Abstract
Background
Ezrin, known as a crosslinker between the plasma membrane and actin cytoskeleton, is closely associated with breast cancer (BC) progression. Here, we explored a novel role of ezrin in breast cancer liver metastasis (BCLM).
Methods
The clinical relevance of ezrin was evaluated using in silico tools and confirmed in BC specimens. The effect of ezrin on proliferation, migration and invasion was examined in vitro and in vivo using murine primary liver-metastatic breast cancer cells (mLM). The molecular mechanism involved in ezrin-mediated activation of the Notch1 signaling pathway was elucidated using in vitro models.
Results
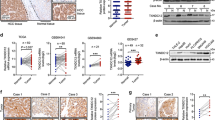
Data-mining demonstrated that ezrin mRNA and protein expression is up-regulated in breast cancer cohorts and has prognostic significance. Ezrin overexpression promotes cell proliferation, migration and invasion in vitro and in vivo. Hairy and enhancer of split-1 (Hes1) is one of the most significantly enriched candidates of differentially expressed genes in ezrin overexpression and control mLM cells. Ezrin can positively regulate Hes1 mRNA and protein expression, and their coexpression was associated with poor prognosis in BC patients. Ezrin promoted BC cell proliferation in a Hes1-dependent manner without directly interacting with Hes1. The functional link between ezrin and Hes1 is dependent on Notch1 activation through promotion of furin-like convertase cleavage.
Conclusion
Our results demonstrated that ezrin drives BCLM through activation of the Notch signaling pathway via furin-like convertase. These findings provide a better understanding of the mechanism of ezrin in breast cancer progression, with the goal of discovering a novel target for the treatment of BCLM in the future.






Similar content being viewed by others
Data availability
The data presented in this study are available in this article.
References
C.M. Perou, T. Sørlie, M.B. Eisen, M. van de Rijn, S.S. Jeffrey, C.A. Rees, J.R. Pollack, D.T. Ross, H. Johnsen, L.A. Akslen, O. Fluge, A. Pergamenschikov, C. Williams, S.X. Zhu, P.E. Lønning, A.L. Børresen-Dale, P.O. Brown, D. Botstein, Molecular portraits of human breast tumours. Nature 406, 747–752 (2000)
M. Sambi, B. Qorri, W. Harless, M.R. Szewczuk, Therapeutic options for metastasis breast cancer. Adv. Exp. Med. Biol. 1152, 131–172 (2019)
H.Q. Cao, Z.W. Zhang, S. Zhao, X.Y. He, H.J. Yu, Q. Yin, Z.P. Zhang, W.W. Gu, L.L. Chen, Y.P. Li, Hydrophobic interaction mediating self-assembled nanoparticles of succinobucol suppress lung metastasis of breast cancer by inhibition of VCAM-1 expression. J. Control Release 205, 162–171 (2015)
Z.C. **ong, G.Z. Deng, X.J. Huang, X. Li, X.H. **e, J. Wang, Z.Y. Shuang, X. Wang, Bone metastasis pattern in initial metastatic breast cancer: a population-based study. Cancer Manag. Res. 10, 287–295 (2018)
M. Smid, Y.X. Wang, Y. Zhang, A.M. Sieuwerts, J. Yu, J.G.M. Klijn, J.A. Foekens, J.W.M. Martens, Subtypes of breast cancer show preferential site of relapse. Cancer Res. 68, 3108–3114 (2008)
G. Pentheroudakis, G. Fountzilas, D. Bafaloukos, V. Koutsoukou, D. Pectasides, D. Skarlos, E. Samantas, H.P. Kalofonos, H. Gogas, N. Pavlidis, Metastatic breast cancer with liver metastases: a registry analysis of clinicopathologic, management and outcome characteristics of 500 women. Breast Cancer Res. Treat. 97, 237–244 (2006)
N.U. Lin, J.R. Bellon, E.P. Winer, CNS metastases in breast cancer. J. Clin. Oncol. 22, 3608–3617 (2004)
Y.L. Tham, K. Sexton, R. Kramer, S. Hilsenbeck, R. Elledge, Primary breast cancer phenotypes associated with propensity for central nervous system metastases. Cancer 107, 696–704 (2006)
M.R. Quigley, O. Fukui, B. Chew, S. Bhatia, S. Karlovits, The shifting landscape of metastatic breast cancer to the CNS. Neurosurg. Rev. 36, 377–382 (2013)
J. Yu, Q.X. Mu, M. Fung, X.L. Xu, L.X. Zhu, R.J.Y. Ho, Challenges and opportunities in metastatic breast cancer treatments: Nano-drug combinations delivered preferentially to metastatic cells may enhance therapeutic response. Pharmacol. Ther. (2022). https://doi.org/10.1016/j.pharmthera.2022.108108
L.M. Yin, T.T. Duan, L. Ulloa, Y.Q. Yang, Ezrin orchestrates signal transduction in airway cells. Rev. Physiol. Biochem. Pharmacol. 174, 1–23 (2018)
S. Yonemura, M. Hirao, Y. Doi, N. Takahashi, T. Kondo, S. Tsukita, Ezrin/radixin/moesin (ERM) proteins bind to a positively charged amino acid cluster in the juxtamembrane cytoplasmic domain of CD44, CD43, and ICAM-2. J. Cell Biol. 140, 885–895 (1998)
L.M. Yin, M. Schnoor, Modulation of membrane-cytoskeleton interactions: ezrin as key player. Trends Cell Biol. 32, 94–97 (2022)
S.I. Muroi, Y. Isohama, Ezrin regulates Ca2+ ionophore-induced plasma membrane translocation of aquaporin-5. Int. J. Mol. Sci. 22, 13505 (2021)
H. Celik, K.P. Sajwan, S.P. Selvanathan, B.J. Marsh, A.V. Pai, Y.S. Kont, J. Han, T.Z. Minas, S. Rahim, H.V. Erkizan, J.A. Toretsky, A. Üren, Ezrin binds to DEAD-box RNA helicase DDX3 and regulates its function and protein level. Mol. Cell Biol. 35, 3145–3162 (2015)
A. Ghaffari, V. Hoskin, G. Turashvili, S. Varma, J. Mewburn, G. Mullins, P.A. Greer, F. Kiefer, A.G. Day, Y. Madarnas, S. SenGupta, B.E. Elliott, Intravital imaging reveals systemic ezrin inhibition impedes cancer cell migration and lymph mode metastasis in breast cancer. Breast Cancer Res. 21, 12 (2019)
R.J. Zhang, S.H. Zhang, R.G. **ng, Q. Zhang, High expression of EZR gene is correlated with the poor overall survival of breast cancer patients. Thorac. Cancer 10, 1953–1961 (2019)
A. Ghaffari, V. Hoskin, A. Szeto, M. Hum, N. Liaghati, K. Nakatsu, D. LeBrun, Y. Madarnas, S. Sengupta, B.E. Elliott, A novel role for ezirn in breast cancer angio/lymphangiogenesis. Breast Cancer Res. 16, 438 (2014)
N. Li, J.N. Kong, Z.H. Lin, Y. Yang, T.F. **, M. Xu, J. Sun, L.Y. Chen, Ezrin promotes breast cancer progression by modulating AKT signals. Br. J. Cancer 120, 703–713 (2019)
R. Limame, A. Wouters, B. Pauwels, E. Fransen, M. Peeters, F. Lardon, O.D. Wever, P. Pauwels, Comparative analysis of dynamic cell viability, migration and invasion assessments by novel real-time technology and classic endpoint assays. PLoS One 7, e46536 (2012)
S. Rahim, A. Üren, A real-time electrical impedance based technique to measure invasion of endothelial cell monolayer by cancer cells. J. Vis. Exp. 2792 (2011)
G.L. Bourne, D.J. Grainger, Development and characterization of an assay for furin activity. J. Immunol. Methods 364, 101–108 (2011)
A. Rani, R. Greenlaw, R.A. Smith, C. Galustian, Hes1 in immunity and cancer. Cytokine Growth Factor Rev. 30, 113–117 (2016)
Y.R. Liang, H.W. Zhang, X.J. Song, Q.F. Yang, Metastatic heterogeneity of breast cancer: molecular mechanism and potential therapeutic targets. Semin. Cancer Biol. 60, 14–27 (2020)
G.D. Venosa, C. Perotti, A. Batlle, A. Casas, The role of cytoskeleton and adhesion proteins in the resistance to photodynamic therapy. Possible therapeutic interventions. Photochem. Photobiol. Sci. 14, 1451–1464 (2015)
A.J. Davidson, W. Wood, Unravelling the actin cytoskeleton: a new competitive edge? Trends Cell Biol. 26, 569–576 (2016)
M.J. Chen, T.F. Liu, L.N. Xu, X.J. Gao, X.H. Liu, C.H. Wang, Q.Y. He, G. Zhang, L.X. Liu, Direct interaction of 14-3-3ζ with ezrin promotes cell migration by regulating the formation of membrane ruffle. J. Mol. Biol. 426, 3118–3133 (2014)
J. Jeong, J. Choi, W. Kim, P. Dann, F. Takyar, J.V. Gefter, P.A. Friedman, J.J. Wysolmerski, Inhibition of ezrin causes PKCα-mediated internalization of erbb2/HER2 tyrosine kinase in breast cancer cells. J. Bio. Chem. 294, 887–901 (2019)
T. Celià-Terrassa, Y.B. Kang, Metastatic niche functions and therapeutic opportunities. Nat. Cell Biol. 20, 868–877 (2018)
H. Rezaeeyan, R. Shirzad, T.D. McKee, N. Saki, Role of chemokines in metastatic niche: new insights along with a diagnostic and prognostic approach. APMIS 26, 359–370 (2018)
Z.H. Liu, X.M. Dai, B. Du, Hes1: a key role in stemness, metastasis and multidrug resistance. Cancer Biol. Ther. 16, 353–359 (2015)
X.Y. Li, Y. Cao, M. Li, F. **, Upregulation of Hes1 promotes cell proliferation and invasion in breast cancer as a prognosis marker and therapy target via the AKT pathway and EMT process. J. Cancer 9, 757–766 (2018)
M. Hartmann, L.M. Parra, A. Ruschel, S. Böhme, Y. Li, H. Morrison, A. Herrlich, P. Herrlich, Tumor suppressor NF2 blocks cellular migration by inhibiting ectodomain cleavage of CD44. Mol. Cancer Res. 13, 879–890 (2015)
A. Darmellah, A. Rayah, R. Auger, M.H. Cuif, M. Prigent, M. Arpin, A. Alcover, C. Delarasse, J.M. Kanellopoulos, Ezrin/radixin/moesin are required for the purinergic P2X7 receptor (P2X7R)-dependent processing of the amyloid precursor protein. J. Biol. Chem. 287, 34583–34595 (2012)
F. Logeat, C. Bessia, C. Brou, O. LeBail, S. Jarriault, N.G. Seidah, The Notch1 receptor is cleaved constitutively by a furin-like convertase. Proc. Natl. Acad. Sci. U.S.A. 95, 8108–8112 (1998)
A. Zolkiewska, ADAM proteases: ligand processing and modulation of the Notch pathway. Cell Mol. Life Sci. 65, 2056–2068 (2008)
J.S. Mumm, E.H. Schroeter, M.T. Saxena, A. Griesemer, X. Tian, D.J. Pan, W.J. Ray, R. Kopan, A ligand-induced extracellular cleavage regulates gamma-secretase-like proteolytic activation of Notch1. Mol. Cell. 5, 197–206 (2000)
T.A. Martin, The role of tight junction in cancer metastasis. Semin. Cell Dev. Biol. 36, 224–231 (2014)
D. Günzel, A.S.L. Yu, Claudins and the modulation of tight junction permeability. Physiol. Rev. 93, 525–569 (2013)
S. Tabariès, P.M. Siegel, The role of claudins in cancer metastasis. Oncogene 36, 1176–1190 (2017)
S. Tabariès, V. Ouellet, B.E. Hsu, M.G. Annis, A.A.N. Rose, L. Meunier, E. Carmona, C.E. Tam, A. Mes-Masson, P.M. Siegel, Granulocytic immune infiltrates are essential for the efficient formation of breast cancer liver metastases. Breast Cancer Res. 17, 45 (2015)
F. Dupuy, S. Tabariès, S. Andrzejewski, Z. Dong, J. Blagih, M.G. Annis, A. Omeroglu, D. Gao, S. Leung, E. Amir, M. Clemons, A. Aguilar-Mahecha, M. Basik, E.E. Vincent, J. St-Pierre, R.G. Jones, P.M. Siegel, PDK1-dependent metabolic reprogramming dictates metastatic potential in breast cancer. Cell Metab. 22, 577–589 (2015)
S. Tabariès, Z. Dong, M.G. Annis, A. Omeroglu, F. Pepin, V. Ouellet, C. Russo, M. Hassanain, P. Metrakos, Z. Diaz, M. Basik, N. Bertos, M. Park, C. Guettier, R. Adam, M. Hallett, P.M. Siegel, Claudin-2 is selectively enriched in and promotes the formation of breast cancer liver metastases through engagement of integrin complexes. Oncogene 30, 1318–1328 (2011)
G. Zheng, G.V. Fon, W. Meixner, A. Creekmore, Y. Zong, M.K. Dame, J. Colacino, P.H. Dedhia, S. Hong, J.W. Wiley, Chronic stress and intestinal barrier dysfunction: Glucocorticoid receptor and transcription repressor HES1 regulate tight junction protein Claudin-1 promoter. Sci Rep. 7, 4502 (2017)
Acknowledgements
This work was supported by National Natural Science Foundation of China (NSFC 81602333, NSFC 31600746, NSFC 81702567, NSFC 81671406).
Author information
Authors and Affiliations
Contributions
Experimentation, M.J.C., Y.P., H.B.L., F.N., X.W.G., H.M.H., M.Z., Y.Y.D.; Cell culture, S.J.L.; IHC analysis, Q.S.L.; Conceiving and designing the study, Y.T.; Writing original draft, M.J.C.; Writing, review and editing, G.E.L. All authors have read and agreed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
The present study was approved by the Ethics Committee of Guangzhou Women and Children’s Medical Center. The animal experiments were approved by the Institutional Animal Care and Use Committee of Guangzhou Medical University.
Consent for publication
The authors declare that they agree to submit the article for publication.
Competing interests
The authors declare that they have no conflicts of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.

Fig. S1
Isolation of murine primary liver-metastatic breast cancer cells and establishment of an animal model. (A) Isolation of murine primary liver-metastatic breast cancer cells. Highly aggressive cells were isolated from a lung metastatic lesion of MMTV-PyMT transgenic mouse and then were orthotopically implanted into the mammary fat pad of BALB/c nude mice. After liver metastasis, murine primary liver-metastatic breast cancer cells (mLM) were obtained from the liver lesions and successfully isolated. (B) Establishment of a breast cancer liver metastasis animal model. The mLM cells mainly metastasized to the liver after orthotopic implantation or injection by tail vein. This experiment was performed more than three times. This is a versatile tool to study breast cancer liver metastasis in vitro and in vivo. (PNG 625 kb)

Fig. S2
The expression of ezrin in human cancers was investigated using the TIMER database. Ezrin was significantly upregulated in BRCA, CHOL, KIRC, KIRP, STAD and UCEC, downregulated in COAD, KICH, LUAD, LUSC, PRAD and THCA, and was not altered in BLCA, ESCA, HNSC, LIHC and READ. Tumor vs. Normal, *P<0.05, **P<0.01, ***P<0.001. ACC, adrenocortical carcinoma; BLCA, bladder urothelial carcinoma; BRCA, breast invasive carcinoma; CESC, cervical squamous cell carcinoma; CHOL, cholangiocarcinoma; COAD, colon adenocarcinoma; DLBC, diffuse large B-cell lymphoma; ESCA, esophageal carcinoma; GBM, glioblastoma multiforme; HNSC, head and neck squamous cell carcinoma; KICH, kidney chromophobe; KIRC, kidney renal clear cell carcinoma; KIRP, kidney renal papillary cell carcinoma; LGG, lower grade glioma; LIHC, liver hepatocellular carcinoma; LUAD, lung adenocarcinoma; LUSC, lung squamous cell carcinoma; MESO, mesothelioma; OV, ovarian serous cystadenocarcinoma; PAAD, pancreatic adenocarcinoma; PCPG, pheochromocytoma and paraganglioma; PRAD, prostate adenocarcinoma; READ, rectum adenocarcinoma; SARC, sarcoma; SKCM, skin cutaneous melanoma; STAD, stomach adenocarcinoma; TGCT, testicular germ cell tumors; THCA, thyroid carcinoma; THYM, thymoma; UCEC, uterine corpus endometrial carcinoma; UCS, uterine carsinosarcoma; UVM, uveal melanoma. (PNG 2848 kb)

Fig. S3
N-cadherin and vimentin expression were detected in ezrin overexpression mLM cells and control cells using western blot. (PNG 55 kb)

Fig. S4
Ezrin expression was confirmed in ezrin overexpression MDA-MB-231 cells. (PNG 39 kb)

Fig. S5
Ezrin expression was detected in MDA-MB-231 and 4T1 cells after transfection with ezrin siRNA pool. (PNG 105 kb)

Fig. S6
Hes1 expression was detected using qPCR after transfection with ezrin siRNA pool ***P<0.001, 4T1 cells transfected with ezrin siRNA pool versus 4T1 cells transfected with NC siRNA, MDA-MB-231 cells transfected with ezrin siRNA pool versus MDA-MB-231 cells transfected with NC siRNA. (PNG 47 kb)

Fig. S7
Hes1 expression was detected using qPCR and western blot after transfection with Hes1 siRNA pool ***P<0.001, mLM cells transfected with Hes siRNA pool versus mLM cells transfected with NC siRNA pool. (PNG 85 kb)

Fig. S8
N-cadherin and vimentin expression were detected in ezrin overexpression mLM cells after transfection with Hes1 siRNA pool using western blot. (PNG 90 kb)

Fig. S9
Colocalization of Notch1 and EEA1 (the marker of endosome). Previous study have shown that endocytosis of Notch is critical for proper NICD production. In order to confirm that the vesicles containing cleaved Notch1 are the endosomes, the endosome marker EEA1 was used to detect the vesicles. The result showed Notch1 and EAA1 colocalized in the vesicles, suggesting they are the endosomes containing Notch1. Conbined with the result of Fig. 5G, ezrin overexpression promoted the number of endosomes containing Notch1. (PNG 1552 kb)

Fig. S10
p-Ezrin and total ezrin expression was detected in MCF-7 cells and MDA-MB-231 cells using western blot. (PNG 86 kb)

Fig. S11
The effect of NSC305787 and decRVKR-CMK treatment on ezrin overexpression in lung metastasis. The mice were sacrificed two weeks after tain vein injection and the lung tissues were collected. (PNG 863 kb)

Fig. S12
GSEA identified significant association between ezrin and cell junction in the ezrin overexpression cells when compared with control cells. (PNG 161 kb)

Fig. S13
Heatmap of claudin family members. Heatmap of gene expression levels of claudin family members in parental 4T1 cells and liver aggressive 4T1 cells. The microarray data was obtained from GSE62598. (B) Heatmap of gene expression levels of claudin family members in control mLM cells and ezrin overexpression mLM cells. Red color represents upregulatd genes, and blue represents downregulated genes. (PNG 98 kb)
Supplementary Table 1
(DOCX 20 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, M., Pan, Y., Liu, H. et al. Ezrin accelerates breast cancer liver metastasis through promoting furin-like convertase-mediated cleavage of Notch1. Cell Oncol. 46, 571–587 (2023). https://doi.org/10.1007/s13402-022-00761-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-022-00761-x