Abstract
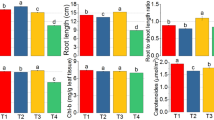
This study was carried out in a tissue culture medium using two salt stress-tolerant varieties (Oryza sativa L. cv. Dhan47 and Kızıltan mutant8) and two stress resistance varieties (Oryza sativa L. cv. Maraş Yellow Rice and Kızıltan) at 50 mM, 100 mM and 200 mM NaCl concentrations in two different periods (fifteen and thirty days). In this study, we analysed root and stem length, antioxidant enzyme activities and gene expressions. Among the varieties, the highest size was observed in the maraş variety, and the least length was observed in the dhan47 rice varieties in the 30-day stem length results. While there was no adverse effect on root lengths as much as 100 mm, it was observed that it had a negative impact on a 200 mM medium. In the 30-day results, it was observed that the SOD activity in the leaves of the Kızıltan variety decreased compared to the control, while it was observed that the activity increased up to 100 mM salt concentration in the roots. It was determined that CAT activity increased in leaves and roots at all concentrations compared to the control at increasing salt concentrations over a 15-day period. At the same time, it was observed that CAT activity was the highest at 100 mM salt concentration on the leaves of dhan 47 plant. In the 30-day results, it was observed that FDH activity increased in leaves and roots in all rice varieties compared to the control. In gene expression analyses, the highest (7 times) gene expression in roots was observed on Kızıltan100 mM salt concentration. The highest relative expression (14 times) in leaves was on the Kiziltan mutant leaf at 200 mM salt concentration. The results showed different responses in physiological effects, antioxidant activities and gene expression analyses as salt concentrations of all rice varieties. With the results obtained in this study, it is expected that different rice varieties that are sensitive and resistant to salt stress will contribute to the usability of agricultural lands where salinity is gradually increasing.





Similar content being viewed by others
Data availability
The authors confirm that data supporting the findings of this study are available in the article and Supplementary material. All data are included in Yunus Emre Arvas doctoral (Ph.D.) thesis. Raw data supporting the findings of this study are available from the corresponding author upon reasonable request.
Abbreviations
- ROS:
-
Reactive oxygen species
- GR:
-
Glutathione reductase
- POX:
-
Peroxidase
- SOD:
-
Superoxide dismutase
- GPX:
-
Guaiacol peroxidase
- CAT:
-
Catalase
- APX:
-
Ascorbate peroxidase
- FDH:
-
Formate dehydrogenase
- NAD+ :
-
Nikotinamid adenin dinücleotide
- MSR:
-
Methionine sulfoxide reductase
- EDTA:
-
Ethylene diamine tetraacetic acid
- Ms:
-
Murashige and Skoog medium
- PBS:
-
Phosphate-buffered saline
- BSA:
-
Bovine serum albumin
- mM:
-
Milimolar
- NBT:
-
Nitro blue tetrazole
- PMS:
-
Phenazine methosulfate
- ANOVA:
-
Analysis of variance
References
Apse MP, Blumwald E (2002) Engineering salt tolerance in plants. Curr Opin Biotechnol 13:146–150. https://doi.org/10.1016/S0958-1669(02)00298-7
Ashraf M (2009) Biotechnological approach of improving plant salt tolerance using antioxidants as markers. Biotechnol Adv 27:84–93. https://doi.org/10.1016/j.biotechadv.2008.09.003
Ayan A, Çelik Ö, Meriç S, Atak Ç (2017) Inducing salt tolerance in rice (Oryza sativa L.) varieties by gamma radiation. J Biotechnol 256:ss. 103-104. https://doi.org/10.1016/j.jbiotec.2017.06.1154
Bechtold U, Murphy DJ, Mullineaux PM (2004) Arabidopsis peptide methionine sulfoxide reductase2 prevents cellular oxidative damage in long nights. Plant Cell 16:908–919. https://doi.org/10.1105/tpc.015818
Bhaduri AM, Fulekar M (2012) Antioxidant enzyme responses of plants to heavy metal stress. Rev Environ Sci Biotechnol 11:55–69. https://doi.org/10.1007/s11157-011-9251-x
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72(1–2):248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Cakmak I, Marschner H (1992) Magnesium deficiency and high light intensity enhance activities of superoxide dismutase ascorbate peroxidase and glutathione reductase in bean leaves. Plant Physiol 98(4):1222–1227. https://doi.org/10.1104/pp.98.4.1222
Chang T-T, Map UB (2002) Origin, Domestication, and. Diversification. In book Rice: Origin, history, technology, and production 3, 3. Wiley Series in Crop Science. ISBN: 978–0–471–34516–9
Cho Y-W, Park E-H, Lim CJ (2000) Catalase, glutathione S-transferase and thioltransferase respond differently to oxidative stress in Schizosaccharomyces pombe. J Biochem Mol Biol 33:344–348
Chunthaburee S, Dongsansuk A, Sanitchon J, Pattanagul W, Theerakulpisut P (2016) Physiological and biochemical parameters for evaluation and clustering of rice cultivars differing in salt tolerance at seedling stage. Saudi J Biol Sci 23:467–477. https://doi.org/10.1016/j.sjbs.2015.05.013
Deinlein U, Aaron BS, Tomoaki H, Wei L, Guohua X, Julian IS (2014) Plant salt-tolerance mechanisms. Trends Plant Sci 19(6):201. s. 371-379. https://doi.org/10.1016/j.tplants.2014.02.001
Dikilitas M, Simsek E, Roychoudhury A (2020) Role of proline and glycine betaine in overcoming abiotic stresses. In book: Protective chemical agents in the amelioration of plant abiotic stress: biochemical and molecular perspectives. Wiley, Hoboken, pp. 1–23. https://doi.org/10.1002/9781119552154.ch1
Ding P, Gao Y, Zhu J, Chen F, **a G (2016) Wheat methionine sulfoxide reductase genes and their response to abiotic stress. Mol Breed 36:1–12. https://doi.org/10.1007/s11032-016-0597-1
Ding P, Fang L, Wang G et al (2019) Wheat methionine sulfoxide reductase A4.1 interacts with heme oxygenase 1 to enhance seedling tolerance to salinity or drought stress. Plant Mol Biol 101:203–220. https://doi.org/10.1007/s11103-019-00901-2
Fariduddin Q, Khalil RR, Mir BA, Yusuf M, Ahmad A (2013) 24-Epibrassinolide regulates photosynthesis, antioxidant enzyme activities and proline content of Cucumis sativus under salt and/or copper stress. Environ Monit Assess 185:7845–7856. https://doi.org/10.1007/s10661-013-3139-x
Farinelli MP, Fry DW, Richardson KE (1983) Isolation, purification, and partial characterization of formate dehydrogenase from soybean seed. Plant Physiol 73:858–859. https://doi.org/10.1104/pp.73.3.858
Flowers TJ (2004) Improving crop salt tolerance. J Exp Bot 55:307–319. https://doi.org/10.1093/jxb/erh003
Guan Q, Takano T, Liu S (2012) Genetic transformation and analysis of rice OsAPx2 gene in Medicago sativa. PLoS One 7:e41233. https://doi.org/10.1371/journal.pone.0041233
Guo X, Wu Y, Wang Y, Chen Y, Chu C (2009) OsMSRA4. 1 and OsMSRB1. 1, two rice plastidial methionine sulfoxide reductases, are involved in abiotic stress responses. Planta 230:227–238. https://doi.org/10.1007/s00425-009-0934-2
Hariadi YC, Nurhayati AY, Soeparjono S, Arif I (2015) Screening six varieties of rice (Oryzasativa) for salinity tolerance. Procedia Environ Sci 28:78–87. https://doi.org/10.1016/j.proenv.2015.07.012
Jalali P, Navabpour S, Yamchi A, Soltanloo H, Bagherikia S (2019) Differential responses of antioxidant system and expression profile of some genes of two rice genotypes in response to salinity stress. Biologia: 1–9. https://doi.org/10.2478/s11756-019-00393-x
Jamil A, Riaz S, Ashraf M, Foolad MR (2011) Gene expression profiling of plants under salt stress. Crit Rev Plant Sci 30(5):435–458. https://doi.org/10.1080/07352689.2011.605739
Kargov I, Kleimenov S, Savin S, Tishkov V, Alekseeva A (2015) Improvement of the soy formate dehydrogenase properties by rational design. Protein Eng Des Sel 28:171–178. https://doi.org/10.1093/protein/gzv007
Kumar V, Shriram V, Nikam T, Jawali N, Shitole MG (2009) Antioxidant enzyme activities and protein profiling under salt stress in indica rice genotypes differing in salt tolerance. Arch Agron Soil Sci 55:379–394. https://doi.org/10.1080/03650340802595543
Kurt-Gür G, Demirci H, Sunulu A, Ordu E (2018) Stress response of NAD+-dependent formate dehydrogenase in Gossypium hirsutum L. grown under copper toxicity. Environ Sci Pollut Res 25:31679–31690. https://doi.org/10.1007/s11356-018-3145-1
Läuchli A, Lüttge U (2002) Salinity: Environment-plants-molecules. Kluwer Academic Publishers, Dordrecht, pp 229–248 (ISBN: 978-1-4020-0492-6)
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2− ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Long R, Li M, Zhang T, Kang J, Sun Y, Cong L, Gao Y, Liu F, Yang Q (2016) Comparative proteomic analysis reveals differential root proteins in medicago sativa and medicago truncatula in response to salt stress. Front Plant Sci 7:424. https://doi.org/10.3389/fpls.2016.00424
Lou HQ, Gong YL, Fan W, Xu JM, Liu Y, Cao MJ, Wang M-H, Yang JL, Zheng SJ (2016) A formate dehydrogenase confers tolerance to aluminum and low pH. Plant Physiol 171:294–305. https://doi.org/10.1104/pp.16.01105
Ma Y, Dias MC, Freitas H (2020) Drought and salinity stress responses and microbe-induced tolerance in plants. Front Plant Sci: 1750. https://doi.org/10.3389/fpls.2020.591911
Mishra P, Bhoomika K, Dubey R (2013) Differential responses of antioxidative defense system to prolonged salinity stress in salt-tolerant and salt-sensitive Indica rice (Oryza sativa L.) seedlings. Protoplasma 250:3–19. https://doi.org/10.1007/s00709-011-0365-3
Miyama M, Tada Y (2008) Transcriptional and physiological study of the response of Burma mangrove (Bruguiera gymnorhiza) to salt and osmotic stress. Plant Mol Biol 68:119–129. https://doi.org/10.1007/s11103-008-9356-y
Monsur MB, Ivy NA, Haque MM, Hasanuzzaman M, El Sabagh A, Rohman MM (2020) Oxidative stress tolerance mechanism in rice under salinity. Phyton 89:497. https://doi.org/10.32604/phyton.2020.09144
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59(1):651–681. https://doi.org/10.1146/annurev.arplant.59.032607.092911
Ologundudu AF, Adelusi AA, Akinwale RO (2014) Effect of Salt Stress on Germination and Growth Parameters of Rice (Oryza sativa L.). Notulae Sci Biol 6. https://doi.org/10.15835/nsb629163
Pan Y, Wu LJ, Yu ZL (2006) Effect of salt and drought stress on antioxidant enzymes activities and SOD isoenzymes of liquorice (Glycyrrhiza uralensis Fisch). Plant Growth Regul 49:157–165. https://doi.org/10.1007/s10725-006-9101-y
Paul K, Pauk J, Kondic-Spika A, Grausgruber H, Allahverdiyev T, Sass L, Vass I (2019) Co-occurrence of mild salinity and drought synergistically enhances biomass and grain retardation in wheat. Front Plant Sci 10:501. https://doi.org/10.3389/fpls.2019.00501
Rahman MH, Naim MA, Matin MA, Anee TI, Hossain MA, Hasanuzzaman M (2016) Variations of growth parameters in some aromatic rice verities under salt stress. Bangladesh Agron J 19:1–10. https://doi.org/10.3329/baj.v19i2.31847
Ramesh R, Ramesh T, Rao P, Shankar V, Bhave M (2017) High temperature stress effected the biochemical parameters of rice (Oryza sativa L.) varieties and hybrids. Int J Pure App Biosci 5:1478–1490. https://doi.org/10.18782/2320-7051.5704
Razzaq A, Ali A, Safdar LB, Zafar MM, Rui Y, Shakeel A, Shaukat A, Ashraf M, Gong W, Yuan Y (2020) Salt stress induces physiochemical alterations in rice grain composition and quality. J Food Sci 85:14–20. https://doi.org/10.1111/1750-3841.14983
Romero HM, Berlett BS, Jensen PJ, Pell EJ, Tien M (2004) Investigations into the role of the plastidial peptide methionine sulfoxide reductase in response to oxidative stress in Arabidopsis. Plant Physiol 136:3784–3794. https://doi.org/10.1104/pp.104.046656
Roychoudhury A, Banerjee A, Lahiri V (2015) Metabolic and molecular-genetic regulation of proline signaling and ist cross-talk with major effectors mediates abiotic stress tolerance in plants. 39: 887-910. https://doi.org/10.3906/bot-1503-27
Sato Y, Tsuda K, Yamagata Y, Matsusaka H, Kajiya-Kanegae H, Yoshida Y, Agata A, Ta KN, Shimizu-Sato S, Suzuki T (2021) Collection, preservation and distribution of Oryza genetic resources by the National Bioresource Project RICE (NBRP-RICE). Breed Sci: 21005. https://doi.org/10.1270/jsbbs.21005
Sharma I, Ahmad P (2014) Catalase: a versatile antioxidant in plants, In book: Oxidative damage to plants. Elsevier, pp. 131–148. https://doi.org/10.1016/B978-0-12-799963-0.00004-6
Sharma S, Verslues PE (2010) Mechanisms independent of abscisic acid (ABA) or proline feedback have a predominant role in transcriptional regulation of proline metabolism during low water potential and stress recovery. Plant Cell Environ 33:1838–1851. https://doi.org/10.1111/j.1365-3040.2010.02188.x
Somboonwatthanaku I, Dorling S, Leung S, McManus MT (2010) Proline biosynthetic gene expression in tissue cultures of rice (Oryza sativa L.) in response to saline treatment. Plant Cell Tiss Org Cult (PCTOC) 103(3):369–376. https://doi.org/10.1007/s11240-010-9790-9
Szabados L, Savoure A (2010) Proline: a multifunctional amino acid. Trends Plant Sci 15:89–97. https://doi.org/10.1016/j.tplants.2009.11.009
Szekely G et al (2008) Duplicated P5CS genes of Arabidopsis play distinct roles in stress regulation and developmental control of proline biosynthesis. Plant J 53:11–28. https://doi.org/10.1111/j.1365-313X.2007.03318.x
Walia H, Wilson C, Condamine P, Liu X, Ismail AM, Zeng L, Wanamaker SI, Mandal J, Xu J, Cui X (2005) Comparative transcriptional profiling of two contrasting rice genotypes under salinity stress during the vegetative growth stage. Plant Physiol 139:822–835. https://doi.org/10.1104/pp.105.065961
Wang Z, Wang B, Zhao F, Cao J (2020) Genome-wide analysis of chitinase gene family in rice and Arabidopsis reveal their mechanisms and diverse roles in defense response. Int J Agric Biol 24:812–822. https://www.cabdirect.org/cabdirect/abstract/20203515042. Accessed 7 Feb 2023
Xu WF, Shi WM (2006) Expression profiling of the 14–3-3 gene family in response to salt stress and potassium and iron deficiencies in young tomato (Solanum lycopersicum) roots: analysis by real-time RT-PCR. Ann Bot 98:965–974. https://doi.org/10.1093/aob/mcl189
Yaghubi M, Nematzadeh G, Pirdashti H, Modarresi M, Motaghian A (2014) The effects of salinity on antioxidant enzymes activity in the leaves of two contrast rice (Oryza sativa L.) cultivars. Int J Biosci 4:116–125. https://www.researchgate.net/publication/276854072_The_effects_of_salinity_on_antioxidant_enzymes_activity_in_the_leaves_of_two_contrast_rice_Oryza_sativa_L_cultivars_International_Journal_of_Biosciences_IJB. Accessed 7 Feb 2023
Zhu JK (2001) Plant salt tolerance. Trends Plant Sci 6:66–71. https://doi.org/10.1016/S1360-1385(00)01838-0
Zhu J, Ding P, Li Q, Gao Y, Chen F, **a G (2015) Molecular characterization and expression profile of methionine sulfoxide reductase gene family in maize (Zea mays) under abiotic stresses. Gene 562:159–168. https://doi.org/10.1016/j.gene.2015.02.066
Acknowledgements
This report is a part of the Ph.D. Thesis of Yunus Emre Arvas, Ph.D. Candidate in Molecular Biology and Genetics Department, Yildiz Technical University. This study was supported by the Research Foundation of Yildiz Technical University, Project No: FBA-2020-3901
Author information
Authors and Affiliations
Contributions
All authors contributed to the study’s conception and design. Material preparation, data collection and analysis were performed by Yunus Emre Arvas, İsmail Kocacaliskan, Emel Ordu and Semiha Erişen. Yunus Emre Arvas and Emel Ordu wrote the first draft of the manuscript, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethical standards
The study is proper with ethical standards.
Consent to participate
Informed consent was obtained from all individual participants included in the study.
Consent for publication
The study does not include human and animal experiments.
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Arvas, Y.E., Kocacaliskan, İ., Erisen, S. et al. Antioxidant and molecular response of mutant and native rice (Oryza sativa L.) varieties grown under salt stress. Biologia 78, 1199–1210 (2023). https://doi.org/10.1007/s11756-023-01342-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11756-023-01342-5