Abstract
Background
Bayesian relaxed-clock dating has significantly influenced our understanding of the timeline of biotic evolution. This approach requires the use of priors on the branching process, yet little is known about their impact on divergence time estimates. We investigated the effect of branching priors using the iconic cycads. We conducted phylogenetic estimations for 237 cycad species using three genes and two calibration strategies incorporating up to six fossil constraints to (i) test the impact of two different branching process priors on age estimates, (ii) assess which branching prior better fits the data, (iii) investigate branching prior impacts on diversification analyses, and (iv) provide insights into the diversification history of cycads.
Results
Using Bayes factors, we compared divergence time estimates and the inferred dynamics of diversification when using Yule versus birth-death priors. Bayes factors were calculated with marginal likelihood estimated with step**-stone sampling. We found striking differences in age estimates and diversification dynamics depending on prior choice. Dating with the Yule prior suggested that extant cycad genera diversified in the Paleogene and with two diversification rate shifts. In contrast, dating with the birth-death prior yielded Neogene diversifications, and four rate shifts, one for each of the four richest genera. Nonetheless, dating with the two priors provided similar age estimates for the divergence of cycads from Ginkgo (Carboniferous) and their crown age (Permian). Of these, Bayes factors clearly supported the birth-death prior.
Conclusions
These results suggest the choice of the branching process prior can have a drastic influence on our understanding of evolutionary radiations. Therefore, all dating analyses must involve a model selection process using Bayes factors to select between a Yule or birth-death prior, in particular on ancient clades with a potential pattern of high extinction. We also provide new insights into the history of cycad diversification because we found (i) periods of extinction along the long branches of the genera consistent with fossil data, and (ii) high diversification rates within the Miocene genus radiations.
Similar content being viewed by others
“Cycads are to the vegetable kingdom what Dinosaurs are to the animal, each representing the culmination in Mesozoic times of the ruling Dynasties in the life of their age.”
Lester Ward, 1900
Background
Our understanding of biotic evolution relies heavily on phylogenetic and dating reconstructions that provide insight into the periods of major diversification [1]. In the last decade, the advent of molecular dating approaches has fostered an explosion of studies constructing time-calibrated trees for diverse plant clades like bryophytes [2], ferns [3,4], gymnosperms [5-8] and angiosperms [9-11]. These dated trees have permitted the study of character evolution via the reconstruction of ancestral traits [11], inference of biogeographical history [7], as well as estimates of diversification rates [2,8,10]. Dated trees are thus pivotal to our understanding of the evolution of plants, and of the groups that interact with them, such as herbivores and pollinators [12].
Despite the importance of reliable estimates of divergence times, our understanding of the temporal patterns of diversification remain in flux for many groups, in part because the methods for estimating evolutionary timescales from DNA sequences are being refined [1,13,14]. Since the introduction of relaxed-clock methods, which allow substitution rates to vary across the tree, a range of molecular dating methods has been developed. Bayesian inference has received the most attention because of the flexibility with which different parameters and prior assumptions can be incorporated, and the fact that priors are updated as part of the analysis [15,16]. The use of explicit prior distributions is central to the Bayesian perspective; however, the critical role of prior selection in Bayesian analysis is not always fully appreciated [17].
In Bayesian relaxed-clock (BRC) approaches, there are various types of priors, including priors on calibration points, branch-lengths, clock models, and branching processes. Priors on calibration points have been well studied [18] and, not surprisingly, the choice of these priors can affect estimates of node ages [19]. The effects of branch-length priors on posterior probabilities have been studied; priors assuming long internal branches cause high posterior probabilities [20]. In comparison, the impact of different branching process priors has been relatively under-explored [15,21].
The branching process prior (BPP), also called the ‘tree shape’ or ‘speciation tree’ prior, is a prior model on how trees are generated. Phylogenetic trees are the result of speciation and extinction events, and their relative roles can be varied and represented as different models of diversification [22]. These models effectively place a prior on how phylogenetic trees grow. Probability distributions over models of diversification were employed in some of the earliest attempts to use likelihood techniques to reconstruct genealogies [23]. The two most commonly used BPPs are the Yule (also called ‘pure-birth’) process, which models tree formation with a constant rate of speciation and no extinction, and the birth-death process, which includes speciation as well as a constant rate of lineage extinction. Birth-death priors have been used in Bayesian phylogenetics [24]; however, most published analyses use the Yule prior, perhaps because it was initially the only prior for the diversification process implemented in the widely used Bayesian software package BEAST [16]. Although the birth-death prior has recently been integrated into this software [25,6] did not assess the support of the birth-death prior versus other priors.
This study has four objectives: (i) investigating the impact of the BPP on the dating of ancient clades using the cycads as an example, (ii) assessing whether the birth-death prior is statistically supported, (iii) studying the difference between the Yule and birth-death prior on our understanding of cycad diversification, and (iv) providing a cycad timetree reconciling fossil and phylogenetic data. We also discuss potential explanations for the differences obtained when using different priors and the consequences of prior choice for Bayesian molecular dating.
Methods
Taxon sampling and molecular dataset
We extended the molecular dataset of Nagalingum et al. [6] that initially contained 199 cycad species. We added molecular data for 38 additional species and the same genes retrieved from Genbank [52-57]. Our dataset comprises three genes covering two plastid genes: the maturase K (matK, 2,387 nucleotides, 82 taxa) and the ribulose 1,5-bisphosphate carboxylase large subunit (rbcL, 1,398 nucleotides, 80 taxa), and one nuclear gene: region 1 of phytochrome (PHYP, 1,802 nucleotides for 201 taxa). This resulted in a total of 237 species out of 331 described species (71%), representing all extant genera with the following number of species per genus: 2 of the 2 Bowenia (B. serrulata and B. spectabilis), 65 of the 107 Cycas, 24 out of 27 Ceratozamia, 8 of the 14 Dioon, all of the 65 Encephalartos, 2 of the 2 Lepidozamia (L. hopei and L. peroffskyana), 26 of the 41 Macrozamia, the monotypic Microcyas calocoma, the single Stangeria eriopus, and 43 out of 71 Zamia (including the junior synonym genus Chigua) (Figure 1). Following Nagalingum et al. [6], the dataset also included six species as outgroups: Ginkgo biloba (Ginkgoales), which is recognised as the sister lineage of cycads [58]; plus five conifer species (Abies firma, Araucaria heterophylla, Cryptomeria japonica, Pinus strobus and Pseudotsuga menziezii). The phylogenetic analyses were thus performed on a dataset containing a total of 243 species (of which 237 are cycads) and 5587 nucleotides. The information on each sampled species is presented on Additional file 1: Table S1.
The node calibration procedure used for dating the cycads. (a) Phylogenetic tree of cycads showing the relationships among the 10 genera. Genera are represented by triangles proportional to their species richness. Numbers in parenthesis are the number of species sampled, and the total number of species, within each genus. Pictures illustrate Cycas, Dioon, Zamia, Macrozamia, and Encephalartos species. Black dots indicate the four ‘traditional’ fossil calibrations used for dating, and red dots indicate the two new fossil calibrations evaluated in this study. (b) Information related to the four fossil calibrations (FC, see the text for more details). Ma, million years ago.
Fossil calibrations
To calibrate the cycad tree, we used two fossil datasets. First, we followed Nagalingum et al. [6] and used four cycad fossil constraints (FC1-4), which were retrieved from a careful examination of the cycad fossil record ([32]; Figure 1). These four fossils have also been used by Salas-Leiva et al. [43]. Hereafter this fossil dataset is referred to as the ‘traditional fossil dataset’ (FC1-4). Second we used these four fossils and added two fossils based on the phylogenetic analyses of Hermsen et al. [32] and Martínez et al. [59]. The fossil record was re-examined after new fossils from the Cretaceous were discovered, which did not affect overall fossil assignments, but allowed the use of new fossils to calibrate the cycad tree [59], two of which are tentatively used here. Hereafter this fossil dataset is referred to as ‘new fossil dataset’ (FC1-6).
Based on the presence of synapomorphies linking the fossils to the extant clades, all fossils were used as minimum age constraints for specific nodes ([32,59], Figure 1). However, since synapomorphies can evolve anywhere along the stem branch and the fossil may attach anywhere along this branch [60], we used a commonly employed approach whereby the fossils constrain the stem rather than crown node [61,62]. The absolute fossil ages we used (detailed below) are slightly different from those used in Nagalingum et al. [6], mainly due to an updated geological timescale [63]. Dating fossils older than ~50,000 years is difficult, which means that those fossils are typically assigned to a stratigraphic interval, for example, the late Miocene. The ages assigned below are the ages of the youngest of the boundaries of stratigraphic interval within which the fossil was found (the fossil will actually be older than this age designation).
Cycad stem (FC1, Younger = 265.1 Ma and Older = 364.7 Ma)
The oldest possible records of the group Cycadophyta are microstrobili (e.g. [64]) and megasporophylls (e.g. [65,66]), of which †Crossozamia (eight known species) is the least equivocal in terms of phylogenetic affinity [32,59,66]. Crossozamia consists of megasporophylls with similar morphology to the extant Cycas [65,66]. Although Hermsen et al. [32] identified Crossozamia as being sister to Cycas, the two characters supporting this relationship do not provide strong evidence for Crossozamia belonging to crown group cycads [59]. In addition, the loosely aggregated cone is most probably an ancestral state within cycads. Therefore the divergence between cycads and Ginkgo biloba was constrained using Crossozamia. The age for the host rock formation (Shihhotse) that preserves the Crossozamia fossils was initially described as lower Permian, but the age of the Shihhotse Formation has been revised to the Roadian-Wordian (265.1-272.3; [63]) in the middle Permian [67]. We used the minimum age of this time interval (265.1 Ma) as a minimum age for the stem of cycads. This age was also used as a conservative maximum possible age for the nodes with internal fossil calibrations (FC2-FC4).
Dioon stem (FC2, Younger = 56 Ma and Older = 265.1 Ma)
The origin of the stem group of Dioon was constrained by fossil leaves of †D. inopinus and †D. praespinulosum [44], which have synapomorphies consistent with the leaflet venation anastomoses and leaflet insertion on the rachis of extant Dioon [32]. While originally described as Eocene by Hollick [44], a more recent interpretation of the composition of the Hamilton Bay flora (Kootznahoo Formation), Kupreanof Island, Alaska, from which fossils of both Dioon were originally described, suggest they are of Paleocene age [6,32]. Therefore the Dioon stem was assigned a minimum age of 56 Ma [63].
Bowenia stem (FC3, Younger = 33.9 Ma and Older = 265.1 Ma)
Bowenia leaf fossils of †B. eocenica and †B. papillosa [68], both found in Australia, were used to constrain the divergence of Bowenia from other genera. These fossils were identified as members of the extant genus based on cuticular characters (i.e. number of subsidiary cells) and leaflet morphology (i.e. venation and serrated margin). Both fossil species were found in Australian Eocene deposits (at Anglesea, Victoria for †B. eocenica, and at Nerriga, New South Wales for †B. papillosa), which confers a minimum age constraint at 33.9 Ma for the Bowenia stem [63].
Lepidozamia stem (FC4, Younger = 33.9 Ma and Older = 265.1 Ma)
Fossil Lepidozamia leaves of †L. hopeites and †L. foveolata are also from Australia [45]. Fossils of this genus can be identified by cuticular characters (orientation of the epidermal cells relative to axis of the pinna) that are unique to Lepidozamia and support affinity to the extant genus [32,45]. Found in the Australian Eocene (at Nerriga, New South Wales for †L. foveolata), this set a minimum age of 33.9 Ma for the Lepidozamia stem [63].
Cycad crown (FC5, Younger = 235 Ma and Older = 364.7 Ma)
The age of modern cycads can be calibrated with †Antarcticycas schopfii [69]. The fossil Antarcticycas has been extensively studied [32,70]. A reconstruction of the plant habit has even been proposed [70]. This makes Antarcticycas the most completely known of the extinct Triassic cycad taxa, if not of all fossil cycads, due to the presence of anatomically preserved organs [70]. Interestingly, phylogenetic studies have also tentatively assigned Antarcticycas within modern cycads, and close to the crown of cycads [32,59]. None of the extant genera are phylogenetically sister to Antarcticycas, which makes it a valuable fossil calibration for the crown of cycads. Antarcticycas was found in the Fremouw Formation of the early Middle Triassic of Antarctica. We used a conservative age of the Middle Triassic, i.e. 235 Ma [63], although there remains uncertainty on the true age of the Fremouw Formation with some authors suggesting an Anisian age (242-247.2 Ma).
Encephalarteae stem (FC6, Younger = 72.1 Ma and Older = 265.1 Ma)
Many fossils have been attributed to the tribe Encephalarteae based on morphological and phylogenetic evidence [32,59]. Among them the recently discovered †Wintucycas stevensonii [59] is one of the best-known fossils. Wintucycas has features that clearly allow us to assign it to Encephalarteae due to their manoxylic wood, centripetal polyxyly, parenchymatous pith, centrifugal polyxyly and medullary vascular bundles. Wintucycas was found in the Allen Formation (middle Campanian to early Maastrichtian) of the Late Cretaceous of Argentina (Patagonia), providing a minimum age for the stem of Encephalarteae at 72.1 Ma [63].
Tree root height
The tree root height is the divergence between Cycadales and Ginkgoales. Following Clarke et al. [62], a maximum age can be established with the first records of seeds in the form of preovules that satisfy the criteria of the seed habit. These criteria are the possession of a single functional megaspore that is enveloped in a nucellus (considered equivalent to the megasporangium), which is surrounded (to some extent) by an integument or pre-integument and has mechanisms enabling the capture of pollen before seed dispersal [62,71]. All the criteria are first met with †Elkinsia polymorpha found in the VCo spore Biozone, Evieux Formation [71] in the Fammenian (Late Devonian). The VCo spore Biozone spans 364.7-360.7 Ma [63]. A maximum age for gymnosperms is thus 364.7 Ma. We also used this age to set maximum ages for the cycad stem (FC1) and crown (FC5).
Phylogenetic reconstructions
To obtain a starting tree for the dating analyses, Bayesian inferences were performed with MrBayes 3.2.3 [72]. To determine the best-fit partitioning scheme of molecular evolution for our dataset, we used PartitionFinder 1.1.1 [73]. For the PartitionFinder analyses, branch lengths were unlinked to allow them to be independently estimated for each partition. The searched for best model, among those available in MrBayes, was performed under the greedy algorithm based on the Bayesian Information Criterion model metric. We used the partitioning scheme and among-site rate variation suggested by PartitionFinder, but instead of selecting one substitution model a priori, we used reversible-jump Markov Chain Monte Carlo (rj-MCMC) to allow sampling across the entire substitution rate model space [74].
MrBayes analyses consisted of four rj-MCMC running for 100 million generations with sampling every 10,000 generations and the first 25% discarded as burn-in. We specified (i) a uniform prior probability of phylogenies (i.e. all possible trees are considered a priori equally probable), and (ii) a uniform prior probability distribution on branch lengths. The convergence of the runs was assessed by checking the potential scale reduction factor (PSRF) values of each parameter in MrBayes and the Effective Sample Size (ESS) values of each parameter in Tracer 1.6 [4). Generic stem ages and some ages of deeper nodes were also inferred to be older with the Yule than with the birth-death prior, although the difference was not as marked as in the case of the genus crown ages (Figure 5). However, the birth-death and Yule priors usually inferred similar ages for the cycad stem and crown ages (Figure 5). Analyses with the birth-death process inferred a very high estimate of the relative extinction rate (ratio of extinction to speciation, or turnover) with a median = 0.966 for the dating with four FC (95% HPD 0.9255-0.9972), and a median = 0.962 for the dating with six FC (95% HPD 0.9153-0.9947).
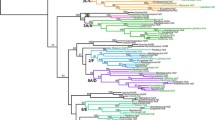
Time-calibrated phylogeny of Cycadales obtained with the four fossil calibrations. Timetree obtained with the Yule (a) or the birth-death (b) model as branching process prior. Each tree is the maximum clade credibility tree with median ages from the Bayesian analyses. The coloured dots highlight nodes on which fossil age constraints were applied. Values are median age estimates for the main nodes, in million years. C, Carboniferous; P, Permian; T, Triassic; J, Jurassic; K, Cretaceous; Pg, Paleogene; N, Neogene. The last geological period, the Quaternary, is missing.
Time-calibrated phylogeny of Cycadales obtained with the six fossil calibrations. Timetree obtained with Yule (a) or the birth-death (b) model as branching process prior. Each tree is the maximum clade credibility tree with median ages from the Bayesian analyses. The coloured dots highlight nodes on which fossil age constraints were applied. Values are median age estimates for the main nodes, in million years. C, Carboniferous; P, Permian; T, Triassic; J, Jurassic; K, Cretaceous; Pg, Paleogene; N, Neogene. The last geological period, the Quaternary, is missing.
Credibility intervals (95% highest posterior density) for the crown ages of the six most species-rich genera. The blue bars depict the age estimates when using the Yule prior, and the green ones show the age estimates obtained with the birth-death prior, shown for the five dating analyses using most fossil calibration points. Analyses with the Yule prior indicate a Paleogene origin, while analyses with the birth-death prior indicate a Neogene origin. Absolute ages are in million years. Q, Quaternary.
Credibility intervals (95% highest posterior density) for the ages of the six deepest nodes. The blue bars depict the age estimates when using the Yule prior, and the green ones show the age estimates obtained with the birth-death prior, shown for the five dating analyses using most fossil calibration points. The numbers associated to each clade correspond to the numbers on the phylogeny. Analyses with the Yule prior consistently indicate older ages. Absolute ages are in million years. The last geological period, the Quaternary, is missing.
Bayes factor values, calculated with the marginal likelihood estimates of the step**-stone analyses, support the birth-death process as the best tree prior (Table 1). Of the two pairs of BEAST analyses (four FC and six FC), BF values are above the standard threshold: BF4FC = 26.66 between the Yule and birth-death prior, and BF6FC = 36.3 between the Yule and birth-death prior. On the contrary, the HME found the opposite, that is the Yule model is the best prior for both datasets for all analyses (Table 1). The results for the cycads thus support previous results showing that the HME estimator is not reliable [76].
Macroevolutionary rates through time
The BAMM analyses converged for both trees (ESSYule = 719.6, Additional file 3: Figure S2; ESSbirth-death = 750.6, Additional file 4: Figure S3). The choice of the BPP had a major influence on the number of different evolutionary regimes detected in the history of the cycads, as well as on the estimation of speciation and extinction rates across the tree (Figure 6). Analyses with the Yule prior supported a model with three evolutionary regimes (i.e. two rate shifts, Additional file 5: Figure S4) located at the crown of the genera Cycas and Encephalartos (according to BF values > 10, and BF = 154.5 over the null model). On the other hand, analyses with the birth-death prior supported a model with five evolutionary regimes (i.e. four rate shifts, Additional file 6: Figure S5) located at (or near) the crown of the four most species-rich genera (according to BF values > 10, and BF = 29.3 over the null model). Moreover, both speciation and extinction rates estimated with the birth-death prior are twice as high as those estimated with the Yule prior (see scales on Figure 6). The credible set of shift configurations with the highest posterior probabilities is provided in Additional file 7: Figure S6 for each tree. The best configuration shift is provided in Additional file 8: Figure S7 for each tree. The macroevolutionary cohort analyses showed distinct evolutionary trajectories for the four richest cycad genera with the birth-death prior (Additional file 9: Figure S8), but not with the Yule prior (Additional file 10: Figure S9).
Diversification pattern of Cycadales. Estimates of speciation (a,c) and extinction (b,d) rates along the cycad phylogeny obtained from BAMM analyses, when considering the chronogram reconstructed with the Yule (a,b) or the birth-death (c,d) prior. Colours at each point in time along branches denote instantaneous rates of speciation or extinction inferred as the mean scenario, with colours indicating mean rates across all the shift configurations sampled in the Bayesian posterior. The diversification scenarios obtained with one versus the other prior are strikingly different. Note the differences for the estimated speciation and extinction rates with each tree prior (rates are twice higher with the tree constrained with a birth-death process). P, Permian; T, Triassic; J, Jurassic; K, Cretaceous; Pg, Paleogene; N, Neogene. The last geological period, the Quaternary, is missing.
Discussion
The branching process prior impacts Bayesian divergence time estimates
Our Bayesian dating analyses highlight an important effect of the branching process prior on the divergence times of cycads. Dating with the Yule prior consistently inferred ages for genus crown groups that were about three times older than those obtained with the birth-death prior. The cycad tree dated with the birth-death process suggested the initiation of crown radiations of the species-rich genera in the Neogene, whereas the Yule model indicated that the radiations began in the Paleogene. These differences in crown ages are noteworthy because the 95% credibility intervals for these nodes did not overlap. It is important to note that the effect of BPP is reduced towards the origin of cycads since, for both priors, genus radiations are subtended by long stem branches originating in the Triassic or Permian. This pattern is recovered for both calibration strategies. Therefore, the BPP is an important influence on the Bayesian dating analyses.
Potential causes of age differences between branching process priors
It is unclear why differing tree priors have a major impact on divergence time estimates. The priors differ principally in the inclusion of an extinction rate. In the Yule prior, there is no extinction, but in the birth-death prior there is a parameter for the extinction rate. The effect of extinction in the birth-death prior is that there are relatively greater nodes toward the present compared to the past, because extinction has not yet had an effect on the more recent nodes, this phenomenon is known as the “pull of the present” [25]. This pattern could explain the contrasting results found when dating the cycad phylogeny with a birth-death versus a Yule prior. Support for the birth-death prior includes fossil evidence indicating that extinctions occurred and were important in sha** cycad evolution. More generally, extinction is a dominant feature of life given that 99.9% of all species that ever existed are now extinct [101]. But the role and effect of extinction could also potentially explain why the effect of BPPs appears to be taxon specific (i.e. little effect in Brassicaceae [28]). Indeed, it is possible that older clades might be more sensitive to the BPP choice because extinction is likely to be a more important component of their evolutionary histories. Earlier simulations and empirical studies have indicated that our understanding of the statistical properties of the diversification prior combined with prior distribution of calibrations is incomplete, particularly when the tree topology is considered [78,102,103]. It is an open question as to how important BPP choice will be for other groups, but further analyses of additional taxa as well as simulation studies will be valuable in understanding the detailed effects that tree priors have on age estimates.
Other potential biases in the branching process priors
While accounting for extinction in BPP priors is an important first step, a birth-death model with homogeneous rates across time and across the various clades within large groups is still an oversimplified model of their evolution. In particular, the hypothesis of rate homogeneity is thought to strongly bias extinction rate estimates [90,93]. We suggest that more complex priors should be developed for implementation in dating software; indeed evidence for age- or time-dependent diversification in numerous clades have resulted in development of methods to take into account rate-heterogeneity through time and clades [93,104].
Another potential bias in BPP priors is the use of models assuming complete or uniform sampling. Phylogenies are often sparsely sampled, particularly in long-branched outgroups. Sparsely sampled outgroups violate the basic assumption of most (if not all) the tree priors implemented in Bayesian dating, namely that the taxon sampling is random and consistent across all clades in the tree (see p. 98 of Drummond and Bouckaert [105]). In this case, it is difficult to find appropriate priors describing the tree: neither the single-parameter Yule model nor the two-parameter birth-death model fit a situation in which some parts of the tree are densely sampled, while others consist of a single long-branched species (i.e. Ginkgo versus the cycads).
Implications for Bayesian dating analyses
The vast majority of molecular dating studies using BEAST have relied on the Yule model, although some recent studies have used both the Yule and birth-death process [27,28,106]. However these studies did not find any differences in age estimates between the two models, nor they did not perform MLE and Bayes factors analyses. Here we tested competing evolutionary hypotheses using a Bayesian relaxed-clock under two different branching priors, and we found stronger support for the birth-death than the Yule process as a prior to reconstruct the cycad tree.
Model selection is a necessary step, because as in the case of cycads, prior choice can have important implications if one simply reconstructs the tree using a Yule model as commonly performed. Conclusions on the age, diversification and/or biogeographic patterns might be different regarding the BPP (see below). Supposedly ancient plant clades might be affected by prior choice as for cycads; specifically, clades with an old and well-documented fossil record that probably experienced periods of extinction such as conifers [7,8], ferns [3,4], gnetophytes [5], and early diverging angiosperm lineages [107,108]. One solution might be to replace the commonly used Yule model by the birth-death process, which is more realistic since all clades are likely to have experienced extinction, even young clades currently diversifying.
As a cautionary tale, we suggest that users routinely test between priors to select the prior that best fit their data. Estimators of the marginal likelihood [82,6). Fossil data and cladistic analyses suggest that, for instance, the lineage today represented by Stangeria might have been more diverse in the past [32,59,118]. The surviving cycads re-diversified from the mid- to late Miocene when five lineages, isolated on different continents, underwent successful independent increases in diversification rate. Therefore our results favour the hypothesis of few and sudden extinction events. As potential explanations, this diversification pattern may be attributed to (i) competition with angiosperms (e.g. [34,35,112,113,119]), (ii) changes in, and final extinction of, non-avian dinosaur faunas that may have acted as seed dispersers (for Mesozoic extinction) [41], and/or (iii) global climate change that led to latitudinal shifts of the main vegetation belts and the elimination of cycads from higher latitudes of Eurasia, North America, and Patagonia [6,110,120], perhaps around the Eocene-Oligocene boundary that marked a significant shift from greenhouse to icehouse climate [38,121].
Discussions on cycad diversity through time are partly hampered due to the difficulty of distinguishing cycad foliage and pollen from that of the extinct (and phylogenetically distinct) clade Bennettitales [39,42]; most previous published diversity curves for Mesozoic plants do not separate Cycadales and Bennettitales, with two clades being grouped together as ‘cycadophytes’ (e.g. [34,39,112]). Also, the Cretaceous and Cenozoic cycad fossil record is probably too limited to enable understanding of the role of extinction, notably around the Cretaceous-Paleogene boundary. Although their apparent post mid-Cretaceous decline is recognized, we do not have quantitative evidence of the diversity decline nor do we know whether Cenozoic cycad diversity remained low until the Neogene radiations, or whether substantial early to mid-Cenozoic diversity existed but was affected by major extinctions. Besides illustrating the potential effect of BPP choice in Bayesian molecular dating, our study re-examines the dating of the cycads, proving a robust phylogenetic framework for future studies aimed at estimating how the diversity of cycads varied through geological time.
Conclusions
Our study highlights that the branching process prior (or tree prior) in Bayesian molecular dating has an important effect on age estimates for cycads. The birth-death process had a better fit than the traditionally used Yule process as determined by marginal likelihood estimates and Bayes factors. It is likely that cycads are not an isolated case and we advocate for a closer investigation of the branching process prior, BPP, in all future molecular phylogenetic dating studies. At minimum we suggest conducting a similar model selection as we did for cycads to select the best-fitting prior and thus the best timetree. However, the birth-death process will not necessarily always better fit the data just because it seems more biologically realistic than a simpler Yule model, especially for shallower divergence times (i.e. recent clades). For instance, two recent studies on Australian diving beetles [106] and St. John’s wort [122] did not show any age differences using different BPPs, and none of the BPP was supported with Bayes factors; both clades originated in the Oligocene (ca. 34-28 Ma). On the contrary for deeper divergence times, age differences might be revealed when comparing a birth-death and a Yule process as exemplified in a dating study of darkling beetles that originated in the Jurassic (≈180 Ma) [27], but those differences were not as marked as in cycads. Based on these studies and our results, we expect that age differences deriving from different BPPs increase when clade age is very old (e.g. Triassic and backward).
As we gain greater understanding of the priors and parameters associated with molecular dating, there have been accordingly advances in our methodology. We note that other avenues for molecular dating are being developed because the standard dating procedure results in overlaying two prior distributions for a calibration node: one from the tree prior and one from the calibration prior [19,21,102]. In the last years, there are new methods that allow the inclusion of fossils in divergence time estimation as non-contemporaneous terminal tips rather than as node calibration points, also known as tip dating [123]. Recently, Heath et al. [103] introduced the fossilized birth–death process, a model for calibrating divergence time estimates in a Bayesian framework, explicitly acknowledging that extant species and fossils are part of the same macroevolutionary process. They argued that a single model that acts as a prior on the speciation times for both calibrated and uncalibrated nodes is a better representation of the lineage diversification process. The approach seems promising and will increasingly be used in future studies, like the application on the royal ferns (family Osmundaceae) [124]. The fossilized birth–death process best fitted the Osmundaceae fossil record and also provided speciation and extinction rates associated with the dated phylogeny.
Availability of supporting data
The data set supporting the results of this article is available in the Dryad repository, http://dx.doi.org/10.5061/dryad.20k7m (see ref. [125]).
Abbreviations
- BPP:
-
Branching process prior
- BF:
-
Bayes factor
- BRC:
-
Bayesian relaxed-clock
- ESS:
-
Effective sample size
- FC:
-
Fossil calibration
- HME:
-
Harmonic mean estimate
- HPD:
-
Height posterior density
- Ma:
-
Million years ago
- MCMC:
-
Markov chain Monte Carlo
- MLE:
-
Marginal likelihood estimate
- PP:
-
Posterior probability
- PSRF:
-
Potential scale reduction factor
- SS:
-
Step**-stone
- UCLD:
-
Uncorrelated lognormal distribution clock model
References
Hedges SB, Kumar S. The Timetree of Life. Oxford: Oxford University Press; 2009.
Laenen B, Shaw B, Schneider H, Goffinet B, Paradis E, Désamoré A, et al. Extant diversity of bryophytes emerged from successive post-Mesozoic diversification bursts. Nature Comm. 2014;5:5134.
Schneider H, Schuettpelz E, Pryer KM, Cranfill R, Magallon S, Lupia R. Ferns diversified in the shadow of angiosperms. Nature. 2004;428:553–7.
Schuettpelz E, Pryer KM. Evidence for a Cenozoic radiation of ferns in an angiosperm-dominated canopy. Proc Natl Acad Sci U S A. 2009;106:11200–5.
Won H, Renner SS. Dating dispersal and radiation in the gymnosperm Gnetum (Gnetales) - Clock calibration when outgroup relationships are uncertain. Syst Biol. 2006;55:610–22.
Nagalingum NS, Marshall CR, Quental TB, Rai HS, Little DP, Mathews S. Recent synchronous radiation of a living fossil. Science. 2011;334:796–9.
Mao K, Milne RI, Zhang L, Peng Y, Liu J, Thomas P, et al. Distribution of living Cupressaceae reflects the breakup of Pangea. Proc Natl Acad Sci U S A. 2012;109:7793–8.
Leslie AB, Beaulieu JM, Rai HS, Crane PR, Donoghue MJ, Mathews S. Hemisphere-scale differences in conifer evolutionary dynamics. Proc Natl Acad Sci U S A. 2012;109:16217–21.
Magallón SA, Castillo A. Angiosperm diversification through time. Am J Bot. 2009;96:349–65.
Fiz-Palacios O, Schneider H, Heinrichs J, Savolainen V. Diversification of land plants: insights from a family-level phylogenetic analysis. BMC Evol Biol. 2011;11:341.
Zanne AE, Tank DC, Cornwell WK, Eastman JM, Smith SA, FitzJohn RG, et al. Three keys to the radiation of angiosperms into freezing environments. Nature. 2014;506:89–92.
Percy DM, Page RDM, Cronk QCB. Plant-insect interactions: double-dating associated insect and plant lineages reveals asynchronous radiations. Syst Biol. 2004;53:120–7.
Kumar S. Molecular clocks: four decades of evolution. Nature Rev Genet. 2005;6:654–62.
Donoghue PCJ, Benton MJ. Rocks and clocks: calibrating the Tree of Life using fossils and molecules. Trends Ecol Evol. 2007;22:424–31.
Alfaro ME, Holder MT. The posterior and the prior in Bayesian phylogenetics. Annu Rev Ecol Evol Syst. 2006;37:19–42.
Drummond AJ, Ho SYW, Phillips MJ, Rambaut A. Relaxed phylogenetics and dating with confidence. PLoS Biol. 2006;4:e88.
Inoue J, Donoghue PCJ, Yang Z. The impact of the representation of fossil calibrations on Bayesian estimation of species divergence times. Syst Biol. 2010;59:74–89.
Yang Z, Rannala B. Bayesian estimation of species divergence times under a molecular clock using fossil calibrations with soft bounds. Mol Biol Evol. 2006;23:212–26.
Ho SYW, Phillips MJ. Accounting for calibration uncertainty in phylogenetic estimation of evolutionary divergence times. Syst Biol. 2009;58:367–80.
Yang Z, Rannala B. Branch-length prior influences Bayesian posterior probability of phylogeny. Syst Biol. 2005;54:455–70.
Heled J, Drummond AJ. Calibrated birth-death phylogenetic time-tree priors for Bayesian inference. Syst Biol. 2015;64:369–383.
Morlon H. Phylogenetic approaches for studying diversification. Ecol Lett. 2014;17:508–25.
Yang Z, Goldman N, Friday A. Maximum likelihood tree from DNA sequences: a peculiar statistical estimation problem. Syst Biol. 1995;44:384–99.
Rannala B, Yang Z. Probability distribution of molecular evolutionary trees: a new method of phylogenetic inference. J Mol Evol. 1996;43:304–11.
Gernhard T. The conditioned reconstructed process. J Theoret Biol. 2008;253:769–78.
Drummond AJ, Suchard MA, **e D, Rambaut A. Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol Biol Evol. 2012;29:1969–73.
Kergoat GJ, Bouchard P, Clamens A-L, Abbate JL, Jourdan H, Zahab R, et al. Cretaceous environmental changes led to high turnovers and decreases in diversification rates in a hyperdiverse non-phytophagous beetle family. BMC Evol Biol. 2014;14:220.
Couvreur TLP, Franzke A, Al-Shehbaz IA, Bakker F, Koch M, Mummenhoff K. Molecular phylogenetics, temporal diversification and principles of evolution in the mustard family (Brassicaceae). Mol Biol Evol. 2010;27:55–71.
Osborne R, Calonje MA, Hill KD, Stanberg L, Stevenson DW. The world list of Cycads. Mem New York Botl Gard. 2012;106:480–510.
Pant DD. The fossil history and phylogeny of the cycadales. Geophytol. 1987;17:125–62.
Axsmith BJ, Serbet R, Krings M, Taylor TN, Taylor EL, Mamay SH. The enigmatic Paleozoic plants Spermopteris and Phasmatocycas reconsidered. Am J Bot. 2003;90:1585–95.
Hermsen EJ, Taylor TN, Taylor EL, Stevenson DW. Cataphylls of the Middle Triassic cycad Antarcticycas schopfii and new insights into cycad evolution. Am J Bot. 2006;93:724–38.
Cúneo NR, Escapa I, Villar-de-Seoane L, Artabe A, Gnaedinger S. Review of the Cycads and Bennettitaleans from the Mesozoic of Argentina. In: Gee CT, editor. Plants in Mesozoic Time: Morphological Innovations, Phylogeny, Ecosystems. Bloomington: Indiana University Press; 2010. p. 187–212.
Crane PR. Vegetational consequences of the angiosperm diversification. In: Friis EM, Chaloner WC, Crane PR, editors. The Origin of Angiosperms and their Biological Consequences. Cambridge: Cambridge University Press; 1987. p. 107–44.
Lupia R, Lidgard S, Crane PR. Comparing palynological abundance and diversity: implications for biotic replacement during the Cretaceous angiosperm radiation. Paleobiol. 1999;25:305–40.
van de Schootbrugge B, Quan TM, Lindström S, Püttmann W, Heunisch C, Pross J, et al. Floral changes across the Triassic/Jurassic boundary linked to flood basalt volcanism. Nature Geosci. 2009;2:589–94.
Coiffard C, Gomez B, Daviero-Gomez V, Dilcher DL. Rise to dominance of angiosperm pioneers in European Cretaceous environments. Proc Natl Acad Sci U S A. 2012;109:20955–9.
Crisp MD, Cook LG. Cenozoic extinctions account for the low diversity of extant gymnosperms compared with angiosperms. New Phytol. 2011;192:997–1009.
Willis KJ, McElwain JC. The Evolution of Plants. Oxford: Oxford University Press; 2002.
McElwain JC, Punyasena SW. Mass extinction events and the fossil record. Trends Ecol Evol. 2007;22:548–57.
Butler RJ, Barrett PM, Kenrick P, Penn MG. Testing co-evolutionary hypotheses over geological timescales: interactions between Mesozoic non-avian dinosaurs and cycads. Biological Rev. 2009;84:73–89.
Taylor TN, Taylor EL, Krings M. Paleobotany: The Biology and Evolution of Fossil Plants. 2nd ed. Burlington: Academic Press; 2009.
Salas-Leiva DE, Meerow AW, Calonje M, Griffith MP, Francisco-Ortega J, Nakamura K, et al. Phylogeny of the cycads based on multiple single-copy nuclear genes: congruence of concatenated parsimony, likelihood and species tree inference methods. Ann Bot. 2013;112:1263–78.
Hollick A. Descriptions of new species of Tertiary cycads, with a review of those previously recorded. Bull Torrey Bot Club. 1932;59:169–89.
Hill RS. Three new Eocene cycads from eastern Australia. Austral J Bot. 1980;28:105–22.
Horiuchi J, Kimura T. Dioonopsis nipponica gen. et sp. nov., a new cycad from the Palaeogene of Japan. Rev Palaeobot Palynol. 1987;51:213–25.
Kvaček Z. A new Tertiary Ceratozamia (Zamiaceae, Cycadopsida) from the European Oligocene. Flora – Morphology, Distribution Functional Ecology of Plants. 2002;197:303–16.
Erdei B, Manchester SR, Kvaček Z. Dioonopsis Horiuchi et Kimura leaves from the Eocene of Western North America: a cycad shared with the Paleogene of Japan. Intl J Plant Sci. 2012;173:81–95.
Kvaček Z. New fossil records of Ceratozamia (Zamiaceae, Cycadales) from the European Oligocene and lower Miocene. Acta Palaeobot. 2014;54:231–47.
Su K, Quan C, Liu Y-S. Cycas fushunensis sp. nov. (Cycadaceae) from the Eocene of northeast China. Rev Palaeobot Palynol. 2014;204:43–9.
Griffith MP, Magellan TM, Tomlinson PB. Variation in leaflet structure in Cycas (Cycadales: Cycadaceae): Does anatomy follow phylogeny and geography? Intl J Plant Sci. 2014;175:241–55.
González D, Vovides AP. Low intralineage divergence in Ceratozamia (Zamiaceae) detected with nuclear ribosomal DNA ITS and chloroplast DNA trnL-F non-coding region. Syst Bot. 2002;27:654–61.
Hill KD, Chase MW, Stevenson DW, Hill HG, Schutzman B. The families and genera of cycads: a molecular phylogenetic analysis of Cycadophyta based on nuclear and plastid DNA sequences. Intl J Plant Sci. 2003;164:933–48.
Rai HS, O’Brien HE, Reeves PA, Olmstead RG, Graham SW. Inference of higher-order relationships in the cycads from a large chloroplast data set. Mol Phylogenet Evol. 2003;29:350–9.
Chaw S-M, Walters TW, Chang C-C, Hu S-H, Chen S-H. A phylogeny of cycads (Cycadales) inferred from chloroplast matK gene, trnK intron, and nuclear rDNA ITS region. Mol Phylogenet Evol. 2005;37:214–34.
González D, Vovides AP, Barcenas C. Phylogenetic relationships of the Neotropical genus Dioon (Cycadales, Zamiaceae) based on nuclear and chloroplast DNA sequence data. Syst Bot. 2008;33:229–36.
Yessoufou K, Bamigboye SO, Daru BH, van der Bank M. Evidence of constant diversification punctuated by a mass extinction in the African cycads. Ecol Evol. 2013;4:50–8.
Wu CH, Chaw SM, Huang YY. Chloroplast phylogenomics indicates that Ginkgo biloba is sister to cycads. Genome Biol Evol. 2013;5:243–54.
Martínez LCA, Artabe AEE, Bodnar J. A new cycad stem from the Cretaceous in Argentina and its phylogenetic relationships with other Cycadales. Bot J Linn Soc. 2012;170:436–58.
Sauquet H, Ho SYW, Gandolfo MA, Jordan GJ, Wilf P, Cantrill DJ, et al. Testing the impact of calibration on molecular divergence times using a fossil-rich group: the case of Nothofagus (Fagales). Syst Biol. 2012;61:289–313.
Near TJ, Meylan PA, Shaffer HB. Assessing concordance of fossil calibration points in molecular clock studies: an example using turtles. Am Nat. 2005;165:137–46.
Clarke JT, Warnock RCM, Donoghue PCJ. Establishing a time-scale for plant evolution. New Phytol. 2011;192:266–301.
Gradstein FM, Ogg JG, Schmitz MD, Ogg GM. The Geologic Time Scale 2012, Volume Set. Boston, USA: Elsevier; 2012.
Zhu JN, Zhang SS, Ma J. A new genus and species – Cycadostrobus paleozoicus Zhu of Cycadaceae from the Permian of China. Acta Phytotaxo Sinica. 1994;32:340–4.
Zhu JN, Du XM. A new cycad – Primocycas chinensis gen. et sp. nov. discovers from the Lower Permian in Sha-nxi, China and its significance. Acta Bot Sinica. 1981;23:401–4.
Zhifeng G, Thomas BA. A review of fossil cycad megasporophylls, with new evidence of Crossozamia Pomel and its associated leaves from the Lower Permian of Taiyuan, China. Rev Palaeobot Palynol. 1989;60:205–23.
Wang J. Late Paleozoic macrofloral assemblages from Weibei Coalfield, with reference to vegetational change through the Late Paleozoic Ice-age in the North China Block. Intl J Coal Geol. 2010;83:292–317.
Hill RS. Two new species of Bowenia Hook, ex Hook, f. from the Eocene of eastern Australia. Austral J Bot. 1978;26:837–46.
Smoot EL, Taylor TN, Delevoryas T. Structurally preserved fossil plants from Antarctica. I. Antarcticycas, gen. nov., a Triassic cycad stem from the Beardmore Glacier area. Am J Bot. 1985;72:1410–23.
Hermsen EJ, Taylor EL, Taylor TN. Morphology and ecology of the Antarcticycas plant. Rev Palaeobot Palynol. 2009;153:108–23.
Rothwell GW, Scheckler SE, Gillespie WH. Elkinsia gen nov, a Late Devonian gymnospermn with cupulate ovules. Bot Gazette. 1989;150:170–89.
Ronquist F, Teslenko M, Van der Mark P, Ayres DL, Darling A, Höhna S, et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol. 2012;61:539–42.
Lanfear R, Calcott B, Ho SYW, Guindon S. PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol Biol Evol. 2012;29:1695–701.
Huelsenbeck JP, Larget B, Alfaro ME. Bayesian phylogenetic model selection using reversible jump Markov chain Monte Carlo. Mol Biol Evol. 2004;21:1123–33.
Rambaut A, Suchard MA, **e D, Drummond AJ. Tracer v1.6. Available at: http://beast.bio.ed.ac.uk/Tracer; 2015
Baele G, Li WLS, Drummond AJ, Suchard MA, Lemey P. Accurate model selection of relaxed molecular clocks in Bayesian phylogenetics. Mol Biol Evol. 2013;30:239–43.
Vanneste K, Baele G, Maere S, Van de Peer Y. Analysis of 41 plant genomes supports a wave of successful genome duplications in association with the Cretaceous–Paleogene boundary. Genome Res. 2014;24:1334–47.
Warnock RCM, Parham JF, Joyce WG, Lyson TR, Donoghue PCJ. Calibration uncertainty in molecular dating analyses: there is no substitute for the prior evaluation of time priors. Proc R Soc B. 2015;282:20141013.
Near TJ, Sanderson MJ. Assessing the quality of molecular divergence time estimates by fossil calibrations and fossil-based model selection. Philos T R Soc B. 2004;359:1477–83.
Ayres DL, Darling A, Zwickl DJ, Beerli P, Holder MT, Lewis PO, et al. BEAGLE: an application programming interface and high-performance computing library for statistical phylogenetics. Syst Biol. 2012;61:170–3.
Miller MA, Schwartz T, Pickett BE, He S, Klem EB, Scheuermann RH, et al. A RESTful API for Access to Phylogenetic Tools via the CIPRES Science Gateway. Evol Bioinformatics. 2015;11:43–8.
Lartillot N, Philippe H. Computing Bayes factors using thermodynamic integration. Syst Biol. 2006;55:195–207.
**e W, Lewis PO, Fan Y, Kuo L, Chen MH. Improving marginal likelihood estimation for Bayesian phylogenetic model selection. Syst Biol. 2011;60:150–60.
Kass RE, Raftery AE. Bayes factors. J Am Stat Assoc. 1995;90:773–95.
Rabosky DL, Santini F, Eastman JM, Smith SA, Sidlauskas B, Chang J, et al. Rates of speciation and morphological evolution are correlated across the largest vertebrate radiation. Nature Comm. 2013;4:1958.
Rabosky DL, Donnellan SC, Grundler M, Lovette IJ. Analysis and visualization of complex macroevolutionary dynamics: an example from Australian scincid lizards. Syst Biol. 2014;63:610–27.
Rabosky DL. Automatic detection of key innovations, rate shifts, and diversity dependence on phylogenetic trees. PLoS One. 2014;9:e89543.
Paradis E. Can extinction rates be estimated without fossils? J Theor Biol. 2004;229:19–30.
Quental TB, Marshall CR. Diversity dynamics: molecular phylogenies need the fossil record. Trends Ecol Evol. 2010;25:434–41.
Rabosky DL. Extinction rates should not be estimated from molecular phylogenies. Evolution. 2010;64:1816–24.
Nee S, Holmes EC, May RM, Harvey PH. Extinction rates can be estimated from molecular phylogenies. Philos Trans R Soc B. 1994;344:77–82.
Purvis A. Phylogenetic approaches to the study of extinction. Annu Rev Ecol Evol Syst. 2008;39:301–19.
Morlon H, Parsons TL, Plotkin JB. Reconciling molecular phylogenies with the fossil record. Proc Natl Acad Sci U S A. 2011;108:16327–32.
Beaulieu JM, O’Meara BC. Extinction can be estimated from moderately sized molecular phylogenies. Evolution. 2015, doi.org/10.1111/evo.12614.
Huang S, Chiang YC, Schaal BA, Chou CH, Chiang TY. Organelle DNA phylogeography of Cycas taitungensis, a relict species in Taiwan. Mol Ecol. 2001;10:2669–81.
**ao LQ, Möller M. Nuclear ribosomal its functional paralogs resolve the phylogenetic relationships of a late-Miocene radiation cycad Cycas (Cycadaceae). PLoS One. 2015;10:e0117971.
Treutlein J, Vorster P, Wink M. Molecular relationships in Encephalartos (Zamiaceae, Cycadales) based on nucleotide sequences of nuclear ITS 1&2, rbcL, and genomic ISSR fingerprinting. Plant Biol. 2005;7:79–90.
Sharma IK, Jones DL, Forster PI. Genetic differentiation and phenetic relatedness among seven species of the Macrozamia plurinervia complex (Zamiaceae). Biochem Syst Ecol. 2004;32:313–27.
Caputo P, Cozzolino S, Gaudio L, Moretti A, Stevenson D. Karyology and phylogeny of some mesoamerican species of Zamia (Zamiaceae). Am J Bot. 1996;83:1513–20.
Zgurski JM, Rai HS, Fai QM, Bogler DJ, Francisco-Ortega J, Graham SW. How well do we understand the overall backbone of cycad phylogeny? New insights from a large, multigene plastid data set. Mol Phylogenet Evol. 2008;47:1232–7.
Raup DM. Extinction: Bad genes or bad luck? New York: W.W. Norton and Company; 1991.
Heled J, Drummond AJ. Calibrated tree priors for relaxed phylogenetics and divergence time estimation. Syst Biol. 2012;61:138–49.
Heath TA, Huelsenbeck JP, Stadler T. The fossilized birth-death process for coherent calibration of divergence-time estimates. Proc Natl Acad Sci U S A. 2014;111:E2957–66.
Hagen O, Hartmann K, Steel M, Stadler T. Age-dependent speciation can explain the shape of empirical phylogenies. Syst Biol. 2015;64:432–440.
Drummond AJ, Bouckaert RR. Bayesian evolutionary analysis with BEAST. Cambridge: Cambridge University Press; 2015.
Toussaint EFA, Condamine FL, Hawlitschek O, Watts CHS, Porch N, Hendrich L, et al. Unveiling the diversification dynamics of Australasian predaceous diving beetles in the Cenozoic. Syst Biol. 2015;64:3–24.
Antonelli A, Sanmartín I. Mass extinction, gradual cooling, or rapid radiation? Reconstructing the spatiotemporal evolution of the ancient angiosperm genus Hedyosmum (Chloranthaceae) using empirical and simulated approaches. Syst Biol. 2011;60:596–615.
Thomas N, Bruhl JJ, Ford A, Weston PH. Molecular dating of Winteraceae reveals a complex biogeographical history involving both ancient Gondwanan vicariance and long-distance dispersal. J Biogeogr. 2014;41:894–904.
Burleigh JG, Brabazuk WD, Davis JM, Morse AM, Soltis PS. Exploring diversification and genome size evolution in extant gymnosperms through phylogenetic synthesis. J Bot. 2012;2012:1–6.
Peñalver E, Labandeira CC, Barrón E, Delclòs X, Nel P, Nel A, et al. Thrips pollination of Mesozoic gymnosperms. Proc Natl Acad Sci U S A. 2012;109:8623–8.
Rainford JL, Hofreiter M, Nicholson DB, Mayhew PJ. Phylogenetic distribution of extant richness suggests metamorphosis is a key innovation driving diversification in insects. PLoS One. 2014;9:e109085.
Niklas KJ, Tiffney BH, Knoll AH. Patterns in vascular land plant diversification. Nature. 1983;303:614–6.
Silvestro D, Cascales-Miñana B, Bacon CD, Antonelli A. Revisiting the origin and diversification of vascular plants through a comprehensive Bayesian analysis of the fossil record. New Phytol. in press, doi.org/10.1111/nph.13247.
Artabe A, Archangelsky S. Las Cycadales Mesodescolea plicata Archangelsky emend. Archangelsky y Petriella 1971 (Cretacio) y Stangeria Moore (Actual). Estudio comparativo de epidermis foliar con microscopia electr6nica de barrido y transmisi6n. Ameghiniana. 1992;28:115–23.
Artabe A, Stevenson DW. Fossil Cycadales of Argentina. Botanical Rev. 1999;65:219–38.
Passalia MG, Del Fueyo G, Archangelsky S. An Early Cretaceous zamiaceous cycad of South West Gondwana: Restrepophyllum nov. gen. from Patagonia, Argentina. Rev Palaeobot Palynol. 2010;161:137–50.
Crisp MD, Cook LG. Explosive radiation or cryptic mass extinction? Interpreting signatures in molecular phylogenies. Evolution. 2009;63:2257–65.
Uzunova K, Palamarev E, Kvaček Z. Eostangeria ruzinciniana (Zamiaceae) from the Middle Miocene of Bulgaria and its relationship to similar taxa of fossil Eostangeria, and extant Chigua and Stangeria (Cycadales). Acta Palaeobot. 2001;41:177–93.
Nagalingum NS, Drinnan AN, Lupia R, McLoughlin S. Fern spore diversity and abundance in Australia during the Cretaceous. Rev Palaeobot Palynol. 2002;119:69–92.
Tang W. The evolutionary history of North American cycads. Cycad Newslett. 2012;35:7–13.
Cramer BS, Miller KG, Barrett PJ, Wright JD. Late Cretaceous–Neogene trends in deep ocean temperature and continental ice volume: Reconciling records of benthic foraminiferal geochemistry (δ18O and Mg/Ca) with sea level history. J Geophys Res: Oceans. 2011;116:1978–2012.
Meseguer AS, Lobo JM, Ree R, Beerling DJ, Sanmartín I. Integrating fossils, phylogenies, and niche models into biogeography to reveal ancient evolutionary history: the case of Hypericum (Hypericaceae). Syst Biol. 2015;64:215–32.
Ronquist F, Klopfstein S, Vilhelmsen L, Schulmeister S, Murray DL, Rasnitsyn AP. A total-evidence approach to dating with fossils, applied to the early radiation of the Hymenoptera. Syst Biol. 2012;61:973–99.
Grimm GW, Kapli P, Bomfleur B, McLoughlin S, Renner SS. Using more than the oldest fossils: Dating Osmundaceae with three Bayesian clock approaches. Syst Biol. 2015;64:96–405.
Condamine FL, Nagalingum NS, Marshall CR, Morlon H. Data from: Origin and diversification of living cycads: A cautionary tale on the impact of the branching process prior in Bayesian molecular dating. BMC Evol Biol. 2015, http://dx.doi.org/10.5061/dryad.20k7m.
Acknowledgements
We thank the Associate Editor Rafael Zardoya, and the two referees Susanne Renner and Diego San Mauro who provided insightful comments on the study. We also thank Frédéric Delsuc and Gael Kergoat for discussions, and Simon Ho and Guy Baele for technical comments on the Bayesian analyses. We also thank Daniel Rabosky for the workshop on the BAMM approach, and Mario Coiro for discussions and feedbacks on the fossil record of cycads. Funding was provided by grant ANR CHEX ECOEVOBIO awarded to HM.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
FLC and HM designed the research. FLC retrieved material or data from GenBank, formerly published by NSN and CRM. FLC performed the analyses. FLC wrote the manuscript, with revisions by HM, NSN and CRM. All authors read and approved the final manuscript.
Authors’ information
Fabien Condamine is a post-doc student who is interested in numerous biodiversity patterns such as the latitudinal diversity gradient, regional difference in diversity and island biogeography. He aims to decipher the main evolutionary and ecological processes that have shaped the present pattern of biodiversity. Nathalie Nagalingum is a research scientist who is interested in cycad diversity, its origin, evolution and conservation using genomic and paleontological tools. Charles Marshall is a professor who is interested in how paleontology can inform our understanding of the history of life, and the processes that control it. Hélène Morlon combines mathematics, bioinformatics and fieldwork to study questions ranging from macroevolution and macroecology to community assembly, biogeography, and conservation.
Additional files
Additional file 1: Table S1.
Taxa sampled and GenBank accession numbers of sequences used in the analyses. All PHYP sequences were generated by Nagalingum et al. [6], and sequences from matK and rbcL were downloaded from GenBank. “—” denotes sequence not available or voucher not indicated.
Additional file 2: Figure S1.
Phylogeny of Cycadales reconstructed (for 237 cycad species and three genes) using MrBayes and reversible jump MCMC. Posterior probabilities are shown at nodes. Only the nodes in the backbone are considered well-supported here. Nodes within generic radiations are generally below the standard threshold of robustness. The low node support within genera is likely due to low genetic divergence between species.
Additional file 3: Figure S2.
Convergence of the BAMM analysis with the chronogram reconstructed with a Yule prior. (A) The stationary of the MCMC before applying a burn-in. (B) The posterior distribution of number of shifts estimated before applying a burn-in. (C) The stationary of the MCMC after removing the burn-in phase. (D) The posterior distribution of number of shifts estimated after removing the burn-in phase.
Additional file 4: Figure S3.
Convergence of the BAMM analysis with the chronogram reconstructed with a birth-death prior. (A) The stationary of the MCMC before applying a burn-in. (B) The posterior distribution of number of shifts estimated before applying a burn-in. (C) The stationary of the MCMC after removing the burn-in phase. (D) The posterior distribution of number of shifts estimated after removing the burn-in phase.
Additional file 5: Figure S4.
Frequency distribution of distinct macroevolutionary rate regimes estimated using BAMM and the tree reconstructed with the Yule prior. (A) Prior distribution of the number of distinct processes. (B) Posterior distribution of the number of distinct processes (including the root process) on the cycad phylogeny reconstructed with a Yule model. A one-process model outperforms a two-process model.
Additional file 6: Figure S5.
Frequency distribution of distinct macroevolutionary rate regimes estimated using BAMM and the tree reconstructed with the birth-death prior. (A) Prior distribution of the number of distinct processes. (B) Posterior distribution of the number of distinct processes (including the root process) on the cycad phylogeny reconstructed with a birth-death model. A one-process model outperforms a two-process model.
Additional file 7: Figure S6.
Credible set of configuration shifts of cycads inferred with BAMM using the (A) tree with the Yule prior, and the (B) tree dated with the birth-death prior. Phylogenies show the distinct shift configurations with the highest posterior probability. For each shift configuration, the locations of rate shifts are shown as red (rate increases) and blue (rate decreases) circles, with circle size proportional to the marginal probability of the shift. Text labels (e.g. f =0.33) denote the posterior probability of each shift configuration.
Additional file 8: Figure S7.
The best shift configuration inferred with BAMM using the (A) tree with the Yule prior, and the (B) tree dated with the birth-death prior. Analysis with the Yule prior indicated a single rate shift (indicated by red circle) near the crowns of Cycas and Encephalartos, whereas the analysis with the birth-death prior indicated four major shifts at the richest genera: Cycas, Zamia, Encephalartos, and Macrozamia.
Additional file 9: Figure S8.
Macroevolutionary cohort matrix for speciation of cycads using the birth-death prior. Each cell in the matrix is coded by a color denoting the pairwise probability that two species share a common macroevolutionary rate regime. The cycad phylogeny reconstructed with a birth-death process is shown for reference on the left and upper margins of each cohort matrix. At least five major cohorts can be identified that are the four most species-rich genera: Cycas, Zamia, Encephalartos, and Macrozamia.
Additional file 10: Figure S9.
Macroevolutionary cohort matrix for speciation of cycads using the Yule prior. Same legend as Additional file 9: Figure S8.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Condamine, F.L., Nagalingum, N.S., Marshall, C.R. et al. Origin and diversification of living cycads: a cautionary tale on the impact of the branching process prior in Bayesian molecular dating. BMC Evol Biol 15, 65 (2015). https://doi.org/10.1186/s12862-015-0347-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12862-015-0347-8