Abstract
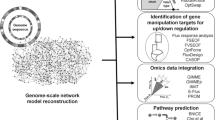
Escherichia coli is a model organism with a clear genetic background that is widely used in metabolic engineering and synthetic biology research. To gain a complete picture of the complexly metabolic and regulatory interactions in E. coli, researchers often need to retrieve information from various databases which cover different types of interactions. A central one-stop service integrating various molecular interactions in E. coli would be helpful for the community. We constructed a database called E. coli integrated network (EcoIN) by integrating known molecular interaction information from databases and literature. EcoIN contains nearly 160,000 pairs of interactions and users can easily search the different types of interacting partners for a metabolite, gene or protein, and thus gain access to a more comprehensive interaction map of E. coli. To illustrate the application of EcoIN, we used the full path algorithm to identify metabolic feedback/feedforward regulatory loops having at least two different types of regulatory interactions. Applying this algorithm to analyze the regulatory loops for the amino acid biosynthetic pathways, we found some multi-step regulation loops which may affect the metabolic flux and are potential new engineering targets. The EcoIN database is freely accessible at http://ecoin.ibiodesign.net/ and analysis codes are available at GitHub: https://github.com/maozhitao/EcoIN.




Similar content being viewed by others
Data availability
The datasets supporting the conclusions of this article are included at GitHub: https://github.com/maozhitao/EcoIN.
Code availability
All analysis codes supporting the conclusions of this article are included at GitHub: https://github.com/maozhitao/EcoIN.
References
Baumler DJ, Peplinski RG, Reed JL, Glasner JD, Perna NT. The evolution of metabolic networks of E. coli. BMC Syst Biol. 2011;5:182.
Zhou X, Wu H, Li Z, Zhou X, Bai L, Deng Z. Over-expression of UDP-glucose pyrophosphorylase increases validamycin A but decreases validoxylamine A production in Streptomyces hygroscopicus var. **ggangensis 5008. Metab Eng. 2011;13(6):768–76.
Dhamankar H, Tarasova Y, Martin CH, Prather KL. Engineering E. coli for the biosynthesis of 3-hydroxy-gamma-butyrolactone (3HBL) and 3,4-dihydroxybutyric acid (3,4-DHBA) as value-added chemicals from glucose as a sole carbon source. Metab Eng. 2014;25:72–81.
Curran KA, Leavitt JM, Karim AS, Alper HS. Metabolic engineering of muconic acid production in Saccharomyces cerevisiae. Metab Eng. 2013;15:55–66.
Covert MW, Schilling CH, Palsson B. Regulation of gene expression in flux balance models of metabolism. J Theor Biol. 2001;213(1):73–88.
Shlomi T, Eisenberg Y, Sharan R, Ruppin E. A genome-scale computational study of the interplay between transcriptional regulation and metabolism. Mol Syst Biol. 2007;3:101.
Chandrasekaran S, Price ND. Probabilistic integrative modeling of genome-scale metabolic and regulatory networks in Escherichia coli and Mycobacterium tuberculosis. PNAS. 2010;107(41):17845–50.
Carrera J, Estrela R, Luo J, Rai N, Tsoukalas A, Tagkopoulos I. An integrative, multi-scale, genome-wide model reveals the phenotypic landscape of Escherichia coli. Mol Syst Biol. 2014;10:735.
Covert MW, **ao N, Chen TJ, Karr JR. Integrating metabolic, transcriptional regulatory and signal transduction models in Escherichia coli. Bioinformatics. 2008;24(18):2044–50.
Lee JM, Gianchandani EP, Eddy JA, Papin JA. Dynamic analysis of integrated signaling, metabolic, and regulatory networks. PLoS Comput Biol. 2008;4(5):e1000086.
Orth JD, Thiele I, Palsson BO. What is flux balance analysis? Nat Biotechnol. 2010;28(3):245–8.
Kuhn M, Szklarczyk D, Pletscher-Frankild S, Blicher TH, von Mering C, Jensen LJ, Bork P. STITCH 4: integration of protein-chemical interactions with user data. Nucleic Acids Res. 2014;42:D401-407.
Whitehead E. The regulation of enzyme activity and allosteric transition. Prog Biophys Mol Biol. 1970;21:321–97.
Latchman DS. Transcription factors: an overview. Int J Biochem Cell Biol. 1997;29(12):1305–12.
Missiakas D, Raina S. The extracytoplasmic function sigma factors: role and regulation. Mol Microbiol. 1998;28(6):1059–66.
Na D, Yoo SM, Chung H, Park H, Park JH, Lee SY. Metabolic engineering of Escherichia coli using synthetic small regulatory RNAs. Nat Biotechnol. 2013;31(2):170–4.
Yanofsky C. Attenuation in the control of expression of bacterial operons. Nature. 1981;289(5800):751–8.
Wolfe AJ. Bacterial protein acetylation: new discoveries unanswered questions. Curr Genet. 2015;62:335–41.
Hu LI, Lima BP, Wolfe AJ. Bacterial protein acetylation: the dawning of a new age. Mol Microbiol. 2010;77(1):15–21.
Su C, Peregrin-Alvarez JM, Butland G, Phanse S, Fong V, Emili A, Parkinson J. Bacteriome.org—an integrated protein interaction database for E. coli. Nucleic Acids Res. 2007;36(suppl_1):D632–6.
Kim H, Shim JE, Shin J, Lee I. EcoliNet: a database of cofunctional gene network for Escherichia coli. Database J Biol Databases Curation. 2015;2015:1–8.
Szklarczyk D, Gable AL, Lyon D, Junge A, Wyder S, Huerta-Cepas J, Simonovic M, Doncheva NT, Morris JH, Bork P, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47(D1):D607–13.
Gumerov VM, Ortega DR, Adebali O, Ulrich LE, Zhulin IB. MiST 3.0: an updated microbial signal transduction database with an emphasis on chemosensory systems. Nucleic Acids Res. 2020;48(D1):D459–64.
Santos-Zavaleta A, Salgado H, Gama-Castro S, Sanchez-Perez M, Gomez-Romero L, Ledezma-Tejeida D, Garcia-Sotelo JS, Alquicira-Hernandez K, Muniz-Rascado LJ, Pena-Loredo P, et al. RegulonDB v 10.5: tackling challenges to unify classic and high throughput knowledge of gene regulation in E. coli K-12. Nucleic Acids Res. 2019;47(D1):D212–20.
Chang A, Jeske L, Ulbrich S, Hofmann J, Koblitz J, Schomburg I, Neumann-Schaal M, Jahn D, Schomburg D. BRENDA, the ELIXIR core data resource in 2021: new developments and updates. Nucleic Acids Res. 2021;49(D1):D498–508.
Keseler IM, Mackie A, Santos-Zavaleta A, Billington R, Bonavides-Martínez C, Caspi R, Fulcher C, Gama-Castro S, Kothari A, Krummenacker M, et al. The EcoCyc database: reflecting new knowledge about Escherichia coli K-12. Nucleic Acids Res. 2016;45(D1):D543–50.
Szklarczyk D, Santos A, von Mering C, Jensen LJ, Bork P, Kuhn M. STITCH 5: augmenting protein-chemical interaction networks with tissue and affinity data. Nucleic Acids Res. 2016;44(D1):D380-384.
AbouElfetouh A, Kuhn ML, Hu LI, Scholle MD, Sorensen DJ, Sahu AK, Becher D, Antelmann H, Mrksich M, Anderson WF, et al. The E. coli sirtuin CobB shows no preference for enzymatic and nonenzymatic lysine acetylation substrate sites. MicrobiologyOpen. 2015;4(1):66–83.
Colak G, **e ZY, Zhu AY, Dai LZ, Lu ZK, Zhang Y, Wan XL, Chen Y, Cha YH, Lin HN, et al. Identification of lysine succinylation substrates and the succinylation regulatory enzyme CobB in Escherichia coli. Mol Cell Proteomics. 2013;12(12):3509–20.
Kuhn ML, Zemaitaitis B, Hu LI, Sahu A, Sorensen D, Minasov G, Lima BP, Scholle M, Mrksich M, Anderson WF, et al. Structural, kinetic and proteomic characterization of acetyl phosphate-dependent bacterial protein acetylation. PLoS One. 2014;9(4):e94816.
Schilling B, Christensen D, Davis R, Sahu AK, Hu LI, Walker-Peddakotla A, Sorensen DJ, Zemaitaitis B, Gibson BW, Wolfe AJ. Protein acetylation dynamics in response to carbon overflow in Escherichia coli. Mol Microbiol. 2015;98(5):847–63.
Schmidt A, Kochanowski K, Vedelaar S, Ahrne E, Volkmer B, Callipo L, Knoops K, Bauer M, Aebersold R, Heinemann M. The quantitative and condition-dependent Escherichia coli proteome. Nat Biotechnol. 2016;34(1):104–10.
Weinert BT, Iesmantavicius V, Wagner SA, Scholz C, Gummesson B, Beli P, Nystrom T, Choudhary C. Acetyl-phosphate is a critical determinant of lysine acetylation in E. coli. Mol Cell. 2013;51(2):265–72.
Zhang K, Zheng S, Yang JS, Chen Y, Cheng Z. Comprehensive profiling of protein lysine acetylation in Escherichia coli. J Proteome Res. 2013;12(2):844–51.
Soares NC, Spat P, Krug K, Macek B. Global dynamics of the Escherichia coli proteome and phosphoproteome during growth in minimal medium. J Proteome Res. 2013;12(6):2611–21.
Potel CM, Lin M-H, Heck AJR, Lemeer S. Widespread bacterial protein histidine phosphorylation revealed by mass spectrometry-based proteomics. Nat Methods. 2018;15:187.
Orth JD, Conrad TM, Na J, Lerman JA, Nam H, Feist AM, Palsson BO. A comprehensive genome-scale reconstruction of Escherichia coli metabolism–2011. Mol Syst Biol. 2011;7:535.
Ortet P, Whitworth DE, Santaella C, Achouak W, Barakat M. P2CS: updates of the prokaryotic two-component systems database. Nucleic Acids Res. 2015;43:D536-541.
Kim M, Rai N, Zorraquino V, Tagkopoulos I. Multi-omics integration accurately predicts cellular state in unexplored conditions for Escherichia coli. Nat Commun. 2016;7:13090.
Hu P, Janga SC, Babu M, Diaz-Mejia JJ, Butland G, Yang W, Pogoutse O, Guo X, Phanse S, Wong P, et al. Global functional atlas of Escherichia coli encompassing previously uncharacterized proteins. PLoS Biol. 2009;7(4):e96.
Ma HW, Zeng AP. Reconstruction of metabolic networks from genome data and analysis of their global structure for various organisms. Bioinformatics. 2003;19(2):270–7.
Lopez-Ibanez J, Pazos F, Chagoyen M. MBROLE 2.0-functional enrichment of chemical compounds. Nucleic Acids Res. 2016;44(W1):W201-204.
Hastings J, Owen G, Dekker A, Ennis M, Kale N, Muthukrishnan V, Turner S, Swainston N, Mendes P, Steinbeck C. ChEBI in 2016: improved services and an expanding collection of metabolites. Nucleic Acids Res. 2016;44(D1):D1214-1219.
Kanehisa M, Furumichi M, Sato Y, Ishiguro-Watanabe M, Tanabe M. KEGG: integrating viruses and cellular organisms. Nucleic Acids Res. 2021;49(D1):D545–51.
Wishart DS, Feunang YD, Marcu A, Guo AC, Liang K, Vazquez-Fresno R, Sajed T, Johnson D, Li C, Karu N, et al. HMDB 4.0: the human metabolome database for 2018. Nucleic Acids Res. 2018;46(D1):D608–17.
von Mering C, Krause R, Snel B, Cornell M, Oliver SG, Fields S, Bork P. Comparative assessment of large-scale data sets of protein-protein interactions. Nature. 2002;417(6887):399–403.
Sprinzak E, Sattath S, Margalit H. How reliable are experimental protein-protein interaction data? J Mol Biol. 2003;327(5):919–23.
Hart GT, Ramani AK, Marcotte EM. How complete are current yeast and human protein-interaction networks? Genome Biol. 2006;7(11):120.
Diallo I, Seve M, Cunin V, Minassian F, Poisson JF, Michelland S, Bourgoin-Voillard S. Current trends in protein acetylation analysis. Expert Rev Proteomic. 2019;16(2):139–59.
Rahman SA, Schomburg D. Observing local and global properties of metabolic pathways: ‘load points’ and ‘choke points’ in the metabolic networks. Bioinformatics. 2006;22(14):1767–74.
Benson DA, Cavanaugh M, Clark K, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW. GenBank. Nucleic Acids Res. 2013;41:D36-42.
Maglott D, Ostell J, Pruitt KD, Tatusova T. Entrez Gene: gene-centered information at NCBI. Nucleic Acids Res. 2011;39:D52–7.
Hagberg A, Swart P, S Chult D. Exploring network structure, dynamics, and function using NetworkX. 2008.
Lu Y, Valentine JS. Engineering metal-binding sites in proteins. Curr Opin Struct Biol. 1997;7(4):495–500.
Gray HB. Biological inorganic chemistry at the beginning of the 21st century. PNAS. 2003;100(7):3563–8.
Wilson CJ, Apiyo D, Wittung-Stafshede P. Role of cofactors in metalloprotein folding. Q Rev Biophys. 2004;37(3–4):285–314.
Chen W, Bailey JE. Application of the cross-regulation system as a metabolic switch. Biotechnol Bioeng. 1994;43(11):1190–3.
Li Y, Cong H, Liu B, Song J, Sun X, Zhang J, Yang Q. Metabolic engineering of Corynebacterium glutamicum for methionine production by removing feedback inhibition and increasing NADPH level. Antonie Van Leeuwenhoek. 2016;109(9):1185–97.
Park JH, Lee KH, Kim TY, Lee SY. Metabolic engineering of Escherichia coli for the production of L-valine based on transcriptome analysis and in silico gene knockout simulation. PNAS. 2007;104(19):7797–802.
Kim SY, Lee J, Lee SY. Metabolic engineering of Corynebacterium glutamicum for the production of L-ornithine. Biotechnol Bioeng. 2015;112(2):416–21.
Chen Z, Bommareddy RR, Frank D, Rappert S, Zeng AP. Deregulation of feedback inhibition of phosphoenolpyruvate carboxylase for improved lysine production in Corynebacterium glutamicum. Appl Environ Microbiol. 2014;80(4):1388–93.
Geng F, Chen Z, Zheng P, Sun J, Zeng AP. Exploring the allosteric mechanism of dihydrodipicolinate synthase by reverse engineering of the allosteric inhibitor binding sites and its application for lysine production. Appl Microbiol Biotechnol. 2013;97(5):1963–71.
Papin JA, Hunter T, Palsson BO, Subramaniam S. Reconstruction of cellular signalling networks and analysis of their properties. Nat Rev Mol Cell Biol. 2005;6:99.
Weng G, Bhalla US, Iyengar R. Complexity in biological signaling systems. Science. 1999;284(5411):92–6.
Urbanowski ML, Stauffer LT, Stauffer GV. The gcvB gene encodes a small untranslated RNA involved in expression of the dipeptide and oligopeptide transport systems in Escherichia coli. Mol Microbiol. 2000;37(4):856–68.
Nielsen AAK, Der BS, Shin J, Vaidyanathan P, Paralanov V, Strychalski EA, Ross D, Densmore D, Voigt CA. Genetic circuit design automation. Science. 2016;352(6281):aac7341.
Zhang F, Carothers JM, Keasling JD. Design of a dynamic sensor-regulator system for production of chemicals and fuels derived from fatty acids. Nat Biotechnol. 2012;30:354.
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2018YFA0900300, 2018YFA0901400); the International Partnership Program of Chinese Academy of Sciences (153D31KYSB20170121); Tian** Synthetic Biotechnology Innovation Capacity Improvement Project (TSBICIP-PTJS-001, TSBICIP-KJGG-005).
Funding
Publication costs are funded by the National Key Research and Development Program of China (2018YFA0900300, 2018YFA0901400); the International Partnership Program of Chinese Academy of Sciences (153D31KYSB20170121); Tian** Synthetic Biotechnology Innovation Capacity Improvement Project (TSBICIP-PTJS-001, TSBICIP-KJGG-005).
Author information
Authors and Affiliations
Contributions
ZM and HM participated in the design of the study. ZM collected and analyzed the data. ZM and TH designed the online website. ZM, HM, TH and QY wrote and edited the manuscript. All authors revised and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interests.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Written informed consent for publication was obtained from all participants.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mao, Z., Huang, T., Yuan, Q. et al. Construction and analysis of an integrated biological network of Escherichia coli. Syst Microbiol and Biomanuf 2, 165–176 (2022). https://doi.org/10.1007/s43393-021-00051-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43393-021-00051-x