Abstract
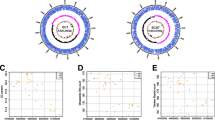
Pectobacterium carotovorum subsp. brasiliense (Pcb) is a gram-negative, plant pathogenic bacterium of the soft rot Enterobacteriaceae (SRE) family. We present the complete genome sequence of Pcb strain BZA12, which reveals that Pcb strain BZA12 carries a single 4,924,809 bp chromosome with 51.97% GC content and comprises 4508 predicted protein-coding genes. Gene annotation of these genes utilized GO, KEGG, and COG databases. In comparison with three closely related soft-rot pathogens, strain BZA12 has 3797 gene families, among which 3107 gene families are identified as orthologous with those of both P. carotovorum subsp. carotovorum PCC21 and P. carotovorum subsp. odoriferum BCS7, as well as 36 putative Unique Gene Families. We selected five putative effectors from the BZA12 genome and transiently expressed them in Nicotiana benthamiana. Candidate effector A12GL002483 was localized in the cell nucleus and induced cell death. This study provides a foundation for a better understanding of the genomic structure and function of Pcb, particularly in the discovery of potential pathogenic factors and for the development of more effective strategies against this pathogen.









Similar content being viewed by others
References
Arnold R, Brandmaier S, Kleine F, Tischler P, Heinz E, Behrens S (2009) Correction: sequence-based prediction of type iii secreted proteins. PLoS Pathog 5:e1000376
Bard J, Winter R (2000) Gene ontology:tool for the unification of biology. Nat Genet 25:25–29
Benson G (1999) Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res 27:573–580
Bent AF, Mackey D (2007) Elicitors, effectors, and R genes: the new paradigm and a lifetime supply of questions. Annu Rev Phytopathol 45:399–436
Caillaud M, Asai S, Rallapalli G, Piquerez S, Fabro G, Jones JDG (2013) A downy mildew effector attenuates salicylic acid–triggered immunity in arabidopsis by interacting with the host mediator complex. PLoS Biol 11:e1001732
Charkowski A, Blanco C, Condemine G, Expert D, Franza T, Hayes C (2012) The role of secretion systems and small molecules in soft-rot Enterobacteriaceae pathogenicity. Annu Rev Phytopathol 50:425–449
Chisholm ST, Coaker G, Day B, Staskawicz BJ (2006) Host-microbe interactions: sha** the evolution of the plant immune response. Cell 124:803–814
Choi O, Kim J (2013) Pectobacterium carotovorum subsp. brasiliense, causing soft rot on paprika in Korea. J Phytopathol 161:125–127
Coburn B, Sekirov I, Finlay BB (2007) Type iii secretion systems and disease. Clin Microbiol Rev 20:535–549
Delcher AL, Bratke KA, Powers EC (2007) Identifying bacterial genes and endosymbiont DNA with glimmer. Bioinformatics 23:673–679
Dong X, Zhang YJ, Zhang Z (2013) Using weakly conserved motifs hidden in secretion signals to identify type-iii effectors from bacterial pathogen genomes. PLoS One 8:e56632
Dou D, Zhou JM (2012) Phytopathogen effectors subverting host immunity: different foes, similar battleground. Cell Host Microbe 12:484–495
Fujimoto T, Yasuoka S, Aono Y, Nakayama T, Ohki T, Sayama M, Maoka T (2016) First report of potato blackleg caused by Pectobacterium carotovorum subsp brasiliense in Japan. Plant Dis 101:241–242
Gardiner L-J (2014) New technologies for high throughput genetic analysis of complex genomes.Ph.D. In: Thesis. University of Liverpool, London
Gardner PP, Daub J, Tate JG (2009) Rfam: updates to the RNA families database. Nucleic Acids Res 37:D136–D140
Glasner JM, Kim HS, Jahn CE, Ma B, Biehl BS, Rissman AI (2008) Niche-specificity and the variable fraction of the Pectobacterium pan-genome. Mol Plant-Microbe Interact 21:1549–1560
Kanehisa M, Goto S, Kawashima S, Okuno Y, Hattori M (2004) The KEGG resource for deciphering the genome. Nucleic Acids Res 32:D277–D280
Kent WJ (2002) Blat--the blast-like alignment tool. Genome Res 12:656–664
Kim HS, Thammarat P, Lommel SA, Hogan CS, Charkowski AO (2011) Pectobacterium carotovorum elicits plant cell death with dspe/f but the p. carotovorum dspe does not suppress callose or induce expression of plant genes early in plant-microbe interactions. Mol Plant-Microbe Interact 24:773–786
Lagesen K, Hallin PF, Rødland E (2007) RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res 35:3100–3108
Laverde AJ, Davey CAH, López EDG (2017) First report of bacterial stem rot of tomatoes caused by Pectobacterium carotovorum subsp. brasiliense in Colombia. Plant Dis 101:830
Li H.L., (2013) Identification of four pathogenic bacteria causing bacterial diseases in vegetables. Master Thesis, Chinese Academy of Agricultural Sciences
Li L, Stoeckert CJ, Roos DS (2003) OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res 13:2178–2189
Lowe TM, Eddy SR (1997) tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25:955–964
Ma B, Hibbing ME, Kim HS, Reedy RM, Yedidia I, Breuer J (2007) Host range and molecular phylogenies of the soft rot enterobacterial genera pectobacterium and dickeya. Phytopathology 97:1150
Magrane M., UniProt Consortium, (2011) UniProt knowledgebase: a hub of integrated protein data. Database : the journal of biological databases and curation, (Oxford), bar009
Mashavha M.L., (2013) Characterisation of Pectobacterium carotovorum subsp. brasiliense isolates causing blackleg and soft rot diseases of potato in south Africa Ph.D. Thesis. Pretoria University, South Africa
Meng X, Chai A, Li B, Ma Z, Shi Y, **e X (2016) Emergence of bacterial soft rot in cucumber caused by Pectobacterium carotovorum subsp. brasiliense in China. Plant Dis 101:279–287
Nabhan S, de Boer SH, Maiss E, Wydra K (2012) Taxonomic relatedness between Pectobacterium carotovorum subsp. carotovorum, Pectobacterium carotovorum subsp. odoriferum and Pectobacterium carotovorum subsp. brasiliense subsp. nov. J Appl Microbiol 113:904–913
Ochiai H, Inoue Y, Takeya M, Sasaki A, Kaku H (2005) Genome sequence of Xanthomonas oryzae pv. oryzae suggests contribution of large numbers of effector genes and insertion sequences to its race diversity. Jpn Agric Res Q 39:275–287
Onkendi EM, Moleleki LN (2014) Characterization of Pectobacterium carotovorum, subsp. carotovorum, and brasiliense, from diseased potatoes in Kenya. Eur J Plant Pathol 139:557–566
Onkendi EM, Ramesh AM, Kwenda S, Naidoo S, Moleleki L (2016) Draft genome sequence of a virulent Pectobacterium carotovorum subsp. brasiliense isolate causing soft rot of cucumber. Genome Announc 4:e01530-15
Pérombelon MCM (2002) Potato diseases caused by soft rot erwinias: an overview of pathogenesis. Plant Pathol 51:1–12
Pérombelon MCM, Kelman A (1980) Ecology of the soft rot erwinias. Annu Rev Phytopathol 18:361–387
Tatusov RL, Fedorova ND (2003) The COG database: an updated version includes eukaryotes. BMC Bioinf 11:4–41
Toth IK, Bell KS, Holeva MC, Birch PRJ (2003) Soft rot erwiniae: from genes to genomes. Mol Plant Pathol 4:17–30
Waldee EL (1945) Comparative studies of some peritrichous phytopathogenic bacteria. Iowa State Coll J Sci 19:435–484
Waleron M, Waleron K, Lojkowska E (2015) First report of Pectobacterium carotovorum subsp. brasiliense causing soft rot on potato and other vegetables in Poland. Plant Dis 99:1271
Wang Y, Huang H, Sun M, Zhang Q, Guo D (2012a) T3db: an integrated database for bacterial type iii secretion system. BMC Bioinf 13:66
Wang Y, Tang H, Debarry JD, Tan X, Li J, Wang X (2012b) Mcscanx: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res 40:e49
Wang J, Wang YH, Zhao T, Dai PG, Chen D, Li XL (2017) First report of tobacco bacterial leaf blight caused by Pectobacterium carotovorum. Subsp. brasiliense in China. Plant Dis 101:830
Werra PD, Bussereau F, Keiser A, Ziegler D (2009) First report of potato blackleg caused by Pectobacterium carotovorum subsp. brasiliense in switzerland. Plant Dis 99:551
Wichmann F, Vorhölter FJ, Hersemann L, Widmer F, Blom J, Niehaus K (2013) The noncanonical type iii secretion system of Xanthomonas translucens pv. graminis is essential for forage grass infection. Mol Plant Pathol 14:576–588
Yang Z (2007) Paml 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Zhang S, Bo H, **a XF, Sun ZR (2007) Bioinformatics research in subcellular localization of protein. Prog Biochem Biophys 34:573–579
Acknowledgements
This project was supported by the National Key R&D Program of China (2017YED0201100).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Huang, Y., Liu, C., Wang, H. et al. Bioinformatic analysis of the complete genome sequence of Pectobacterium carotovorum subsp. brasiliense BZA12 and candidate effector screening. J Plant Pathol 101, 39–49 (2019). https://doi.org/10.1007/s42161-018-0126-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42161-018-0126-7