Abstract
Since December 2019, the Coronavirus Disease (COVID-19) pandemic has caused world-wide turmoil in a short period of time, and the infection, caused by SARS-CoV-2, is spreading rapidly. AI-driven tools are used to identify Coronavirus outbreaks as well as forecast their nature of spread, where imaging techniques are widely used, such as CT scans and chest X-rays (CXRs). In this paper, motivated by the fact that X-ray imaging systems are more prevalent and cheaper than CT scan systems, a deep learning-based Convolutional Neural Network (CNN) model, which we call Truncated Inception Net, is proposed to screen COVID-19 positive CXRs from other non-COVID and/or healthy cases. To validate our proposal, six different types of datasets were employed by taking the following CXRs: COVID-19 positive, Pneumonia positive, Tuberculosis positive, and healthy cases into account. The proposed model achieved an accuracy of 99.96% (AUC of 1.0) in classifying COVID-19 positive cases from combined Pneumonia and healthy cases. Similarly, it achieved an accuracy of 99.92% (AUC of 0.99) in classifying COVID-19 positive cases from combined Pneumonia, Tuberculosis, and healthy CXRs. To the best of our knowledge, as of now, the achieved results outperform the existing AI-driven tools for screening COVID-19 using the acquired CXRs, and proves the viability of using the proposed Truncated Inception Net as a screening tool.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coronavirus Disease 2019 (COVID-19) is an infectious disease caused by Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) [1]. The disease was first identified in 2019 in Wuhan, China, and has since spread globally, resulting in the 2019–2020 Coronavirus pandemic [2]. With more than 5.92 million confirmed cases of infection and 364,000 cases of death by the fifth month of its discovery (as on May 30, 2020), the SARS-CoV-2 continues to infect people worldwide [3]. The virus is primarily transmitted among individuals through respiratory droplets. Studies have also shown that the virus can persist on surfaces which an infected individual might have touched. As a consequence, by the end of March 2020, the spread of this virus had been described as exponential [3].
The gold standard for the diagnosis and detection of COVID-19 is the polymerase chain reaction (PCR). It can detect the SARS-CoV-2 RNA from respiratory specimens through nasopharyngeal or oropharyngeal swabs. Despite the high sensitivity and accuracy of the PCR technique, the method is highly time-consuming and resource-intensive. Therefore, considering the unprecedented spread rate of the virus across the globe and the rapid temporal progression of the disease throughout a subject’s body [4], a faster screening tool is necessary for COVID-19 outbreaks. As an alternative to the traditional PCR technique, researchers have proposed the use of radiography techniques such as Computed Tomography (CT) scans and chest X-rays (CXRs) for COVID-19 screening. Early studies of COVID-19 positive patients have shown that their CT scans and CXRs show identifiable abnormalities [
Experimental setup
Datasets
Collecting COVID-19 dataset is not trivial. We, however, collect a number of CXR benchmark collections (C1 to C3) from the literature (See Table 1). They help to showcase/validate the usability and robustness of our model.
-
C1:
COVID-19 collection[28] is an open-source collection that is made available and maintained by Joseph Paul Cohen. At the time of the present study, it is composed of 162 COVID-19 positive CXRs, along with some other CXRs of diseases like MERS, SARS, and viral Pneumonia. For our purpose, only COVID-19 positive posteroanterior CXRs are considered.
-
C2:
Pneumonia collection[29] (Kaggle CXR collection) is composed of 5863 CXRs. Out of this, 1583 CXRs are normal or healthy CXRs and the remaining 4280 CXRs show various manifestations of viral and bacterial Pneumonia.
-
C3:
Two publicly available Tuberculosis (TB) collections[30] are considered: (a) Shenzhen, China and (b) Montgomery County, USA. These CXR benchmark collections were made available by the U.S. National Library of Medicine, National Institutes of Health (NIH). The Shenzhen, China collection is composed of 340 normal cases and 342 positive cases of TB. The Montgomery County, USA collection is composed of 80 normal CXRs and 58 TB positive CXRs.
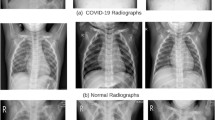
A few samples from the aforementioned collections are visualized in Fig. 5. Using aforementioned collections, we constructed six different combinations of data to train and validate our model. As provided in Table 2, these six different combinations of datasets (D1 to D6) are enlisted below:
-
D1:
In dataset D1, 162 COVID-19 positive CXRs and 340 healthy CXRs from the Shenzhen, China collections are considered.
-
D2:
For this dataset D2, 162 COVID-19 positive CXRs and 80 healthy CXRs from the Montgomery County, USA are considered.
-
D3:
D3 consists of 162 COVID-19 positive CXRs and 1583 healthy CXRs from the Pneumonia collections are considered.
-
D2:
D4 contains 162 COVID-19 positive CXRs and 2003 healthy CXRs, combined from the Shenzhen, Montgomery and Pneumonia collections are considered.
-
D5:
In dataset D5, 162 COVID-19 positive CXRs, 4280 Pneumonia positive CXRs and 1583 healthy CXRs from the Pneumonia collections are considered.
-
D6:
In dataset D6, 162 COVID-19 positive CXRs and 6683 non-COVID CXRs (comprising of 4280 Pneumonia positive, 400 TB positive and 2003 healthy CXRs) are considered.
The primary motivation behind constructing the various data combinations (D1 to D6) is to show the robustness of the Truncated Inception Net to detect COVID-19 positive cases. Further, COVID-19 is believed to have a close relationship with traditional Pneumonia. Therefore, a separate dataset (D5) was constructed to show whether our proposed model is able to differentiate COVID-19 positive cases from those traditional Pneumonia positive cases. Besides, CXRs of Tuberculosis manifestation were also added in D6 to prove that our model is robust enough to identify COVID-19 from other diseases like TB, Pneumonia, and healthy CXRs. The robustness also lies in the way we collect data, where regional variation can be considered as a crucial element. In our datasets, the healthy CXRs in D1, D2, and D3 are collected from different regions of the world. Considering multiple combination of data from different places can help develop cross-population train/test models.Footnote 1
As an input to our model, CXR images were scaled down to the size of 224\(\times \)224\(\times \)3 to match the input dimensions of the Truncated Inception Net. Such a resizing can also reduce computational complexity. Since the pixels of the CXRs have bounded discrete values, the images were normalized using the min-max scaling scheme. The choice is further backed by the fact that standardization (zero-mean unit variance) assumes the data to always have a Gaussian distribution that might not always be the case. Additionally, pixel intensities of COVID-19 features like GGO patches and consolidation falls in the same range of bones in CXRs, as demonstrated by quick preliminary experiments. So histogram matching was also excluded as a normalization scheme, since it decreases the signal to noise ratio in this scenario.
Validation protocol and evaluation metrics
To validate our proposed model, a 10 fold cross-validation scheme was opted for training and testing purposes on all six datasets: D1–D6. The process of 10 fold cross-validation works in the following way: say there are 100 data samples in the total dataset. Then samples 1–10 are made a subset and labelled as fold-1, samples 11–20 are labelled as fold-2 and so on. These creates 10 disjoint subsets of the original dataset. Following this, the model to be tested is first trained on subsets 1–9 and tested on subset 10. Similarly, in the second trial the model is tested on subset 9 after being trained on the remaining subsets. This scheme ensures that the model’s performance is not biased by the presence of outlier data samples in the training or testing datasets. Following this strategy, each of the constructed datasets (D1–D6) was subdivided into 10 subsets of almost equal number of data samples. Then the model was trained on 9 subsets and tested on the remaining 1 subset. This process was repeated using each of the subsets as a test set for once. After the ten separate trials of training and testing, the result was averaged over the ten trials to assess the mean (and standard deviation) performance of the model on that dataset. This procedure can be well understood by observing the result pattern in Table 4, which tabulates the tenfold cross-validation performance of the model on dataset D6. For each of the 10 folds, six different evaluation metrics were employed: (a) Accuracy (ACC); (b) Area under the ROC curve (AUC); (c) Sensitivity (SEN); (d) Specificity (SPEC); (e) Precision (PREC); and (f) F1 score. These can be computed as follows:
where \(t_{p}\), \(f_{p}\), \(t_{n}\), and \(f_{n}\) are the total number of true positives, false positives, true negatives, and false negatives. The mean scores from all 10 folds were taken for each of the above metrics, to get the final results on a particular dataset.
In traditional deep learning tasks, a primary metric like accuracy is sufficient to judge the performance of a deep learning model as binary classifier. On the contrary, such an assumption does not work well when considering imbalanced datasets. In such cases (like, in medical datasets), the positive class to be predicted often has much lower data samples than the negative class. Therefore, accuracy would demonstrate a fairly high value even if the model labels all the test data to be negative. Therefore, special attention is given to metrics like Sensitivity/Recall, Precision, and F1 score here.
In the context of COVID-19, the Sensitivity metric plays a very crucial role when deploying a model for screening patients in the early stages of a pandemic. Sensitivity measures the likelihood that the model would not miss classifying COVID-19 positive samples/patients. This prevents the further spreading of the infection. Secondly, the precision measures the likelihood that a model would not make a mistake to classify normal patients as COVID-19 positive. This metric becomes very important in the later stages of a pandemic, when medical resources are limited, and they are available only to the patients that are in need. Besides, F1 score is used to extract the combined performance score of a model, which is the harmonic mean of the precision and sensitivity of a model.
Results and analysis
Before providing quantitative results, we first provide activation maps generated by our proposed model for a COVID-19 positive, Pneumonia positive, and TB positive CXR can be visualized in Fig. 6. It can be observed that in the preliminary layers (like Conv2D), the lung region is clearly visible in the activation map for normal CXR, while the clarity gradually decreases for pneumonia and further for COVID-19 CXR. This corresponds to the growth of GGO patches in COVID-19 positive CXRs. However, in the later layers of the model, the activation maps become more abstract, for which the terminal dense layer is used in the model to map these abstract feature representations to their corresponding labels (COVID+ or COVID−).
Following the validation protocol and evaluation metrics mentioned in the previous “Validation protocol and evaluation metrics” section, we present the mean scores that were achieved using tenfold cross-validation train-test scheme, on each of the six different datasets: D1–D6. The experimental results are well documented in Table 3. Also, standard deviation (\(\sigma \)) is reported in all cases, whose very low value proves the statistical robustness of our model. Our proposed Truncated Inception Net model achieves a classification ACC, AUC, SEN, SPEC, PREC, and F1 score of 99.96%, 1.0, 0.98, 0.99, 0.98, and 0.98, respectively, on the dataset: D5 (COVID-19 positive case detection against Pneumonia and healthy cases) and that of 99.92%, 0.99, 0.93, 1.0, 1.0, and 0.96, respectively, on the D6 dataset (COVID-19 positive case detection against Pneumonia, TB, and healthy CXRs). Since the custom datasets being used were highly imbalanced in terms of class representation, sensitivity and precision are the most significant metrics in our case, as said in “Validation protocol and evaluation metrics” section. Consequently, the proposed model achieves high sensitivity and precision on these datasets. For a better understanding of the results, six different ROC curves are shown in Fig. 7; one for each dataset, starting from D1 to D6.
Additionally, since for every dataset we computed tenfold cross-validation, for better understanding of how average scores and their standard deviation were computed, the results obtained from each fold on the dataset: D6 are provided in Table 4. Besides, the proposed Truncated Inception Net model performs \(2.3\pm 0.18\) times on an average faster than Inception Net V3 model. In Table 5, computational times (by taking 10 different CXR samples as input) are used to demonstrate the differences between them. The primary reason being the large number of parameters in the original Inception Net V3 model. Precisely, this model contains more than 21.7 million trainable parameters in contrast our model which contains only 2.1 million trainable parameters, making it a better choice for training on small datasets and also for active learning. Therefore, for mass screening in resource-constrained areas, employing a faster tool is the must.
Discussion
Since COVID-19 outbreak, very few pieces of works have been proposed/reported using CXRs to detect COVID-19 positive cases (see “Introduction” section): In our comparison, ResNet50 and SVM [19], COVID-Net [20], ResNet50 [21], and Inception Net V3 [21] are considered even though they are not peer-reviewed research articles. We have compared with these pieces works using exact same evaluation metrics (ACC in %, AUC, SEN, SPEC, PREC, and F1 score) and nature of dataset. Like other works, we take COVID-19 positive and healthy CXRs from Pneumonia dataset (D3 in our case), and used this result as a comparison to other works. Besides, since all models were based on deep learning models, we consider an essential element i.e., number of parameters in our comparison. Table 6 provides a complete comparative study. Not all the authors reported AUC, SPEC, and F1 score. Note that, our model was used as a binary classifier to screen a CXR as COVID+ or COVID-, while not all the stated works performed the same. The mentioned results belong to the COVID+ positive class, wherever multi-class classification was done instead of binary classification. On the whole, considering the number of parameters, the proposed Truncated Inception Net outperforms all. Note that, since our model is the derivative of Inception Net V3 model, it is worth to compare between them. We observe that, in both computational time (Table 5) and performance scores (Table 6), Truncated Inception Net performs better than Inception Net V3 [21]. For a better understanding, three different performance scores: poor, the best and average are considered from Table 3. This suggests that the Truncated Inception Net is not only more computationally effective in terms of training and usability, but also more flexible for the purpose of active learning [14].
Even though the performed experiments validate that the proposed deep learning model for screening COVID-19 positive CXRs, it is important to understand that the system relies completely on visual cues in the input data. Therefore, in the early stages of COVID-19, when the radiologically observable cues have not yet developed, the system might fail to perform as stated. A detailed study on this is a scope for future work, where the input data shall be additionally labeled with the stage of COVID-19 it depicts as well. However, data acquired for this work did not contain any explicit information regarding the stages of COVID-19 in the individual CXRs. Further, the system is limited by its capacity to localize the disease in the CXR. As seen in the activation maps of deeper layers (Fig. 6), the model develops an intrinsic representation of the CXR features rather than accurate spatial heat-map, which is then mapped to the output using a dense layer classifier. The mentioned goal can be achieved by using increased number of data or a deep learning model(s) that is/are pre-trained on a large number of CXRs of different diseases (like CheXNet [31]), which shall be our future goal.
Conclusion and future works
In this work, we have proposed the Truncated Inception Net deep learning model to detect COVID-19 positive patients using chest X-rays. For validation, experimental tests were done on six different experimental datasets by combining COVID-19 positive, Pneumonia positive, Tuberculosis positive, and healthy CXRs. The proposed model outperforms the state-of-the-art results in detecting COVID-19 cases from non-COVID ones. Besides, considering the number of parameters used in our proposed model, it is computationally efficient as compared to original Inception Net V3 model and other works proposed in the literature. It is important to note that the study has no clinical implications. Instead, we solely aimed to check whether the proposed Truncated Inception Net could be used in detecting COVID-19 positive cases using CXRs.
Observing the performance scores, the Truncated Inception Net can serve as a milestone for screening COVID-19 under active-learning framework on multitudinal/multimodal data [14]. It also motivates to work on cross-population train/test models. Integrating this model with CheXNet model [31] will be our immediate plan, since ChexNet is primarily employed to analyze CXRs.