Abstract
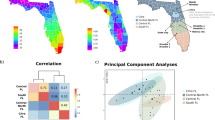
High-throughput phenoty** (HTP) enables breeders and researchers to have massive data sets accurately and objectively. It could be applied to plant breeding for screening stress tolerance and biodiversity among wild species in the gene bank, which can be a breakthrough in the phenoty** bottleneck. However, there are many factors to be considered. Thus, this study is designed to show an example of phenoty** traits using yield and image data in citrus using the Normalized Difference Vegetation Index (NDVI) and Red, Green, and Blue (RGB) images. The results using image analysis showed that R2 in linear regression ranged from 0.79 to 0.91, depending on the methods which were used in the current study. However, the results from NDVI were proven to be false, unlike those of RGB images. This means that researchers and breeders must be very cautious when dealing with new technologies to avoid being misled to the wrong conclusion when they try to associate this data with genomic data.






Similar content being viewed by others
Data availability
The datasets generated and analysed during the current study are available from the corresponding author on reasonable request.
References
Apolo-Apolo O, Martínez-Guanter J, Egea G, Raja P, Pérez-Ruiz M (2020) Deep learning techniques for estimation of the yield and size of citrus fruits using a UAV. Eur J Agron 115:126030
Feng L, Chen S, Zhang C, Zhang Y, He Y (2021) A comprehensive review on recent applications of unmanned aerial vehicle remote sensing with various sensors for high-throughput plant phenoty**. Comput Electron Agric 182:106033. https://doi.org/10.1016/j.compag.2021.106033
Iglesias DJ, Tadeo FR, Legaz F, Primo-Millo E, Talon M (2001) In vivo sucrose stimulation of colour change in citrus fruit epicarps: interactions between nutritional and hormonal signals. Physiol Plant 112:244–250
Kim M, Lee C, Hong S, Kim SL, Baek J-H, Kim K-H (2021) High-throughput phenoty** methods for breeding drought-tolerant crops. Int J Mol Sci 22:8266. https://doi.org/10.3390/ijms22158266
Otsu N (1979) A threshold selection method from gray-level histograms. IEEE Trans Syst Man Cybern 9:62–66
Van Rossum G, Drake F (2009) Python 3 reference manual createspace. Scotts Valley, CA
Shakoor N, Lee S, Mockler TC (2017) High throughput phenoty** to accelerate crop breeding and monitoring of diseases in the field. Curr Opin Plant Biol 38:184–192. https://doi.org/10.1016/j.pbi.2017.05.006
Tardieu F, Cabrera-Bosquet L, Pridmore T, Bennett M (2017) Plant phenomics, from sensors to knowledge. Curr Biol 27:R770–R783
Tattaris M, Reynolds MP, Chapman SC (2016) A direct comparison of remote sensing approaches for high-throughput phenoty** in plant breeding. Front Plant Sci 7:1131. https://doi.org/10.3389/fpls.2016.01131
Team RC (2013) R: a language and environment for statistical computing.
Vijayakumar V, Ampatzidis Y, Costa L (2023) Tree-level Citrus yield prediction utilizing ground and aerial machine vision and machine learning. Smart Agric Technol 3:100077
** traits using UAV-based sensors. Comput Electron Agric 178:105731. https://doi.org/10.1016/j.compag.2020.105731
Xu H, Ye Z, Ying Y (2005) Identification of citrus fruit in a tree canopy using color information. Trans Chin Soc Agric Eng 21:98–101
Acknowledgements
This research was funded by Project No. PJ01666701 for horticultural science and technological development from the National Institute of Horticultural and Herbal Science, Rural Development Administration, Republic of Korea.
Funding
Rural Development Administration, Project No. PJ01666701.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kwon, SH., Ku, K.B., Tomar, V. et al. Case study: things to be considered for high-throughput phenoty** in genomic studies. Plant Biotechnol Rep 17, 415–420 (2023). https://doi.org/10.1007/s11816-023-00834-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-023-00834-9