Abstract
Purpose
GAS41 is a YEATS domain protein that binds to acetylated histone H3 to promote the chromatin deposition of H2A.Z in non-small cell lung cancer. The role of GAS41 in pancreatic cancer is still unknown. Here, we aimed to reveal this role.
Methods
GAS41 expression in pancreatic cancer tissues and cell lines was examined using qRT-PCR, Western blotting and immunohistochemistry. MTT, colony formation, spheroid formation and in vivo tumorigenesis assays were performed to assess the proliferation, tumorigenesis, stemness and gemcitabine (GEM) resistance of pancreatic cancer cells. Mechanistically, co-immunoprecipitation (co-IP) and chromatin immunoprecipitation (ChIP) assays were used to evaluate the roles of GAS41, H2A.Z.2 and Notch1 in pancreatic cancer.
Results
We found that GAS41 is overexpressed in human pancreatic cancer tissues and cell lines, and that its expression increases following the acquisition of GEM resistance. We also found that GAS41 up-regulates Notch, as well as pancreatic cancer cell stemness and GEM resistance in vitro and in vivo. We show that GAS41 binds to H2A.Z.2 and activates Notch and its downstream mediators, thereby regulating stemness and drug resistance. Depletion of GAS41 or H2A.Z.2 was found to down-regulate Notch and to sensitize pancreatic cancer cells to GEM.
Conclusion
Our data indicate that GAS41 mediates proliferation and GEM resistance in pancreatic cancer cells via H2A.Z.2 and Notch1.









Similar content being viewed by others
Data availability
The datasets supporting the conclusions of this article are included in the article (and its additional files 1 to 17: Supplementary Material).
Abbreviations
- GEM :
-
Gemcitabine
- CSCs:
-
Cancer stem cells
- GAS41:
-
Glioma amplified sequence 41
- GR:
-
GEM-resistant
- BrDs:
-
Bromodomains
- ChIP:
-
Chromatin immunoprecipitation
- NSCLC:
-
non-small cell lung cancer
References
P. Rawla, T. Sunkara, V. Gaduputi, Epidemiology of Pancreatic Cancer: Global Trends, Etiology and Risk Factors. World J Oncol. 10, 10–27 (2019)
F. Bray, J. Ferlay, I. Soerjomataram, R.L. Siegel, L.A. Torre, A. Jemal, Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 68, 394–424 (2018)
R.L. Siegel, K.D. Miller, H.E. Fuchs, A. Jemal, Cancer Statistics, 2021. CA: A Cancer Journal for Clinicians. 71 (2021)
M. Manrai, T. Tilak, S. Dawra, S. Srivastava, A. Singh, Current and emerging therapeutic strategies in pancreatic cancer: Challenges and opportunities. World J. Gastroenterol. 27, 6572–6589 (2021)
M.P. Kim, G.E. Gallick, Gemcitabine resistance in pancreatic cancer: picking the key players. Clin Cancer Res. 14, 1284–1285 (2008)
A. Jain, V. Bhardwaj, Therapeutic resistance in pancreatic ductal adenocarcinoma: Current challenges and future opportunities. World J. Gastroenterol. 27, 6527–6550 (2021)
B. Cpa, A. Bf, C. Bba, B. Dc, D. Gjpa, B. Pd, E. Yga, F. Ega, GSK3β as a novel promising target to overcome chemoresistance in pancreatic cancer. Drug Resistance Updates. 58, 100779 (2021)
A. Bvksl, B. Bf, C. Sl, D. Sp, E. Nsy, F. Pvb, I. Makgh, J. Mss, B. Gpn, Small molecule tyrosine kinase inhibitors and pancreatic cancer—Trials and troubles. Seminars in Cancer Biology. 54, 149–167 (2019)
S.R. Martins-Neves, A.M. Cleton-Jansen, C.M.F. Gomes, Therapy-induced enrichment of cancer stem-like cells in solid human tumors: Where do we stand? Pharmacol Res. 137, 193–204 (2018)
S. Chatterjee, P.C. Sil, Targeting the crosstalks of Wnt pathway with Hedgehog and Notch for cancer therapy. Pharmacol Res. 142, 251–261 (2019)
V. Venkatesh, R. Nataraj, G.S. Thangaraj, M. Karthikeyan, A. Gnanasekaran, S.B. Kaginelli, G. Kuppanna, C.G. Kallappa, K.M. Basalingappa, Targeting Notch signalling pathway of cancer stem cells. Stem Cell Investig. 5, 5 (2018)
Z. Wang, Y. Li, D. Kong, S. Banerjee, A. Ahmad, A.S. Azmi, S. Ali, J.L. Abbruzzese, G.E. Gallick, F.H. Sarkar, Acquisition of epithelial-mesenchymal transition phenotype of gemcitabine-resistant pancreatic cancer cells is linked with activation of the notch signaling pathway. Cancer Res. 69, 2400–2407 (2009)
B.D. Giaimo, F. Ferrante, A. Herchenrother, S.B. Hake, T. Borggrefe, The histone variant H2A.Z in gene regulation. Epigenetics Chromatin. 12, 37 (2019)
D. Corujo, M. Buschbeck, Post-Translational Modifications of H2A Histone Variants and Their Role in Cancer. Cancers (Basel). 10 (2018)
B.D. Giaimo, F. Ferrante, D.M. Vallejo, K. Hein, I. Gutierrez-Perez, A. Nist, T. Stiewe, G. Mittler, S. Herold, T. Zimmermann, M. Bartkuhn, P. Schwarz, F. Oswald, M. Dominguez, T. Borggrefe, Histone variant H2A.Z deposition and acetylation directs the canonical Notch signaling response. Nucleic Acids Res. 46, 8197–8215 (2018)
J.M. Eirin-Lopez, R. Gonzalez-Romero, D. Dryhurst, T. Ishibashi, J. Ausio, The evolutionary differentiation of two histone H2A.Z variants in chordates (H2A.Z-1 and H2A.Z-2) is mediated by a stepwise mutation process that affects three amino acid residues. BMC Evol Biol. 9, 31 (2009)
C. Vardabasso, A. Gaspar-Maia, D. Hasson, S. Punzeler, D. Valle-Garcia, T. Straub, E.C. Keilhauer, T. Strub, J. Dong, T. Panda, C.Y. Chung, J.L. Yao, R. Singh, M.F. Segura, B. Fontanals-Cirera, A. Verma, M. Mann, E. Hernando, S.B. Hake, E. Bernstein, Histone Variant H2A.Z.2 Mediates Proliferation and Drug Sensitivity of Malignant Melanoma. Mol Cell. 59, 75–88 (2015)
M.M. Wong, L.K. Cox, J.C. Chrivia, The chromatin remodeling protein, SRCAP, is critical for deposition of the histone variant H2A.Z at promoters. J Biol Chem. 282, 26132–26139 (2007)
A. Cuadrado, N. Corrado, E. Perdiguero, V. Lafarga, P. Munoz-Canoves, A.R. Nebreda, Essential role of p18Hamlet/SRCAP-mediated histone H2A.Z chromatin incorporation in muscle differentiation. EMBO J. 29, 2014–2025 (2010)
C.C. Hsu, J. Shi, C. Yuan, D. Zhao, S. Jiang, J. Lyu, X. Wang, H. Li, H. Wen, W. Li, X. Shi, Recognition of histone acetylation by the GAS41 YEATS domain promotes H2A.Z deposition in non-small cell lung cancer. Genes Dev. 32, 58–69 (2018)
D.G. Berta, H. Kuisma, N. Vlimki, M. Risnen, L.A. Aaltonen, Deficient H2A.Z deposition is associated with genesis of uterine leiomyoma. Nature. 596, 398–403 (2021)
G.M. Christman, H. Tang, I. Ahmad, J.M. Stribley, Differential expression of the Notch signal transduction pathway: ligands, receptors and Numb in uterine leiomyomas vs. myometrium. Fertility & Sterility. 88, S72–S72 (2007)
X. Hu, P. Chen, Y. Wu, K. Wang, Y. Xu, H. Chen, L. Zhang, R. Wu, K.A. Webster, H. Yu, W. Zhu, J. Wang, MiR-211/STAT5A Signaling Modulates Migration of Mesenchymal Stem Cells to Improve its Therapeutic Efficacy. Stem Cells. 34, 1846–1858 (2016)
Y. Li, H. Wen, Y. **, K. Tanaka, H. Wang, D. Peng, Y. Ren, Q. **, S.Y. Dent, W. Li, H. Li, X. Shi, AF9 YEATS domain links histone acetylation to DOT1L-mediated H3K79 methylation. Cell. 159, 558–571 (2014)
Z.H. Liu, X.M. Dai, B. Du, Hes1: a key role in stemness, metastasis and multidrug resistance. Cancer Biol Ther. 16, 353–359 (2015)
C.B. Benton, W. Fiskus, K.N. Bhalla, Targeting Histone Acetylation: Readers and Writers in Leukemia and Cancer. Cancer J. 23, 286–291 (2017)
T. Fujisawa, P. Filippakopoulos, Functions of bromodomain-containing proteins and their roles in homeostasis and cancer. Nat Rev Mol Cell Biol. 18, 246–262 (2017)
J. Qi, Bromodomain and extraterminal domain inhibitors (BETi) for cancer therapy: chemical modulation of chromatin structure. Cold Spring Harb Perspect Biol. 6, a018663 (2014)
D. Zhao, H. Guan, S. Zhao, W. Mi, H. Wen, Y. Li, Y. Zhao, C.D. Allis, X. Shi, H. Li, YEATS2 is a selective histone crotonylation reader. Cell Res. 26, 629–632 (2016)
L. Wan, H. Wen, Y. Li, J. Lyu, Y. **, T. Hoshii, J.K. Joseph, X. Wang, Y.E. Loh, M.A. Erb, A.L. Souza, J.E. Bradner, L. Shen, W. Li, H. Li, C.D. Allis, S.A. Armstrong, X. Shi, ENL links histone acetylation to oncogenic gene expression in acute myeloid leukaemia. Nature. 543, 265–269 (2017)
K. Kim, V. Punj, J. Choi, K. Heo, J.M. Kim, P.W. Laird, W. An, Gene dysregulation by histone variant H2A.Z in bladder cancer. Epigenetics Chromatin. 6, 34 (2013)
H.D. Yang, P.J. Kim, J.W. Eun, Q. Shen, H.S. Kim, W.C. Shin, Y.M. Ahn, W.S. Park, J.Y. Lee, S.W. Nam, Oncogenic potential of histone-variant H2A.Z.1 and its regulatory role in cell cycle and epithelial-mesenchymal transition in liver cancer. Oncotarget. 7, 11412–11423 (2016)
U. Fischer, D. Heckel, A. Michel, M. Janka, T. Hulsebos, E. Meese, Cloning of a novel transcription factor-like gene amplified in human glioma including astrocytoma grade I. Hum Mol Genet. 6, 1817–1822 (1997)
Acknowledgements
The authors are grateful for the support provided by the Tongji University School of Medicine and Shanghai Tenth People’s Hospital.
Funding
This study was supported by the National Natural Science Foundation of China (No. 81801805), the Shanghai Sailing Program (No.16YF1409000) and the Climbing Talent Program of Shanghai Tenth People’s Hospital (No. 2021SYPDRC014).
Author information
Authors and Affiliations
Contributions
Shilong Han, Chuanwu Cao, Rui Liu: Conceptualization, Methodology, Software; Maoquan Li and ** Zhang: Supervision, Project administration; Shilong Han, Chuanwu Cao, Rui Liu: Data curation, Writing-Original draft preparation, Investigation; YifengYuan, Long Pan, Minjie Xu, Chao Hu, ** Zhang: Writing-Reviewing and Editing, Funding acquisition.
Corresponding authors
Ethics declarations
Ethical approval and consent to participate
All participating patients provided written informed consent and the human studies were approved by the Institute Research Ethics Committee of Shanghai Tenth People’s Hospital. The study methodologies conformed to the standards set by the Declaration of Helsinki. The animal studies were performed in accordance with ARRIVE guidelines and were approved by the Institute Research Ethics Committee of the Shanghai Tenth People’s Hospital.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information

ESM 1.
Supplement Figure 1. GAS41 aggravates resistance of GR cells. (A) The IC50 values of GEM with 24-hour treatment. (B) Cell viability after 24-hour treatment with indicated concentrations of GEM was evaluated with the MTT assay. (C-D) The mRNA levels of the stemness-related genes ALDH, ABCG2, CD133, and Nestin by qRT-PCR. (E) The GAS41 mRNA and protein levels in parental and GR BXPC3 cells transfected with GAS41 or vector. (F) The GAS41 mRNA and protein levels in parental and GR MiaPaCa2 cells transfected with shGAS41 or scr. Parental (G) and GR (H) BXPC3 and MiaPaCa2 cells were transfected with GAS41, vector, shGAS41, or scr for indicated hours and indicated concentrations. (I-J) The mRNA and protein levels of the chemoresistance-related genes by qRT-PCR and western blot analysis, respectively. (K-L) The mRNA and protein levels of the stemness-related genes by qRT-PCR and western blot analysis, respectively. n=3, &, #P<0.05, **, &&, ##, $$P<0.01. (PNG 2654 kb)

ESM 2.
Supplement Figure 2. GAS41 aggravates resistance of GR cells by upregulating Notch. (A, B) GR BXPC3 cells were transfected with GAS41 or vector for 24 hours. Untransfected resistant cells were included as control. The mRNA (A) and protein (B) levels of Notch1 and Hes1 were determined with qRT-PCR and western blot analysis, respectively. (C, D) GR MiaPaCa2 cells were transfected with shGAS41 or scr for 24 hours. Untransfected resistant cells were included as control. The mRNA (C) and protein (D) levels of Notch1 and Hes1 were determined with qRT-PCR and western blot analysis, respectively. (E, F) The Notch1 mRNA (E) and protein (F) levels in parental and GR BXPC3 cells transfected with shNotch1 or scr. (G, H) The Notch1 mRNA (G) and protein (H) levels in parental and GR MiaPaCa2 cells transfected with Notch1 or vector. (I) GR BXPC3 cells were co-transfected with GAS41 and scr or GAS41 and shNotch1 for 24 hours. GR MiaPaCa2 cells were co-transfected with shGAS41 and vector or shGAS41 and Notch1 for 24 hours. The cells were treated with GEM at indicated concentrations for 24 hours. The cell viability was determined with the MTT assay. n=3, *P<0.05, **, ##P<0.01. (PNG 519 kb)

ESM 3.
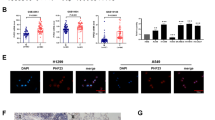
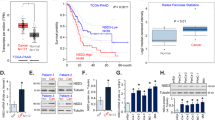
Supplement Figure 3. H2A.Z.2.1 and H2AZ.1 was up-regulated in human pancreatic cancer (A, B) The H2A.Z.2 mRNA levels in human pancreatic cancer and adjacent normal tissues were examined by qRT-PCR. n=87, **P<0.01. (C-F) H2AZ protein levels after H2AZ knockdown or overexpression by Western Blot. (G) H2AZ protein levels were examined by Western Blot in BXPC3 and MiaPaCa2. (PNG 368 kb)

ESM 4.
Supplement Figure 4. H2A.Z.2 knockdown in GR cells restores sensitivity to GEM. GR BXPC3 (A, B, E – G) and MiaPaCa2 (C, D, H – J) cells were transfected with shH2A.Z.2-1, shH2A.Z.2-2, or scr for 24 hours. (A, C) The interference efficiency of H2A.Z.2 is tested by qRT-PCR. (B, D) Cell viability after 24-hour treatment with indicated concentrations of GEM by the MTT assay. (E, H) The mRNA levels of the stemness- and chemoresistance-related genes by qRT-PCR. (F-G, I-J) The protein levels of the stemness- and chemoresistance-related genes by western blot analysis. n=3, *, #, $P<0.05, **, &&, ##, $$P<0.01. (PNG 738 kb)

ESM 5.
Supplement Figure 5. H2A.Z.2 upregulates Notch in GR cells. GR BXPC3 (A, B) and MiaPaCa2 (C, D) cells were transfected with shH2A.Z.2 or scr for 24 hours. Untransfected cells were included as control. (A, C) The mRNA levels of Notch1 and Hes1 by qRT-PCR. n=3, *P<0.05, **, ##P<0.01. (B, D) The protein levels of Notch1 and Hes1 by western blot analysis. (E, F) ChIP analysis was performed with primers targeted to the promoter region of Notch1 and Hes1. IgG was used as control. (PNG 369 kb)

ESM 6
(PNG 1556 kb)

ESM 7
(PNG 1922 kb)

ESM 8
(PNG 542 kb)

ESM 9
(PNG 1014 kb)

ESM 10
(PNG 1489 kb)

ESM 11
(PNG 1231 kb)

ESM 12
(PNG 846 kb)

ESM 13
(PNG 1284 kb)

ESM 14
(PNG 1163 kb)

ESM 15
(PNG 476 kb)
ESM 16
(DOCX 17 kb)
Rights and permissions
About this article
Cite this article
Han, S., Cao, C., Liu, R. et al. GAS41 mediates proliferation and GEM chemoresistance via H2A.Z.2 and Notch1 in pancreatic cancer. Cell Oncol. 45, 429–446 (2022). https://doi.org/10.1007/s13402-022-00675-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-022-00675-8