2008. https://doi.org/10.1111/j.1365-2133.2008.08715.x. Monheit G, Cognetta AB, Ferris L, Rabinovitz H, Gross K, Martini M, et al. The performance of MelaFind: a prospective multicenter study. 2011; doi: https://doi.org/10.1001/archdermatol.2010.302. Moher D, Liberati A, Tetzlaff J, Altman DG, PRISMA Group. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. PLoS Med. 2009. https://doi.org/10.1371/journal.pmed.1000097. Zortea M, Schopf TR, Thon K, Geilhufe M, Hindberg K, Kirchesch H, et al. Performance of a dermoscopy-based computer vision system for the diagnosis of pigmented skin lesions compared with visual evaluation by experienced dermatologists. Artif Intell Med. 2014. https://doi.org/10.1016/j.artmed.2013.11.006. Not applicable. JBC, SH, and AM designed the study. AM and ET acquired, analyzed, and interpreted the data. All authors contributed to the drafting of the manuscript. All authors read and approved the final manuscript. Not applicable. Not applicable. Not applicable. Not applicable. The authors have no competing interests. Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations. PRISMA checklist. (DOC 66 kb) Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated. Marka, A., Carter, J.B., Toto, E. et al. Automated detection of nonmelanoma skin cancer using digital images: a systematic review.
BMC Med Imaging 19, 21 (2019). https://doi.org/10.1186/s12880-019-0307-7 Received: Accepted: Published: DOI: https://doi.org/10.1186/s12880-019-0307-7Abbreviations
References
Acknowledgements
Author contributions
Funding
Availability of data and materials
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Consent for publication
Competing interests
Publisher’s Note
Additional file
Additional file 1:
Rights and permissions
About this article
Cite this article
Keywords