Abstract
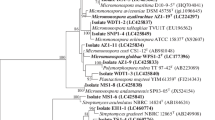
In this study, actinomycetes from roots and rhizospheric soils of leguminous plants were isolated using starch casein agar supplemented with antifungal and antibacterial antibiotics. Three hundred and seventeen actinomycetes were isolated with 77 isolates obtained from plant roots and 240 isolates from rhizospheric soils. Analysis of whole-organism hydrolysates showed that 289 strains were rich in the LL-isomer of diaminopimelic acid, a result consistent with their assignment to the streptomycetes. The remaining 28 strains were assigned to non-streptomycetes based on the presence of meso-isomer of diaminopimelic acid in cell wall. Sixty-four isolates (20.2 %) showed antagonistic activity against soybean pathogen Xanthomonas campestris pv. glycine by agar overlay method. Isolate RM 365 showed the highest activity with an inhibition ratio of 3.79, with no inhibitory activity on the growth of Rhizobium japonicum TISTR 079, Rhizobium sp. TISTR 061 and Rhizobium sp. TISTR 063. The 16S rRNA gene sequence analysis revealed that isolate RM 365 shared 99.28 % similarity to Streptomyces caeruleatus GIMN4T (GQ329712). In addition, isolates which contained meso-DAP were also identified by 16S rRNA gene sequence analysis. The results showed that they were members of the genus Amycolatopsis, Isoptericola, Micromonospora, Microbispora, Nocardia, Nonomuraea, Promicromonospora and Pseudonocardia.



Similar content being viewed by others
References
Adegboye FM, Babalola OO (2012) Taxonomy and ecology of antibiotic producing actinomycetes. Afr J Agric Res 7:2255–2261
Anand TP, Bhat AW, Shouche YS, Roy U, Siddharth J, Sarma SP (2006) Antimicrobial activity of marine bacteria associated with sponges from the waters off the coast of South East India. Microbiol Res 161:252–262
Arun B, Gopinath B, Sharma S (2012) Plant growth promoting potential of bacteria isolated on N free media from rhizosphere of Cassia occidentalis. World J Microbiol Biotechnol 28:2849–2857
Baltz RH (1998) Genetic manipulation of antibiotic-producing Streptomyces. Trends Microbiol 6:76–83
Basilio A, Gonzalez I, Vicente MF, Gorrochategui J, Cabello A, Gonzalez A, Genilloud O (2003) Patterns of antimicrobial activities from soil actinomycetes isolated under different conditions of pH and salinity. J Appl Microbiol 95:814–823
Becker B, Lechevalier MP, Gordon RE, Lechevalier HA (1964) Rapid differentiation between Nocardia and Streptomyces by paper chromatography of whole-cell hydrolysates. Appl Microbiol 12:421–423
Cao L, Qiu Z, Dai X, Tan H, Lin Y, Zhou S (2004) Isolation of endophytic actinomycetes from roots and leaves of banana (Musa acuminata) plants and their activities against Fusarium oxysporum f. sp. cubense. World J Microbiol Biotechnol 20:501–504
Castejón-Muñoz M, Oyarzun PJ (1995) Soil receptivity to Fusarium solani f. sp. pisi and biological control of root rot of pea. Eur J Plant Pathol 101:35–49
Castillo UF, Strobel GA, Ford EJ, Hess WM, Porter H, Jensen JB, Albert H, Robison R, Condron MA, Teplow DB, Stevens D, Yaver D (2002) Munumbicins, wide-spectrum antibiotics produced by Streptomyces NRRL 30562, endophytic on Kennedia nigriscans. Microbiol 148:2675–2685
Coombs JT, Franco CM (2003) Isolation and identification of actinobacteria from surface-sterilized wheat roots. Appl Environ Microbiol 69:5603–5608
Crawford DL, Lynch JM, Whipps JM, Ousley MA (1993) Isolation and characterization of actinomycete antagonists of a fungal root pathogen. Appl Environ Microbiol 59:3899–3905
Cross T, Maciver AM, Lacey J (1968) The thermophilic actinomycetes in mouldy hay: Micropolyspora faeni sp. nov. J Gen Microbiol 50:351–359
DeFrank J, Putnam AR (1985) Screening procedures to identify soil-borne actinomycetes that can produce herbicidal compounds. Weed Sci 33:271–274
Donghua J, Qinying L, Yiming S, Hao J (2013) Antimicrobial compound from a novel Streptomyces termitum strain ATC-2 against Xanthomonas oryzae pv. oryzae. Res J Biotech 8:66–70
Duangmal K, Mingma R, Pathom-Aree W, Thamchaipenet A, Inahashi Y, Matsumoto A, Takahashi Y (2011) Amycolatopsis samaneae sp. nov., isolated from roots of Samanea saman (Jacq.) Merr. Int J Syst Evol Microbiol 61:951–955
El-Tarabily KA, Nassar AH, Hardy GE, Sivasithamparam K (2009) Plant growth promotion and biological control of Pythium aphanidermatum, a pathogen of cucumber, by endophytic actinomycetes. J Appl Microbiol 106:13–26
Felsenstein J (1989) PHYLIP—phylogeny inference package (version 3.2). Cladistics 5:164–166
Germida JJ, Siciliano SD, Renato de Freitas J, Seib AM (1998) Diversity of root-associated bacteria associated with field-grown canola (Brassica napus L.) and wheat (Triticum aestivum L.). FEMS Microbiol Ecol 26:43–50
Gregor AK, Klubek B, Varsa EC (2003) Identification and use of actinomycetes for enhanced nodulation of soybean co-inoculated with Bradyrhizobium japonicum. Can J Microbiol 49:483–491
Gronemeyer JL, Burbano CS, Hurek T, Reinhold-Hurek B (2012) Isolation and characterization of root-associated bacteria from agricultural crops in the Kavango region of Namibia. Plant Soil 356:67–82
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hartman G, West E, Herman T (2011) Crops that feed the World 2. Soybean-worldwide production, use, and constraints caused by pathogens and pests. Food Sec 3:5–17
Hasegawa T, Takizawa M, Tanida S (1983) A rapid analysis for chemical grou** of aerobic actinomycetes. J Gen Appl Microbiol 29:319–322
Heisey RM, Defrank J, Putnam AR (1985) A survey of soil microorganisms for herbicidal activity. In: The chemistry of allelopathy. ACS Symposium Series, vol 268. American Chemical Society, pp 337–349
Jeffrey LSH, Sahilah AM, Son R, Tosiah S (2007) Isolation and screening of actinomycetes from Malaysian soil for their enzymatic and antimicrobial activities. J Trop Agric Food Sci 35:159–164
Kang YS, Lee Y, Cho SK, Lee KH, Kim BJ, Kim M, Lim Y, Cho M (2009) Antibacterial activity of a disaccharide isolated from Streptomyces sp. strain JJ45 against Xanthomonas sp. FEMS Microbiol Lett 294:119–125
Katz D, Sneh B, Friedman J (1987) The allelopathic potential of Coridothymus capitatus L. (Labiatae). Preliminary studies on the roles of the shrub in the inhibition of annuals germination and/or to promote allelopathically active actinomycetes. Plant Soil 98:53–66
Khamna S, Yokota A, Lumyong S (2009) Actinomycetes isolated from medicinal plant rhizosphere soils: diversity and screening of antifungal compounds, indole-3-acetic acid and siderophore production. World J Microbiol Biotechnol 25:649–655
Kieser T, Bibb MJ, Buttner MJ, Chater KF, Hopwood DA (2000) Practical Streptomyces genetics. John Innes Foundation, Norwich
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Küster E, Williams ST (1964) Selection of media for isolation of streptomycetes. Nature 202:928–929
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, New York, pp 115–175
Lechevalier HA, Lechevalier MP (1967) Biology of actinomycetes. Annu Rev Microbiol 21:71–100
Merckx R, Dijkstra A, Hartog A, Veen JA (1987) Production of root-derived material and associated microbial growth in soil at different nutrient levels. Biol Fertil Soils 5:126–132
Patil HJ, Srivastava AK, Singh DP, Chaudhari BL, Arora DK (2011) Actinomycetes mediated biochemical responses in tomato (Solanum lycopersicum) enhances bioprotection against Rhizoctonia solani. Crop Prot 30:1269–1273
Pillay VK, Nowak J (1997) Inoculum density, temperature, and genotype effects on in vitro growth promotion and epiphytic and endophytic colonization of tomato (Lycopersicon esculentum L.) seedlings inoculated with a pseudomonad bacterium. Can J Microbiol 43:354–361
Qin S, **ng K, Jiang JH, Xu LH, Li WJ (2011) Biodiversity, bioactive natural products and biotechnological potential of plant-associated endophytic actinobacteria. Appl Microbiol Biotechnol 89:457–473
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Samac DA, Willert AM, McBride MJ, Kinkel LL (2003) Effects of antibiotic-producing Streptomyces on nodulation and leaf spot in alfalfa. Appl Soil Ecol 22:55–66
Sharma S, Aneja MK, Mayer J, Munch JC, Schloter M (2005) Characterization of bacterial community structure in rhizosphere soil of grain legumes. Microb Ecol 49:407–415
Shirling EB, Gottlieb D (1966) Methods for characterization of Streptomyces species. Int J Syst Bacteriol 16:313–340
Soe KM, Bhromsiri A, Karladee D (2010) Effects of selected endophytic actinomycetes (Streptomyces sp.) and bradyrhizobia from Myanmar on growth, nodulation, nitrogen fixation and yield of different soybean varieties. CMU J Nat Sci 9:95–109
Somasegaran P, Hoben HJ (1985) Methods in legume-rhizobium technology. University of Hawaii NifTAL Project and MIRCEN, Dept. of Agronomy and Soil Science, Hawaii Institute of Tropical Agriculture and Human Resources, College of Tropical Agriculture and Human Resources
Sutherland ED, Lockwood JL (1984) Hyperparasitism of oospores of some peronosporales by Actinoplanes missouriensis and Humicola fuscoatra and other actinomycetes and fungi. Can J Plant Pathol 6:139–145
Taechowisan T, Lumyong S (2003) Activity of endophytic actinomycetes from roots of Zingiber officinale and Alpinia galangal against phytopathogenic fungi. Ann Microbiol 53:291–298
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Valan AM, Ignacimuthu S, Agastian P (2012) Actinomycetes from Western Ghats of Tamil Nadu with its antimicrobial properties. Asian Pac J Trop Biomed 2:830–837
Wellington EMH, Williams ST (1977) Preservation of actinomycete inoculum in frozen glycerol. Microbios Lett 6:151–157
Williams ST, Sharpe ME, Holt JG (1989) Bergey’s mannual of systematic bacteriology, vol 4, 2nd edn. Williams & Wilkins, Baltimore
Yuan WM, Crawford DL (1995) Characterization of Streptomyces lydicus WYEC108 as a potential biocontrol agent against fungal root and seed rots. Appl Environ Microbiol 61:3119–3128
Zhao K, Penttinen P, Chen Q, Guan T, Lindstrom K, Ao X, Zhang L, Zhang X (2012) The rhizospheres of traditional medicinal plants in Panxi, China, host a diverse selection of actinobacteria with antimicrobial properties. Appl Microbiol Biotechnol 94:1321–1335
Acknowledgments
This work was supported by the Higher Education Research Promotion and National Research University Project of Thailand, Office of the Higher Education Commission, and Kasetsart University Research and Development Institute (KURDI). We would also like to thank two anonymous referees for their constructive comments to improve the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mingma, R., Pathom-aree, W., Trakulnaleamsai, S. et al. Isolation of rhizospheric and roots endophytic actinomycetes from Leguminosae plant and their activities to inhibit soybean pathogen, Xanthomonas campestris pv. glycine . World J Microbiol Biotechnol 30, 271–280 (2014). https://doi.org/10.1007/s11274-013-1451-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-013-1451-9