Abstract
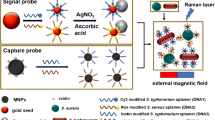
A novel surface-enhanced Raman scattering (SERS) response emerging from aptamers, complementary DNA (cDNA), p-aminothiophenol (PATP), and Au nanorods (GNRs) for detection of foodborne pathogens sensitivity is proposed here. In the presence of Salmonella typhimurium (ST) and cDNA simultaneously, ST and cDNA would competitively combine with the S. typhimurium aptamer (STA), inducing a highly conformation changes of STA. Accordingly, STA no longer stabilizes the GNRs in salt solution, leading to the varying aggregation extent of GNRs. The varying GNR aggregation will give rise to the plasmonic coupling and display a strong SERS signal, which can be distinctly reflected by the attached PATP via a transition of signals typical from ST peaks to PATP peaks. Under optimal conditions, the SERS intensity was observed to increase linearly with ST concentration from 56 to 56 × 107 cfu/mL (R 2 = 0.971), with a LOD of 9 cfu/mL. Additionally, this aptasensor exhibits a high selectivity to other similar pathogens, and the ability of the bioassay to detect ST was also confirmed in adulterated milk samples.





Similar content being viewed by others
References
Andrade MO, Farah CS, Wang N (2014) The post-transcriptional regulator rsmA/csrA activates T3SS by stabilizing the 5′ UTR of hrpG, the master regulator of hrp/hrc genes, in Xanthomonas. PLoS Pathog 10(2):e1003945–e1003945
Chen Z, Tabakman SM, Goodwin AP, Kattah MG, Dan D, Wang X, Zhang G, Li X, Liu Z, Utz PJ (2008) Protein microarrays with carbon nanotubes as multicolor Raman labels. Nat Biotechnol 26(11):1285–1292
Cho IH, Bhandari P, Patel P, Irudayaraj J (2015) Membrane filter-assisted surface enhanced Raman spectroscopy for the rapid detection of E. coli O157:H7 in ground beef. Biosens Bioelectron 64(64C):171–176
Cho IH, Mauer L, Irudayaraj J (2014) In-situ fluorescent immunomagnetic multiplex detection of foodborne pathogens in very low numbers. Biosens Bioelectron 57(5):143–148
Chung E, Jeon J, Yu J, Lee C, Choo J (2015) Surface-enhanced Raman scattering aptasensor for ultrasensitive trace analysis of bisphenol A. Biosens Bioelectron 64:560–565
Dastider S G, Barizuddin S, Yuksek N S, Dweik M, Almasri M F (2015) Efficient and rapid detection of salmonella using microfluidic impedance based sensing. Journal of Sensors 2015(7):861–866
Duan N, Wu S, Dai S, Miao T, Chen J, Wang Z (2015) Simultaneous detection of pathogenic bacteria using an aptamer based biosensor and dual fluorescence resonance energy transfer from quantum dots to carbon nanoparticles. Microchim Acta 182(5):917–923
Duan N, Chang B, Zhang H, Wang Z, Wu S (2016) Salmonella typhimurium detection using a surface-enhanced raman scattering-based aptasensor. Int J Food Microbiol 218:38
Farka Z, Juřík T, Pastucha M, Kovář D, Lacina K, Skládal P (2016)Rapid immunosensing of salmonella typhimurium using electrochemical impedance spectroscopy: the effect of sample treatment. Electroanalysis 28(8):1803–1809.
Fu C, Wang Y, Chen G, Yang L, Xu S, Xu W (2015) An aptamer-based SERS-microfluidic sensor for sensitive and selective polychlorinated biphenyls detection. Anal Chem 87(19):9555–9558
Gan Q, Zhang G, Wu Z, Shen A, Wang J, Hu J (2014) A “turn-off” SERS assay of heparin with high selectivity based on heparin–peptide complex and Raman labelled gold nanoparticles. Biosens Bioelectron 60C(60C):124–129
Giammanco GM, Aleo A, Guida I, Mammina C (2011) Molecular epidemiological survey of Citrobacter freundii misidentified as Cronobacter spp. (Enterobacter sakazakii) and Enterobacter hormaechei isolated from powdered infant milk formula. Foodborne Pathogens & Disease 8(4):517–525
Guntupalli R, Hu J, Lakshmanan R S, Huang T S, Barbaree J M, Chin B A (2007) A magnetoelastic resonance biosensor immobilized with polyclonal antibody for the detection of salmonella typhimurium. Biosens Bioelectron 22(7):1474–9
Hafner M, Vianini E, Albertoni B, Marchetti L, Grüne I, Gloeckner C, Famulok M (2008) Displacement of protein-bound aptamers with small molecules screened by fluorescence polarization. Nature Protocol 3(4):579–587
Hu W, Chen Q, Li H, Ouyang Q, Zhao J (2016) Fabricating a novel label-free aptasensor for acetamiprid by fluorescence resonance energy transfer between NH2-NaYF4: Yb, Ho@ SiO2 and Au nanoparticles. Biosens Bioelectron 80:398–404
Huang Y, Zhang H, Chen X, Wang X, Duan N, Wu S, Xu B, Wang Z (2015) A multicolor time-resolved fluorescence aptasensor for the simultaneous detection of multiplex Staphylococcus aureus enterotoxins in the milk. Biosens Bioelectron 74:170–176
Hui Z, Ma X, Ying L, Duan N, Wu S, Wang Z, Xu B (2015) Gold nanoparticles enhanced SERS aptasensor for the simultaneous detection of Salmonella typhimurium and Staphylococcus aureus. Biosens Bioelectron 74:872–877
**g N, Harpster MH, Wilson WC, Johnson PA (2013) Surface-enhanced Raman scattering (SERS) detection of multiple viral antigens using magnetic capture of SERS-active nanoparticles. Biosens Bioelectron 41(6):316–321
Kim NH, Lee SJ, Moskovits M (2010) Aptamer-mediated surface-enhanced Raman spectroscopy intensity amplification. Nano Lett 10(10):4181–4185
Kim YS, Niazi JH, Yun JC, Ko UR, Man BG (2011) Aptamers-in-liposomes for selective and multiplexed capture of small organic compounds. Macromol Rapid Commun 32(15):1169–1173
Ko J, Lee C, Choo J (2015) Highly sensitive SERS-based immunoassay of aflatoxin B1 using silica-encapsulated hollow gold nanoparticles. J Hazard Mater 285:11–17
Lai YH, Chen SW, Hayashi M, Shiu YJ, Huang CC, Chuang WT, Su CJ, Jeng HC, Chang JW, Lee YC (2014) Mesostructured arrays of nanometer-spaced gold nanoparticles for ultrahigh number density of SERS hot spots. Adv Funct Mater 24(17):2544–2552
Lantz PG, Hahn-Hägerdal B, Rådström P (1994) Sample preparation methods in PCR-based detection of food pathogens. Trends Food Sci Technol 5(12):384–389
Lermo A, Zacco E, Barak J, Delwiche M, Campoy S, Barbé J, Alegret S, Pividori MI (2008) Towards Q-PCR of pathogenic bacteria with improved electrochemical double-tagged genosensing detection. Biosens Bioelectron 23(12):1805–1811
Levy M (2011) Abstract LB-234: aptamer-prodrugs for targeted cancer therapy. Cancer Res 71(8):LB-234–LB-234
Li H, Chen Q, Mehedi Hassan M, Chen X, Ouyang Q, Guo Z, Zhao J (2017) A magnetite/PMAA nanospheres-targeting SERS aptasensor for tetracycline sensing using mercapto molecules embedded core/shell nanoparticles for signal amplification. Biosens Bioelectron 92:192–199
Miyahara T, Nakatsuji H (2015) Indicator of the stacking interaction in the DNA double-helical structure: ChiraSac study. J Phys Chem A 119(30):8269–8278
Mokhtarzadeh A, Dolatabadi JEN, Abnous K, Guardia MDL, Ramezani M (2015) Nanomaterial-based cocaine aptasensors. Biosens Bioelectron 68(68):95–106
Nakashima H, Furukawa K, Kashimura Y, Torimitsu K (2007) Anisotropic assembly of gold nanorods assisted by selective ion recognition of surface-anchored crown ether derivatives. Chem Commun 10(10):1080–1082
Nikoobakht B, El-Sayed MA (2003) Preparation and growth mechanism of gold nanorods (NRs) using seed-mediated growth method. Chem Mater 15(10):1957–1962
Palomo G, Campos MJ, Ugarte M, Porrero MC, Alonso JM, Borge C, Vadillo S, Domínguez L, Quesada A, Píriz S (2013) Dissemination of antimicrobial-resistant clones of Salmonella enterica among domestic animals, wild animals, and humans. Foodborne Pathogens & Disease 10(2):171–176
Pan W, Zhao J, Chen Q (2015) Fabricating upconversion fluorescent probes for rapidly sensing foodborne pathogens. Journal of Agricultural & Food Chemistry 63(36):8068
Perron GG, Quessy S, Bell G (2008) A reservoir of drug-resistant pathogenic bacteria in asymptomatic hosts. PLoS One 3(11):e3749
Pividori MI, Liébana S, Campoy S, Lermo A, Barbé J, Alegret S (2009) Magneto immunoseparation of pathogenic bacteria and electrochemical magneto genosensing of the double-tagged amplicon. Anal Chem 25(14):5812–5820
Rérole AL, Gobbo J, De TA, Schmitt E, Jp PDB, Hammann A, Lanneau D, Fourmaux E, Demidov ON, Deminov O (2011) Peptides and aptamers targeting HSP70: a novel approach for anticancer chemotherapy. Cancer Res 71(2):484–495
Seo K H, Brackett R E, Hartman N F, Campbell D P (1999) Development of a rapid response biosensor for detection of salmonella typhimurium. J Food Prot 62(5):431–437
Shlyahovsky B, Di L, Katz E, Willner I (2007) Proteins modified with DNAzymes or aptamers act as biosensors or biosensor labels. Biosens Bioelectron 22(11):2570–2576
Wang Y, Ye Z, ** J, **g S, Ying Y (2014) Development of an aptamer-based impedimetric bioassay using microfluidic system and magnetic separation for protein detection. Biosens Bioelectron 59(13):106–111
Wu S, Duan N, Shi Z, Fang C, Wang Z (2014) Simultaneous aptasensor for multiplex pathogenic bacteria detection based on multicolor upconversion nanoparticles labels. Anal Chem 86(6):3100–3107
** Z, Huang R, Li Z, He N, Wang T, Su E, Deng Y (2015) Selection of HBsAg-specific DNA aptamers based on carboxylated magnetic nanoparticles and their application in the rapid and simple detection of hepatitis B virus infection. ACS Appl Mater Interfaces 7(21):51–58
Yang C, Wang Y, Marty JL, Yang X (2011) Aptamer-based colorimetric biosensing of Ochratoxin a using unmodified gold nanoparticles indicator. Biosens Bioelectron 26(5):2724–2727
Yilmaz M, Ozdemir M, Erdogan H, Tamer U, Sen U, Facchetti A, Usta H, Demirel G (2015) Micro-/nanostructured highly crystalline organic semiconductor films for surface-enhanced Raman spectroscopy applications. Adv Funct Mater 25(35):5669–5676
Zeng Z, Liu Y, Wei J (2015) Recent advances in surface enhanced Raman spectroscopy (SERS): finite-difference time-domain (FDTD) method for SERS and sensing applications. TrAC Trends Anal Chem 75:162–173
Zhang L, Xu J, Mi L, Gong H, Jiang S, Yu Q (2012) Multifunctional magnetic-plasmonic nanoparticles for fast concentration and sensitive detection of bacteria using SERS. Biosens Bioelectron 31(1):130–136
Zor E, Moralesnarváez E, Zamoragálvez A, Bingol H, Ersoz M, Merkoçi A (2015) Graphene quantum dots-based Photoluminescent sensor: a multifunctional composite for pesticide detection. ACS Appl Mater Interfaces 7(36):20272–20279
Acknowledgements
This work has been financially supported by the National Natural Science Foundation of China (31471646), the Key R&D Program of Jiangsu Province (BE2015302), Postgraduate Innovative Program for Higher Education Institutions in Jiangsu Province (KYLX16_0913), the Natural Science Foundation of Jiangsu Province (Youth) (BK20150502), the China Postdoctoral Science Foundation (2015 M571698), and the Advanced Talents Science Foundation of Jiangsu University (15JDG064).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Huanhuan Li declares that she has no conflict of interest. Quansheng Chen declares that he has no conflict of interest. Qin Ouyang declares that she has no conflict of interest. Jiewen Zhao declares that he has no conflict of interest.
Funding
This study has no financial relationship with organization that sponsored the research.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed Consent
Not applicable.
Electronic supplementary material
ESM 1
(DOC 1887 kb)
Rights and permissions
About this article
Cite this article
Li, H., Chen, Q., Ouyang, Q. et al. Fabricating a Novel Raman Spectroscopy-Based Aptasensor for Rapidly Sensing Salmonella typhimurium . Food Anal. Methods 10, 3032–3041 (2017). https://doi.org/10.1007/s12161-017-0864-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12161-017-0864-8