Abstract
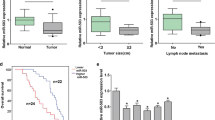
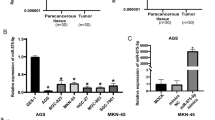
MicroRNAs (miRNAs) are a class of small noncoding RNAs that negatively regulate protein expression by binding protein-coding mRNAs and repressing translation. Accumulating evidence suggests that miRNAs are involved in cancer development and progression, acting as either tumor suppressors or oncogenes. Intriguingly, it has been shown that miR-133b was significantly downregulated in several types of cancers. However, its role and relevance in gastric cancer are still largely unknown. We showed that miR-133b was downregulated in human gastric cancer tissues and cell lines compared with nontumor counterparts by quantitative RT-PCR analysis. Overexpression of miR-133b could inhibit cell proliferation and colony formation of the gastric cancer cell lines MKN-45 and SGC-7901. Bioinformatics analysis indicated two putative miR-133b binding sites in the 3′-untranslated region of fibroblast growth factor receptor 1 (FGFR1) mRNA. In dual-luciferase reporter assay, miR-133b reduced the luciferase activity of Luc-FGFR1-wt, and mutation of miR-133b binding sites abolished the inhibitory effect of miR-133b. In this study, we found that miR-133b reduced the protein but not the mRNA levels of endogenous FGFR1. Furthermore, FGFR1 expression was upregulated in gastric cancer tissues and inversely correlated with miR-133b expression. Finally, knockdown of FGFR1 inhibited the growth of MKN-45 cells in a dose-dependent manner and overexpression of FGFR1 promoted the growth of GES-1 cells. These results indicate that miR-133b targets FGFR1 and inhibits gastric cancer cell growth, suggesting that it may serve as a tumor suppressive target in gastric cancer therapy.






Similar content being viewed by others
References
Wu CW, Hsiung CA, Lo SS, Hsieh MC, Chen JH, Li AF, et al. Nodal dissection for patients with gastric cancer: a randomised controlled trial. Lancet Oncol. 2006;7(4):309–15. doi:10.1016/S1470-2045(06)70623-4.
Lehnert T, Rudek B, Buhl K, Golling M. Surgical therapy for loco-regional recurrence and distant metastasis of gastric cancer. Eur J Surg Oncol. 2002;28(4):455–61. doi:S0748798302912606.
Varadhachary G, Ajani JA. Gastric cancer. Clin Adv Hematol Oncol. 2005;3(2):118–24.
Petrocca F, Visone R, Onelli MR, Shah MH, Nicoloso MS, de Martino I, et al. E2F1-regulated microRNAs impair TGFbeta-dependent cell-cycle arrest and apoptosis in gastric cancer. Cancer Cell. 2008;13(3):272–86. doi:10.1016/j.ccr.2008.02.013.
Ambros V. The functions of animal microRNAs. Nature. 2004;431(7006):350–5. doi:10.1038/nature02871nature02871.
Farh KK, Grimson A, Jan C, Lewis BP, Johnston WK, Lim LP, et al. The widespread impact of mammalian microRNAs on mRNA repression and evolution. Science. 2005;310(5755):1817–21. doi:10.1126/science.1121158.
Houbaviy HB, Murray MF, Sharp PA. Embryonic stem cell-specific microRNAs. Dev Cell. 2003;5(2):351–8. doi:S1534580703002272.
Croce CM. Causes and consequences of microRNA dysregulation in cancer. Nat Rev Genet. 2009;10(10):704–14. doi:10.1038/nrg2634.
Kent OA, Mendell JT. A small piece in the cancer puzzle: microRNAs as tumor suppressors and oncogenes. Oncogene. 2006;25(46):6188–96. doi:10.1038/sj.onc.1209913.
Volinia S, Calin GA, Liu CG, Ambs S, Cimmino A, Petrocca F, et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci U S A. 2006;103(7):2257–61. doi:10.1073/pnas.0510565103.
Panguluri SK, Bhatnagar S, Kumar A, McCarthy JJ, Srivastava AK, Cooper NG, et al. Genomic profiling of messenger RNAs and microRNAs reveals potential mechanisms of TWEAK-induced skeletal muscle wasting in mice. PLoS One. 2010;5(1):e8760. doi:10.1371/journal.pone.0008760.
Koutsoulidou A, Mastroyiannopoulos NP, Furling D, Uney JB, Phylactou LA. Expression of miR-1, miR-133a, miR-133b and miR-206 increases during development of human skeletal muscle. BMC Dev Biol. 2011;11:34. doi:10.1186/1471-213X-11-34.
Sucharov C, Bristow MR, Port JD. miRNA expression in the failing human heart: functional correlates. J Mol Cell Cardiol. 2008;45(2):185–92. doi:10.1016/j.yjmcc.2008.04.014.
Kim J, Inoue K, Ishii J, Vanti WB, Voronov SV, Murchison E, et al. A MicroRNA feedback circuit in midbrain dopamine neurons. Science. 2007;317(5842):1220–4. doi:10.1126/science.1140481.
Sanchez-Simon FM, Zhang XX, Loh HH, Law PY, Rodriguez RE. Morphine regulates dopaminergic neuron differentiation via miR-133b. Mol Pharmacol. 2010;78(5):935–42. doi:10.1124/mol.110.066837.
Bandres E, Cubedo E, Agirre X, Malumbres R, Zarate R, Ramirez N, et al. Identification by Real-time PCR of 13 mature microRNAs differentially expressed in colorectal cancer and non-tumoral tissues. Mol Cancer. 2006;5:29. doi:10.1186/1476-4598-5-29.
Wong TS, Liu XB, Chung-Wai Ho A, Po-Wing Yuen A, Wai-Man Ng R, Ignace Wei W. Identification of pyruvate kinase type M2 as potential oncoprotein in squamous cell carcinoma of tongue through microRNA profiling. Int J Cancer. 2008;123(2):251–7. doi:10.1002/ijc.23583.
Crawford M, Batte K, Yu L, Wu X, Nuovo GJ, Marsh CB, et al. MicroRNA 133B targets pro-survival molecules MCL-1 and BCL2L2 in lung cancer. Biochem Biophys Res Commun. 2009;388(3):483–9. doi:10.1016/j.bbrc.2009.07.143.
Ichimi T, Enokida H, Okuno Y, Kunimoto R, Chiyomaru T, Kawamoto K, et al. Identification of novel microRNA targets based on microRNA signatures in bladder cancer. Int J Cancer. 2009;125(2):345–52. doi:10.1002/ijc.24390.
Hu G, Chen D, Li X, Yang K, Wang H, Wu W. miR-133b regulates the MET proto-oncogene and inhibits the growth of colorectal cancer cells in vitro and in vivo. Cancer Biol Ther. 2010;10(2):190–7. doi:12186.
Akcakaya P, Ekelund S, Kolosenko I, Caramuta S, Ozata DM, **e H, et al. miR-185 and miR-133b deregulation is associated with overall survival and metastasis in colorectal cancer. Int J Oncol. 2011;39(2):311–8. doi:10.3892/ijo.2011.1043.
Qin W, Dong P, Ma C, Mitchelson K, Deng T, Zhang L, et al. MicroRNA-133b is a key promoter of cervical carcinoma development through the activation of the ERK and AKT1 pathways. Oncogene. 2011. doi:10.1038/onc.2011.561onc2011561.
Kano M, Seki N, Kikkawa N, Fujimura L, Hoshino I, Akutsu Y, et al. miR-145, miR-133a and miR-133b: tumor-suppressive miRNAs target FSCN1 in esophageal squamous cell carcinoma. Int J Cancer. 2010;127(12):2804–14. doi:10.1002/ijc.25284.
Patron JP, Fendler A, Bild M, Jung U, Muller H, Arntzen MO, et al. MiR-133b targets antiapoptotic genes and enhances death receptor-induced apoptosis. PLoS One. 2012;7(4):e35345. doi:10.1371/journal.pone.0035345PONE-D-11-20757.
Tao J, Wu D, Xu B, Qian W, Li P, Lu Q, et al. microRNA-133 inhibits cell proliferation, migration and invasion in prostate cancer cells by targeting the epidermal growth factor receptor. Oncol Rep. 2012;27(6):1967–75. doi:10.3892/or.2012.1711.
Guo J, Miao Y, **ao B, Huan R, Jiang Z, Meng D, et al. Differential expression of microRNA species in human gastric cancer versus non-tumorous tissues. J Gastroenterol Hepatol. 2009;24(4):652–7. doi:10.1111/j.1440-1746.2008.05666.x.
Freier K, Schwaenen C, Sticht C, Flechtenmacher C, Muhling J, Hofele C, et al. Recurrent FGFR1 amplification and high FGFR1 protein expression in oral squamous cell carcinoma (OSCC). Oral Oncol. 2007;43(1):60–6. doi:10.1016/j.oraloncology.2006.01.005.
Ishizuka T, Tanabe C, Sakamoto H, Aoyagi K, Maekawa M, Matsukura N, et al. Gene amplification profiling of esophageal squamous cell carcinomas by DNA array CGH. Biochem Biophys Res Commun. 2002;296(1):152–5. doi:S0006291X02008367.
Gorringe KL, Jacobs S, Thompson ER, Sridhar A, Qiu W, Choong DY, et al. High-resolution single nucleotide polymorphism array analysis of epithelial ovarian cancer reveals numerous microdeletions and amplifications. Clin Cancer Res. 2007;13(16):4731–9. doi:10.1158/1078-0432.CCR-07-0502.
Simon R, Richter J, Wagner U, Fijan A, Bruderer J, Schmid U, et al. High-throughput tissue microarray analysis of 3p25 (RAF1) and 8p12 (FGFR1) copy number alterations in urinary bladder cancer. Cancer Res. 2001;61(11):4514–9.
Edwards J, Krishna NS, Witton CJ, Bartlett JM. Gene amplifications associated with the development of hormone-resistant prostate cancer. Clin Cancer Res. 2003;9(14):5271–81.
Missiaglia E, Selfe J, Hamdi M, Williamson D, Schaaf G, Fang C, et al. Genomic imbalances in rhabdomyosarcoma cell lines affect expression of genes frequently altered in primary tumors: an approach to identify candidate genes involved in tumor development. Genes Chromosomes Cancer. 2009;48(6):455–67. doi:10.1002/gcc.20655.
Weir BA, Woo MS, Getz G, Perner S, Ding L, Beroukhim R, et al. Characterizing the cancer genome in lung adenocarcinoma. Nature. 2007;450(7171):893–8. doi:10.1038/nature06358.
Turner N, Pearson A, Sharpe R, Lambros M, Geyer F, Lopez-Garcia MA, et al. FGFR1 amplification drives endocrine therapy resistance and is a therapeutic target in breast cancer. Cancer Res. 2010;70(5):2085–94. doi:10.1158/0008-5472.CAN-09-3746.
Weiss J, Sos ML, Seidel D, Peifer M, Zander T, Heuckmann JM, et al. Frequent and focal FGFR1 amplification associates with therapeutically tractable FGFR1 dependency in squamous cell lung cancer. Sci Transl Med. 2010;2(62):62ra93. doi:10.1126/scitranslmed.3001451.
Tomlinson DC, Lamont FR, Shnyder SD, Knowles MA. Fibroblast growth factor receptor 1 promotes proliferation and survival via activation of the mitogen-activated protein kinase pathway in bladder cancer. Cancer Res. 2009;69(11):4613–20. doi:10.1158/0008-5472.CAN-08-2816.
**an W, Schwertfeger KL, Vargo-Gogola T, Rosen JM. Pleiotropic effects of FGFR1 on cell proliferation, survival, and migration in a 3D mammary epithelial cell model. J Cell Biol. 2005;171(4):663–73. doi:10.1083/jcb.200505098.
Acevedo VD, Gangula RD, Freeman KW, Li R, Zhang Y, Wang F, et al. Inducible FGFR-1 activation leads to irreversible prostate adenocarcinoma and an epithelial-to-mesenchymal transition. Cancer Cell. 2007;12(6):559–71. doi:10.1016/j.ccr.2007.11.004.
Funding
This work was supported by the Natural Science Foundation of Jilin Province (201215083) and the Special Foundation for Industrial Technology Research and Development of Jilin Province (2010018-5).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Wen, D., Li, S., Ji, F. et al. miR-133b acts as a tumor suppressor and negatively regulates FGFR1 in gastric cancer. Tumor Biol. 34, 793–803 (2013). https://doi.org/10.1007/s13277-012-0609-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-012-0609-7