Abstract
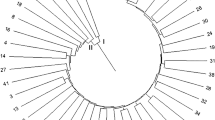
To assess the genetic diversity and relationship between varieties eighteen elite Indian sesame varieties were analysed using thirty sesame-specific SSR markers. A total of 134 alleles with an average number of 4.5 alleles were observed with PIC value ranging from 0.39 (CUESTSSR563) to 0.87 (CUSSR29). The dendrogram was constructed using the software NTSYS Pc (Ver. 2.20). The varieties were grouped into 5 clusters. The dissimilarity coefficients between sesame varieties based on molecular data ranged from 0.466 to 0.967. GT 10 and RT 351 showed highest distance value i.e. 0.967 and the lowest distance value was estimated between the genotypes Prachi and Nirmala. The study revealed that geographic origin is not related to genetic diversity as genotypes belonging to different geographical origin grouped in the same cluster. DNA profile of varieties exhibited good polymorphism which helps to distinctly identify and characterize sesame varieties.

Similar content being viewed by others
References
Ali ML, Rajewski JF, Baenziger PS, Gill KS, Eskridge KM, Dweikat I. Assessment of genetic diversity and relationship among a collection of US sweet sorghum germplasm by SSR markers. Mol Breed. 2008;21(4):497–509.
Ashri A. Sesame breeding. Plant Breed Rev. 1998;16:179–228.
Badri J, Yepuri V, Ghanta A, Siva S, Siddiq EA. Development of microsatellite markers in sesame (Sesamum indicum L.). Turk J Agric For. 2014;38:603–14.
Bedigian D, Seigler DS, Harlan JR. Sesamin, sesamolin and the origin of sesame. Biochem Syst Ecol. 1985;13:133–9.
Bedigian D. Evolution of sesame revisited: domestication, diversity and prospects. Genet Resour Crop Evol. 2003;50:779–87.
Bhattacharjee M, Prakash SH, Roy S, Begum T, Dasgupta T. DNA fingerprinting of CUMS 17 (Suprava): a newly developed variety of sesame. Int J Pure Appl Biosci. 2018;6(5):161–6.
Bhattacharyya U, Pandey SK, Dasgupta T. Identification of EST-SSRs and FDMin sesame (Sesamum indicum L.) through data mining. J Agric Sci. 2014;4:60–9.
Botstein D, White LR, Sholnick M, Davis RW. Construction of a genetic linkage map in man using restriction fragment length polymorphism. Am J Hum Genet. 1980;32:314–31.
Brar GS, Ahuja KL. Sesame, its culture, genetics, breeding and biochemistry. Annu Rev Plant Sci. 1979;1:245–313.
Dixit A, ** MH, Chung JW, Yu JW, Chung HK, Ma KH, Cho EG. Development of polymorphic microsatellite markers in sesame (Sesamum indicum L.). Mol Ecol Resour. 2005;5:736–8.
Dossa K, Yu J, Liao B, Cisse N, Zhang X. Development of highly informative genome-wide single sequence repeat markers for breeding applications in sesame and construction of a web resource: SisatBase. Front Plant Sci. 2017;8:1470.
Faostat, FAO Statistics Division. Food and Agriculture Organization of the United Nations. 2016.
Gao YL, Zhu RS, Liu CY, Li WF, Jiang HW, Li CD, Chen QS. Constructing molecular identity for soybean varieties from Heilongjiang Province, China. Acta Agron Sin. 2009;35:211–8.
Kola VSR, Yepuri V, Surapaneni M, Badri J, Vemireddy LR, Ghanta A, Siddiq EA. Genetic diversity and DNA fingerprinting in sesame (Sesamum indicum L.) cultivars of ANGRAU. Asian Australas J Plant Sci Biotechnol. 2012;16:98–101.
Kumar V, Sharma SN. Comparative potential of phenotypic, ISSR and SSR markers for characterization of sesame (Sesamum indicum L.) varieties from India. J Crop Sci Biotechnol. 2011;14:163–71.
Laurentin HE, Karlovsky P. Genetic relationship and diversity in a sesame (Sesamum indicum L.) germplasm collection using amplified fragment length polymorphism (AFLP). BMC Genet. 2006;7(1):10.
Laurentin H, Karlovsky P. AFLP fingerprinting of sesame (Sesamum indicum L.) cultivars: identification, genetic relationship and comparison of AFLP informativeness parameters. Genet Resour Crop Evol. 2007;54:1437–46.
Pandey SK, Das A, Rai P, Dasgupta T. Morphological and genetic diversity assessment of sesame (Sesamum indicum L.) accessions differing in origin. Physiol Mol Biol Plants. 2015;21:519–29.
Rajora O, Rahman M. Microsatellite DNA and RAPD fingerprinting, identification and genetic relationships of hybrid poplar (Populus canadensis) cultivars. Theor Appl Genet. 2003;106:470–7.
Rani A, Kumar V, Shukla S, Jha P, Rawal R. DNA barcoding of Indian soybean varieties as constructed through SSR markers. Seed Sci Technol. 2016;44:357–69.
Rohlf FJ NTSYS-pc. Numerical taxonomy and multivariate analysis system. Setauket: Exeter Publishing; 2000.
Spandana B, Reddy VP, John Prasanna G, Anuradha G, Sivaramakrishnan S. Development and characterization of microsatellite markers (SSR) in sesamum (Sesamum indicum L.) species. Appl Biochem Biotechnol. 2012;168:1594–607.
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW. Ribosomal DNA spacer-length polymorphisms in barley: mendelian inheritance, chromosomal location, and population dynamics. Proc Natl Acad Sci. 1984;81:8014–8.
Vijayarajan S, Siruthaiyur KG, Mahalingam G. Generation mean analysis for quantitative traits in sesam (Sesamum indicum L.) crosses. Genet Mol Biol. 2007;30:80–4.
Wei YB, Zhang HY, Zheng YZ, Miao HM, Zhang TZ, Guo WZ. A genetic linkage map construction for sesame (Sesamum indicum L.). Genes Genomics. 2009;31:207–16.
Wu K, Yang M, Liu H, Tao Y, Mei J, Zhao Y. Genetic analysis and molecular characterization of Chinese sesame [Sesamum indicum L.] cultivars using insertion–deletion [InDel] and simple sequence repeat [SSR] markers. BMC Genet. 2014;15(1):35.
Yepuri V, Surapaneni M, Kola VSR, Vemireddy LR, Jyothi B, Dineshkumar V, Siddiq EA. Assessment of genetic diversity in sesame (Sesamum indicum L.) genotypes, using EST-derived SSR markers. J Crop Sci Biotechnol. 2013;16:93–103.
Zhang L, Cai R, Yuan M, Tao A, Xu J, Lin L, Qi J. Genetic diversity and DNA fingerprinting in jute (Corchorus spp.) based on SSR markers. Crop J. 2015;3:416–22.
Acknowledgements
All authors are thankful to Dr. A.R.G. Ranganatha, former Project Co-coordinator (Sesame and Niger), Indian Council of Agricultural Research for providing us seeds of sesame genotypes. This study was carried out with the financial support of the Higher Education, Science and Technology and Biotechnology (Sci & Tech.), Government of West Bengal.
Funding
Funding was provided by Department of Science and Technology, Government of West Bengal (Grant No. (135 (Sanc.)/ST/P/S&T/ 1G-4/2014)).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Bhattacharjee, M., Prakash, S.H., Roy, S. et al. SSR-based DNA fingerprinting of 18 elite Indian varieties of sesame (Sesamum indicum L.). Nucleus 63, 67–73 (2020). https://doi.org/10.1007/s13237-019-00290-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13237-019-00290-3