Abstract
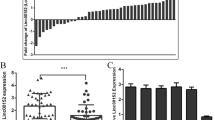
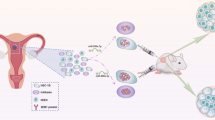
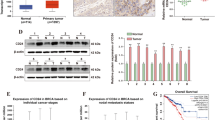
Epithelial ovarian cancer (EOC) is a common type of gynecologic cancer, which accounts for the majority of deaths among all gynecologic malignant tumors in developed countries. A series of recent studies suggested that miR-381 might play important roles in the development of various cancer types. However, the biological function of miR-381 in EOC remains to be investigated. We examined the levels of miR-381 expression in EOC tissues and cell lines. We identified miR-381 target genes by bioinformatic prediction. We also characterized the phenotype regarding cell proliferation, cell migration, and cell invasion in EOC cells lines with altered expression levels of both miR-381 and its target gene, YY1. The expression levels of miR-381 were downregulated in EOC tissues and cell lines. Overexpression of miR-381 significantly inhibited EOC cell proliferation, migration, and invasion. Restoration of YY1 expression partially reversed the phenotype induced by miR-381 overexpression. Knockdown of miR-381 target gene, YY1, mimicked the phenotype induced by miR-381 overexpression. MiR-381 regulated EOC cell through miR-381/YY1/p53 and miR-381/YY1/Wnt signaling axis. We concluded that miR-381 inhibited EOC cell proliferation, migration, and invasion, at least in part, via suppressing YY1 expression.







Similar content being viewed by others
References
Granato T, Midulla C, Longo F, Colaprisca B, Frati L, Anastasi E. Role of he4, ca72.4, and ca125 in monitoring ovarian cancer. Tumour Biol J Int Soc Oncodevelopmental Biol Med. 2012;33:1335–9.
Bristow RE. Surgical standards in the management of ovarian cancer. Curr Opin Oncol. 2000;12:474–80.
Chang SJ, Bristow RE. Evolution of surgical treatment paradigms for advanced-stage ovarian cancer: redefining 'optimal' residual disease. Gynecol Oncol. 2012;125:483–92.
Harter P, Muallem ZM, Buhrmann C, Lorenz D, Kaub C, Hils R, et al. Impact of a structured quality management program on surgical outcome in primary advanced ovarian cancer. Gynecol Oncol. 2011;121:615–9.
Harries M, Gore M. Part i: chemotherapy for epithelial ovarian cancer-treatment at first diagnosis. Lancet Oncol. 2002;3:529–36.
Matei DE, Nephew KP. Epigenetic therapies for chemoresensitization of epithelial ovarian cancer. Gynecol Oncol. 2010;116:195–201.
Winter J, Jung S, Keller S, Gregory RI, Diederichs S. Many roads to maturity: microrna biogenesis pathways and their regulation. Nat Cell Biol. 2009;11:228–34.
Osada H, Takahashi T. Micrornas in biological processes and carcinogenesis. Carcinogenesis. 2007;28:2–12.
Zhang Q, Tang Q, Qin D, Yu L, Huang R, Lv G, et al. Role of microrna 30a targeting insulin receptor substrate 2 in colorectal tumorigenesis. Mol Cell Biol. 2015;35:988–1000.
Sui X, Wang X, Han W, Li D, Xu Y, Lou F, et al. Micrornas-mediated cell fate in triple negative breast cancers. Cancer Lett. 2015;361:8–12.
Iorio MV, Croce CM. Microrna involvement in human cancer. Carcinogenesis. 2012;33:1126–33.
Lee SH, Jung YD, Choi YS, Lee YM. Targeting of runx3 by mir-130a and mir-495 cooperatively increases cell proliferation and tumor angiogenesis in gastric cancer cells. Oncotarget. 2015.
Guo M, Zhang X, Wang G, Sun J, Jiang Z, Khadarian K, et al. Mir-603 promotes glioma cell growth via wnt/beta-catenin pathway by inhibiting wif1 and ctnnbip1. Cancer Lett. 2015;360:76–86.
Liu YN, Yin J, Barrett B, Sheppard-Tillman H, Li D, Casey OM, et al. Loss of androgen-regulated microrna 1 activates src and promotes prostate cancer bone metastasis. Mol Cell Biol. 2015;35:1940–51.
Li X, Li H, Zhang R, Liu J. Microrna-449a inhibits proliferation and induces apoptosis by directly repressing e2f3 in gastric cancer. Cell Physiol Biochem Int J Exp Cell Physiol, Biochem Pharmacol. 2015;35:2033–42.
Tang H, Liu X, Wang Z, She X, Zeng X, Deng M, et al. Interaction of hsa-mir-381 and glioma suppressor lrrc4 is involved in glioma growth. Brain Res. 2011;1390:21–32.
Liang Y, Zhao Q, Fan L, Zhang Z, Tan B, Liu Y, et al. Down-regulation of microrna-381 promotes cell proliferation and invasion in colon cancer through up-regulation of lrh-1. Biomed Pharmacother Biomed Pharmacotherapie. 2015;75:137–41.
Rothschild SI, Tschan MP, Jaggi R, Fey MF, Gugger M, Gautschi O. Microrna-381 represses id1 and is deregulated in lung adenocarcinoma. J Thorac Oncol : Off Publ Int Assoc Study Lung Cancer. 2012;7:1069–77.
Chen B, Liu B. [mirna-381 inhibits the invasion of renal carcinoma and the underlying mechanisms]. Zhong nan da xue xue bao Yi xue ban J Cent South Univ Med Sci. 2015;40:1053–9.
Chen B, Duan L, Yin G, Tan J, Jiang X. Simultaneously expressed mir-424 and mir-381 synergistically suppress the proliferation and survival of renal cancer cells—cdc2 activity is up-regulated by targeting wee1. Clinics. 2013;68:825–33.
Donohoe ME, Zhang X, McGinnis L, Biggers J, Li E, Shi Y. Targeted disruption of mouse yin yang 1 transcription factor results in peri-implantation lethality. Mol Cell Biol. 1999;19:7237–44.
Shi Y, Seto E, Chang LS, Shenk T. Transcriptional repression by yy1, a human gli-kruppel-related protein, and relief of repression by adenovirus e1a protein. Cell. 1991;67:377–88.
Gordon S, Akopyan G, Garban H, Bonavida B. Transcription factor yy1: structure, function, and therapeutic implications in cancer biology. Oncogene. 2006;25:1125–42.
Zhang Q, Stovall DB, Inoue K, Sui G. The oncogenic role of yin yang 1. Crit Rev Oncog. 2011;16:163–97.
Sui G, el Affar B, Shi Y, Brignone C, Wall NR, Yin P, et al. Yin yang 1 is a negative regulator of p53. Cell. 2004;117:859–72.
Yakovleva T, Kolesnikova L, Vukojevic V, Gileva I, Tan-No K, Austen M, et al. Yy1 binding to a subset of p53 DNA-target sites regulates p53-dependent transcription. Biochem Biophys Res Commun. 2004;318:615–24.
Yokoyama NN, Pate KT, Sprowl S, Waterman ML. A role for yy1 in repression of dominant negative lef-1 expression in colon cancer. Nucleic Acids Res. 2010;38:6375–88.
Zhang N, Li X, Wu CW, Dong Y, Cai M, Mok MT, et al. Microrna-7 is a novel inhibitor of yy1 contributing to colorectal tumorigenesis. Oncogene. 2013;32:5078–88.
Kang W, Tong JH, Chan AW, Zhao J, Dong Y, Wang S, et al. Yin yang 1 contributes to gastric carcinogenesis and its nuclear expression correlates with shorter survival in patients with early stage gastric adenocarcinoma. J Transl Med. 2014;12:80.
Tang H, Wang Z, Liu Q, Liu X, Wu M, Li G. Disturbing mir-182 and −381 inhibits brd7 transcription and glioma growth by directly targeting lrrc4. PLoS One. 2014;9, e84146.
Liang HQ, Wang RJ, Diao CF, Li JW, Su JL, Zhang S. The pttg1-targeting mirnas mir-329, mir-300, mir-381, and mir-655 inhibit pituitary tumor cell tumorigenesis and are involved in a p53/pttg1 regulation feedback loop. Oncotarget. 2015;6:29413–27.
Vilming Elgaaen B, Olstad OK, Haug KB, Brusletto B, Sandvik L, Staff AC, et al. Global mirna expression analysis of serous and clear cell ovarian carcinomas identifies differentially expressed mirnas including mir-200c-3p as a prognostic marker. BMC Cancer. 2014;14:80.
**a B, Yang S, Liu T, Lou G. Mir-211 suppresses epithelial ovarian cancer proliferation and cell-cycle progression by targeting cyclin d1 and cdk6. Mol Cancer. 2015;14:57.
Wang CC, Chen JJ, Yang PC. Multifunctional transcription factor yy1: a therapeutic target in human cancer? Expert Opin Ther Targets. 2006;10:253–66.
Xu Y, Ohms SJ, Li Z, Wang Q, Gong G, Hu Y, et al. Changes in the expression of mir-381 and mir-495 are inversely associated with the expression of the mdr1 gene and development of multi-drug resistance. PLoS One. 2013;8, e82062.
Ramos YF, Stad R, Attema J, Peltenburg LT, van der Eb AJ, Jochemsen AG. Aberrant expression of hdmx proteins in tumor cells correlates with wild-type p53. Cancer Res. 2001;61:1839–42.
Ryu SY, Kim K, Lee WS, Kwon HC, Lee KH, Kim CM, et al. Synergistic growth inhibition by combination of adenovirus mediated p53 transfer and cisplatin in ovarian cancer cell lines. J Gynecol Oncol. 2009;20:48–54.
Liu Z, Gersbach E, Zhang X, Xu X, Dong R, Lee P, et al. Mir-106a represses the rb tumor suppressor p130 to regulate cellular proliferation and differentiation in high-grade serous ovarian carcinoma. Mol Cancer Res : MCR. 2013;11:1314–25.
Lee MH, Choi BY, Cho YY, Lee SY, Huang Z, Kundu JK, et al. Tumor suppressor p16(ink4a) inhibits cancer cell growth by downregulating eef1a2 through a direct interaction. J Cell Sci. 2013;126:1744–52.
Huang L, Wang HY, Li JD, Wang JH, Zhou Y, Luo RZ, et al. Kpna2 promotes cell proliferation and tumorigenicity in epithelial ovarian carcinoma through upregulation of c-myc and downregulation of foxo3a. Cell Death Dis. 2013;4, e745.
Gan Y, Mo Y, Johnston J, Lu J, Wientjes MG, Au JL. Telomere maintenance in telomerase-positive human ovarian skov-3 cells cannot be retarded by complete inhibition of telomerase. FEBS Lett. 2002;527:10–4.
Oishi T, Kigawa J, Minagawa Y, Shimada M, Takahashi M, Terakawa N. Alteration of telomerase activity associated with development and extension of epithelial ovarian cancer. Obstet Gynecol. 1998;91:568–71.
Gatcliffe TA, Monk BJ, Planutis K, Holcombe RF. Wnt signaling in ovarian tumorigenesis. Int J Gynecol Cancer : Off J Int Gynecol Cancer Soc. 2008;18:954–62.
Friedman RC, Farh KK, Burge CB, Bartel DP. Most mammalian mrnas are conserved targets of micrornas. Genome Res. 2009;19:92–105.
Krek A, Grun D, Poy MN, Wolf R, Rosenberg L, Epstein EJ, et al. Combinatorial microrna target predictions. Nat Genet. 2005;37:495–500.
Acknowledgments
This study was supported by grants from National Natural Science Foundation of China (no. 81472028).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This study was approved by the Medical Ethics Committee of the Affiliated Tumor Hospital of Harbin Medical University and all patients were provided informed consent.
Conflicts of interest
None
Additional information
Bairong **a and Huiyan Li contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplemental figure 1
miR-381 decreases downstream genes of Wnt signaling pathway in both SKOV3 and COV644 cells. *p < 0.05 compared to miR-Ctrl transfected cells (GIF 17 kb)
Rights and permissions
About this article
Cite this article
**a, B., Li, H., Yang, S. et al. MiR-381 inhibits epithelial ovarian cancer malignancy via YY1 suppression. Tumor Biol. 37, 9157–9167 (2016). https://doi.org/10.1007/s13277-016-4805-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-016-4805-8