Abstract
Background
Schisandra chinensis, an ancient member of the most basal angiosperm lineage which is known as the ANITA, is a fruit-bearing vine with the pharmacological effects of a multidrug system, such as antioxidant, anti-inflammatory, cardioprotective, neuroprotective, anti-osteoporosis effects. Its major bioactive compound is represented by lignans such as schisandrin. Molecular characterization of lignan biosynthesis in S. chinensis is of great importance for improving the production of this class of active compound. However, the biosynthetic mechanism of schisandrin remains largely unknown.
Results
To understand the potential key catalytic steps and their regulation of schisandrin biosynthesis, we generated genome-wide transcriptome data from three different tissues of S. chinensis cultivar Cheongsoon, including leaf, root, and fruit, via long- and short-read sequencing technologies. A total of 132,856 assembled transcripts were generated with an average length of 1.9 kb and high assembly completeness. Overall, our data presented effective, accurate gene annotation in the prediction of functional pathways. In particular, the annotation revealed the abundance of transcripts related to phenylpropanoid biosynthesis. Remarkably, transcriptome profiling during fruit development of S. chinensis cultivar Cheongsoon revealed that the phenylpropanoid biosynthetic pathway, specific to coniferyl alcohol biosynthesis, showed a tendency to be upregulated at the postfruit development stage. Further the analysis also revealed that the pathway forms a transcriptional network with fruit ripening-related genes, especially the ABA signaling-related pathway. Finally, candidate unigenes homologous to isoeugenol synthase 1 (IGS1) and dirigent-like protein (DIR), which are subsequently activated by phenylpropanoid biosynthesis and thus catalyze key upstream steps in schisandrin biosynthesis, were identified. Their expression was increased at the postfruit development stage, suggesting that they may be involved in the regulation of schisandrin biosynthesis in S. chinensis.
Conclusions
Our results provide new insights into the production and accumulation of schisandrin in S. chinensis berries and will be utilized as a valuable transcriptomic resource for improving the schisandrin content.
Similar content being viewed by others
Background
Schisandra chinensis, also known as ‘Omija’ (Korea), ‘Wuweizi’ (China), or ‘Gomishi’ (Japan), is a climbing species with a generation time of five years that belongs to the order Austrobaileyales, which consists of early-diverging angiosperms [1]. The natural habitats of S. chinensis are mostly within northeastern Asia; they show a uniform distribution of genetic diversity and low genetic differentiation because of extensive gene flow [2]. The berry fruits of S. chinensis are clustered in grape-like bunches and exhibit five flavors: salty, sweet, sour, pungent (spicy), and bitter. The fruits contain many bioactive compounds, including lignans, triterpenes, phenolic acids, flavonoids, essential oils, and polysaccharides [3, 4], and they have been used as an important traditional medicine in northeastern Asia. In particular, dibenzocyclooctadiene lignans, including schisandrin and gomisin, present in S. chinensis fruit extracts exhibit pharmacological effects such as antitumor, anti-inflammatory, antioxidative, hepatoprotective, antihypertensive, and anti-osteoporosis activities [3,4,5,6,7]. Moreover, fruits have the potential to effectively protect against neuronal cell damage and to significantly enhance cognitive performance, suggesting their usefulness as new therapeutic agents for treating neurodegenerative diseases [8].
In general, biosynthesis of lignans in plants is linked to phenylpropanoid pathway in an upstream function and branches after synthesis of coniferyl alcohol. A succession of specific steps, involving in catalytic reactions of dirigent (DIR), pinoresinol-lariciresinol reductase (PLR), secoisolariciresinol dehydrogenase (SILD), O-methyltransferases (OMTs; i.e. OMT1 and OMT3), cytochrome P450 families (CYPs; i.e. CYP719A23, CYP71CU1, CYP71BE54, CYP82D61), and UDP-glucose-dependent glucosyltransferases (UGTs) [9,10,11,12,27] (from Jilin, China), CS_F_40 and CS_F_120 (sampled at 40 and 120 DAFs of fruit in Cheongsoon), and SB_F (Sobaeksan), and two leaf samples, including CS_L (Cheongsoon) and SB_L (Sobaeksan), were analyzed
We assessed the quality of the S. chinensis unigenes. First, the Core Eukaryotic Genes Map** Approach (CEGMA), which assesses a highly reliable set of gene annotations in genome and transcriptome assemblies, revealed a completeness of 93.6% (Fig. 1d). Second, various RNA-Seq samples that were generated from leaves and fruits of Cheongsoon and Sobaeksan were mapped to the unigene dataset with coverage ranging from 91.0 to 97.1% (Fig. 1e), showing high data map** coverage. Therefore, our data indicate the high quality of the assembly and map** coverage of the reads for S. chinensis, which is a helpful transcriptome reference dataset.
Functional annotation of unigenes
All unigenes were annotated using homology-based searches. Of all unigenes, 90,930 unigenes (68.4%) showed significant homology with protein sequences in the UniProt (69,782 hits), TAIR (78,952 hits), NCBI NR (87,851 hits), and InterPro (46,222 hits) databases. In the searches, 83,228 (91.5%) annotated unigenes were predicted to be known genes based on UniProt and TAIR, which accept primary sequences derived from sequencing experiments with functional information, showing good annotation quality. In particular, 39,776 of the known unigenes were predicted to be known genes based on the identification of protein domains (Fig. 2a), showing outstanding hit scores. Of the annotated unigenes, 75,748 were identified to be derived from Iso-Seq data, showing efficient transcript annotation by long-read sequencing. We also examined unigenes homologous to known transcription factor (TF) sequences by searching against PlantTFDB using BLASTX with a cutoff of 1E-05. A total of 21,800 unigenes classified into 58 TF families were identified. In the search, the bHLH, NAC, WRKY, MYB/−related, FAR1, C3H, B3, C2H2, ERF, GRAS, bZIP, HB-other, TALE, G2-like, YABBY, and Trihelix families were abundant in S. chinensis, making up the top 30% of all families according to abundance (Fig. 2b). Interestingly, bHLH, NAC, MYB, ERF, GRAS, bZIP, and TALE are involved in fruit development and/or plant growth [30]. The predominant proteins in terms of Pfam domains included P-loop-containing nucleotide triphosphate hydrolase (5.4%), various protein kinases (17.1%), tetra- and penta-tricopeptides (5.7%), and NAD(P)-binding protein (2.1%) (Additional file 4: Fig. S2a). In addition, UniProtKB keywords associated with functional membranes (7%), alternative splicing (3.7%), transferases (3.7%), the nucleus (3.6%), and binding (6.2%) were highly abundant (Additional file 4: Fig. S2b).
Overview of the functional annotation of unigenes in S. chinensis. a. Overlap of hits among known homologous genes obtained from searches against different databases, including UniProt, TAIR, NR, and InterPro. b. The primary TF families identified in unigenes by PlantTFDB search. c. Abundance of unigenes assigned to the top 20 ranked KEGG pathways (left panel) and enrichment of unigenes related to secondary metabolite biosynthesis in the S. chinensis transcriptome dataset (right panel) [28, 29]. KEGG enrichment analysis was performed by using DAVID according to an EASE score < 0.001. Cross mark (‘†’) indicates the primary KEGG pathway focused on in this study
To further predict and classify unigene functions, the annotated unigenes were analyzed according to KEGG pathway assignments. A total of 12,940 unigenes were assigned to 20 primary pathways. The most common pathways were ‘metabolic pathways’, ‘biosynthesis of secondary metabolites’, ‘biosynthesis of antibiotics’, ‘carbon metabolism’, ‘terpenoid biosynthesis’, and ‘biosynthesis of amino acids’ (left panel in Fig. 2c). In particular, the enrichment analysis for transcripts related to ‘biosynthesis of secondary metabolites’ revealed the significance of pathways related to ‘glycolysis/gluconeogenesis’, ‘TCA cycle’, ‘glyoxylate and dicarboxylate metabolism’, ‘phenylpropanoid biosynthesis’, ‘amino acid-related metabolism’, ‘lipid-related metabolism’, ‘terpenoid biosynthesis’, and ‘carotenoid biosynthesis’ (right panel in Fig. 2c). Interestingly, a high abundance of transcripts related to phenylpropanoid biosynthesis was identified. This result seems to suggest that the activation of the phenylpropanoid pathway in S. chinensis is associated with the biosynthesis of polyphenols such as lignans or flavonoids [22, 23]. Our data show that long-read-based transcriptome sequencing resulted in more effective, accurate annotation or prediction for elucidating genes associated with interesting traits or functional pathways in S. chinensis.
Characterization of transcripts in phenylpropanoid and lignan biosynthesis
In relation to the lignan biosynthetic pathway, in the podophyllotoxin pathway, especially the pathway leading to etoposide aglycone, (−)-4′-desmethylepi podophyllotoxin, an identified biosynthetic gene, has been studied in detail [14]. However, the biosynthetic mechanism of schisandrin production remains largely unknown, and only early steps for catalyzing the synthesis of dihydroguaiaretic acid that is converted to the dibenzocyclooctadiene skeleton have been studied [31]. Nevertheless, to understand lignan biosynthesis in S. chinensis, we identified transcripts related to lignan biosynthetic pathways for deoxypodophyllotoxin and dihydroguaiaretic acid and the phenylpropanoid biosynthetic pathway for coniferyl alcohol, which is utilized for activating the lignan biosynthetic pathway (Fig. 3b). For phenylpropanoid biosynthesis, a total of 61 homologous unigenes were identified: 13 PALs, 5 C4Hs, 9 4CLs, 14 HCTs, 4 C3Hs, 5 CCoAOMTs, 3 CCRs, and 8 CADs (Fig. 3a). For lignan biosynthesis, 76 homologous unigenes were also identified: 4 IGSs, 9 DIRs, 4 PLRs, 11 SILDs, 8 OMT1/3 s-like, 31 CYP71CU1s-like, 6 CYP719A23/24 s-like, and 3 2-ODDs (Fig. 3a and b). This result shows the conservation of those biosynthetic pathways in S. chinensis. Of those unigenes identified, unigenes homologous to PAL, HCT, 4CL and CAD for phenylpropanoid biosynthesis and DIR, SILD, OMT1/3, and CYP71CU1 for lignan biosynthesis were relatively abundant (Fig. 3a and b).
Identification of candidate genes related to phenylpropanoid and lignan biosynthetic pathways in S. chinensis. a. Identification of unigenes homologous to genes related to these two biosynthetic pathways. b. Expression of candidate unigenes in the two pathways in leaf and fruit tissues. After quantifying gene expression from RNA-Seq data derived from the leaves and fruits of S. chinensis (Cheongsoon) with five replicates, the log2-scale fold changes of each of gene between leaf and fruit were calculated. In Figure, each small square cell indicates unigene homologous to gene encoding enzyme involved in the pathways. In addition, expression change of each unigene between leaf and fruit is shown with heatmap. Mark ‘x’ in small square cell indicates no unigene. c. Multiple-sequence alignment of DIR, IGS1, and PLR homologs. Sequence conservation was identified among targeted genes (UniProt accession number: Q9SDR7 for DIR, Q15G13 for IGS1, and O65679 for PLR), the corresponding primary protein domains (dirigent for DIR, PCBER_SDR_a for IGS1, and NmrA-like family for PLR), and the corresponding homologous unigenes. d. Phylogeny of IGS1 homologs on the basis of the relationships among major linages of angiosperms sequenced, including basal angiosperms (GenBank accession number: XP_006859191.1 and XP_020532181.1 Amborella trichopoda and RWR83219.1 and RWR85316.1 for Cinnamomum micranthum), rosids (XP_021900462.1 for Carica papaya, XP_007022888.1 for Theobroma cacao, XP_007213116.1 and XP_007212642.2 for Prunus persica, XP_002310455.1 for Populus trichocarpa, and RVX21189.1 for Vitis vinifera), asterids (Q15GI3.1 for Petunia x hybryda and CAA2957992.1 and CAA2959183.1 for Olea europaea) reported by the Amborella Genome Project [32]. e. Phylogenetic analysis between IGS1 and PLR homologs via the maximum likelihood-based method
Among the identified unigenes, full-length ORFs homologous to IGS, DIR and PLR, which essentially contribute to the early step in the biosynthesis of the two lignans, were identified by analyzing conserved protein domains through multiple-sequence alignment with previously reported target genes (Fig. 3c). First, four IGS1 unigenes, KSC_ISO-074408, − 090301, − 004209 and − 08352, which were distinct from eugenol synthase 1 (EGS1) or isoflavone reductase, were identified to contain a phenylcoumaran benzylic ether reductase (PCBER)-like protein domain. IGS1 is an atypical short-chain dehydrogenase/reductase (SDR) identified in Petunia hybrid [33] that acts as an NADPH-dependent aromatic alcohol reductase and functions to catalyze the synthesis of phenylpropene isoeugenol from coniferyl alcohol. IGS1 homologs identified in S. chinensis were subordinate to the basal angiosperm group (Fig. 3d) compared with other key lineages that correspond to mostly tree genomes. Second, five DIR unigenes, including KSC_ISO-007672, − 009738, − 010220, − 008873 and − 002318, were identified with high sequence conservation to the dirigent protein domains of psd_Fi1 and PI206 identified in Forsythia intermedia [34] and Pisum sativum [35]. DIR plays a role in selectively biosynthesizing pinoresinol or verrucosin from two molecules of coniferyl alcohol. Sequence divergences of those DIR unigenes in S. chinensis were identified in their 5′ and 3′ regions. Third, four PLR unigenes, including KSC_ISO-014507, − 042181, − 004036 and − 070096, were identified with high sequence conservation to AtPLR3 of Arabidopsis, which contains a NmrA-type oxidoreductase family protein domain and might be involved in the reduction of lariciresinol into secoisolariciresinol [10, 36]. Interestingly, although IGS1 and PLR harbored NAD(P)-binding motifs (as members of the SDR family), they showed clear sequence divergence (Fig. 3e). These identified genes are potential candidates for elucidating the regulation of lignan biosynthesis in S. chinensis.
Differential expression of transcripts involved in phenylpropanoid and lignan biosynthesis in the fruit and leaf of S. chinensis
We also examined the differential expression patterns of transcripts involved in phenylpropanoid and lignan biosynthesis by analyzing RNA-Seq data derived from the deep red-colored fruit and yellow green leaf of S. chinensis cultivar Cheongsoon at 120 days after flowering (DAF) that may lead to the two biosynthesis at the high level. Overall, most unigenes showed divergent expression in the two tissues (Fig. 3b). Transcriptome diversity was even identified among paralogs or isoforms. These results suggest that the tissue-specific expression of transcripts is involved in these two biosynthetic pathways. In phenylpropanoid biosynthesis, most PAL and HCT homologs showed higher expression patterns in the leaves, but all CAD homologs were remarkably upregulated in the fruit (left panel of Fig. 3b), suggesting the predominant production of coniferyl alcohol by CAD in the fruit S. chinensis. Although indirect evidence was obtained, the high content of schisandrin in the fruit of S. chinensis was actually identified by high-performance liquid chromatography (HPLC) profiling (significance at P < 0.01) (Additional file 4: Fig. S3), supporting this hypothesis. In those two pathways for deoxypodophyllotoxin and dihydroguaiaretic acid, three IGS1 unigenes showed slightly high expression in the leaf, but three DIR and two PLR unigenes showed selectively high expression in the fruit with small fold-changes (right panel of Fig. 3b; KSC_ISO-009738 (1.7-fold change) and − 010220 (2.0-fold change) for DIR, and KSC_ISO-014507 (1.8-fold change) and − 042181 (2.2-fold change) for PLR). Interestingly, most SILD unigenes, which are involved in matairesinol biosynthesis, were upregulated in the fruit, especially KSC_ISO_081641 (71.4-fold change in the fruit) (right panel of Fig. 3b), suggesting the predominant production of matairesinol in the fruit of S. chinensis. Our results show the tissue-specific expression of transcripts in the phenylpropanoid and lignan biosynthetic pathways in S. chinensis and their expression diversity.
Transcriptome profiling during fruit development in S. chinensis
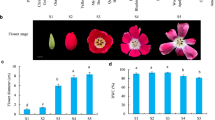
The amounts of dibenzocyclooctadiene lignans in the fruits of S. chinensis are known to be influenced by the degree of fruit maturity and harvest time [3]. To understand transcriptomic changes during fruit development in S. chinensis, we analyzed differentially expressed genes (DEGs) using RNA-Seq data derived from two different fruit development stages in S. chinensis cultivar Cheongsoon, 40 (green-colored and circle-shaped fruit berry) and 120 (deep red-colored and circle-shaped fruit berry) DAF (Fig. 4a), with cutoffs of q < 0.05, an absolute fold change ≥1.5, and an TPM ≥ 1 as the minimum average expression value between genes in the two stages. A total of 16,698 DEGs, consisting of 9344 downregulated and 7354 upregulated genes at 120 DAF, were identified and subjected to functional categories with Gene Ontology (GO) enrichment analysis. The GO enrichment analysis for genes upregulated at 120 DAF, 904 DEGs, revealed the significance of the functions associated with ‘fruit development’ (1.39 × 10− 12), ‘Golgi vesicle transport’ (1.04 × 10− 8), ‘response to plant hormone signaling pathways’ such as abscisic acid (ABA) (4.95 × 10− 16), ethylene (1.83 × 10− 5), and jasmonic acid (JA) (1.6 × 10− 7), ‘phenylpropanoid biosynthetic process’ (6.87 × 10− 5), ‘vitamin biosynthetic process’ (2.69 × 10− 4), and ‘response to hydrogen peroxide (H2O2)’ (3.34 × 10− 4) (Fig. 4b). Remarkably, the pathway for coniferyl alcohol production within the phenylpropanoid biosynthetic pathway is likely to be activated as a specific synthesis route of lignan. In this pathway, the expression of 10 PAL, 3 C4H, 9 4CL2, 1 HCT, 2 C3H, 1 CCoAOMT, 2 CCR and 4 CAD unigenes was significantly increased (Fig. 4c). In particular, three CAD homologous unigenes (KSC_ISO-012053, − 040169 and − 069272) showed an increase in expression with ≥2.7 ~ 10-fold changes (Additional file 4: Fig. S4). As presented in Additional file 4: Fig. S6, the qRT-PCR data for these DEGs were generally similar with the RNA-seq data, showing a positive correlation (r = 0.601). Interestingly, the phenylpropanoid biosynthetic pathway was extensively linked to biosynthetic processes such as those for lignin (FDR = 5.33 × 10− 30), suberin (3.54 × 10− 11), proanthocyanidin (1.04 × 10− 8), flavonoid (8.04 × 10− 9), phenol-containing compound (2.09 × 10− 8), cinnamic acid (2.53 × 10− 6), karrikin (1 × 10− 4), and cell wall (2.2 × 10− 3) (Fig. 4d). This result supports the potential for the activation of other secondary metabolites by the genes involved in the phenylpropanoid biosynthetic pathway at the postfruit development stage of S. chinensis. Therefore, our results indicate the activation of the phenylpropanoid biosynthetic process for lignan biosynthesis at the postfruit development stage of S. chinensis. Moreover, genes related to the response to ABA, ethylene-activated signaling, and response to H2O2, which are known to be associated with fruit ripening [37], are upregulated, implicating a functional link between lignan biosynthesis and ripening. We also identified unigenes upregulated in lignan biosynthetic pathways at 120 DAF (Fig. 4e): 2 IGS1s, 1 DIR, 3 SILDs, 3 OMT1/3, and 10 CYP71CU1. However, unlike IGS1 and DIR unigenes, PLR unigenes showed upregulation at 40 DAF. Additionally, most SILD and CYP719A23 unigenes were upregulated at 40 DAF. Collectively, our results show the transcriptional changes in the phenylpropanoid and lignan biosynthetic pathways at postfruit development in S. chinensis.
Profile of differentially expressed genes (DEGs) between the green- and red-colored berry stages of S. chinensis. a. Fruit shape at 40 (green-colored berry) and 120 (red-colored fruit berry) days after flowering (DAF) in S. chinensis. b. Gene ontology (GO) enrichment for DEGs between 40 DAF and 120 DAF. The GO enrichment analysis for up- (904 genes) and down- (921 genes) regulated genes at 120 DAF (left panel in Figure) was performed by using DAVID according to an EASE score < 0.001, and significant, uniquely represented GO terms were then selected. c. Activation of the phenylpropanoid biosynthetic pathway to coniferyl alcohol at 120 DAF and identification of the corresponding unigenes. d. Map** to genes in the phenylpropanoid biosynthetic process to other closely related biosynthetic processes, including lignin, suberin, flavonoid, proanthocyanidin, phenol-containing compound, cinnamic acid, karrikin, and cell wall biosynthetic processes. e. Identification of DEGs related to lignan biosynthetic pathways at 40 and 120 DAF. f. Transcriptional network among genes upregulated at 120 DAF related to phenylpropanoid biosynthesis and the responses to ABA, JA, ethylene, and hydrogen peroxide. After selecting up-regulated genes enriched in those five GO terms, the interaction between those genes was identified by using STRING and further analyzed using Cytoscape on the basis of the degree of connectivity of the nodes. In the network, the interaction between genes in ‘phenylpropanoid biosynthetic process’ and other functional terms, including ‘responses to ABA’, ‘JA mediated signaling’, ‘ethylene-activated signaling’, or ‘response to hydrogen peroxide’, are indicated with red-color dot line. In addition, genes in phenylpropanoid pathway are indicated with orange-color letter
Transcriptional network between phenylpropanoid biosynthesis and plant hormone signaling
To investigate the functional links between phenylpropanoid biosynthesis and ripening-related plant hormone signaling in S. chinensis, a transcriptional network of genes upregulated at 120 DAF, which were enriched in the functional categories of ‘phenylpropanoid biosynthetic process’ and ‘responses to plant hormone signaling pathways’, including ABA, ethylene, and JA, and ‘response to H2O2’, was analyzed. The analysis showed that the phenylpropanoid biosynthetic pathway highly interacted with the ABA and JA signaling pathways (Fig. 4f). In the network, CYP84A1 (FAH1), LDOX, CAD5, TT2, IRX12, AHA10, and OMT1, which are involved in the phenylpropanoid biosynthetic process, interacted with ABI3, LHY, CCA1, MSS1, FLDH, MYB43, and ARF2 during ABA signaling and IRX1, GL3, and EGL3 during JA signaling. Three genes, AHA10, MTO3, and MAT3, for the phenylpropanoid biosynthetic process, also interacted with RCI1A during ethylene signaling, which is linked to ABA signaling. In connection with plant hormone signaling, IGS1 and DIR interacted with CCoAOMT1 and POM1 in the phenylpropanoid biosynthetic process, respectively, and IGS1 was linked to VLN4. Additionally, the hydrogen peroxide (H2O2) signaling pathway, including CAT1, ADH1 and MIPS2, also interacted with genes involved in phenylpropanoid biosynthetic processes, including TT4, CCoAOMT1, C4H, and 4CL5. Hydrogen peroxide signaling play an important role in promoting ripening at the molecular level [38]. However, the functions of those three genes associated with fruit ripening remain unknown. These results indicate that the phenylpropanoid biosynthetic pathway is closely connected with the ABA, JA, ethylene, and hydrogen peroxide signaling pathways in the postfruit development stage of S. chinensis and that crosstalk may affect the activation of lignan biosynthesis.