Abstract
Phytocystatins are proteinaceous inhibitors of cysteine proteases. They have been implicated in the regulation of plant protein turnover and in defense against pathogens and insects. Here, we have characterized an Arabidopsis phytocystatin family gene, Arabidopsis thaliana phytocystatin 4 (AtCYS4). AtCYS4 was induced by heat stress. The heat shock tolerance of AtCYS4-overexpressing transgenic plants was greater than that of wild-type and cys4 knock-down plants, as measured by fresh weight and root length. Although no heat shock elements were identified in the 5′-flanking region of the AtCYS4 gene, canonical ABA-responsive elements (ABREs) and dehydration-responsive elements (DREs) were found. Transient promoter activity measurements showed that AtCYS4 expression was up-regulated in unstressed protoplasts by co-expression of DRE-binding factor 2s (DREB2s), especially by DREB2C, but not by bZIP transcription factors that bind to ABREs (ABFs, ABI5 and AREBs). DREB2C bound to and activated transcription from the two DREs on the AtCYS4 promoter although some preference was observed for the GCCGAC DRE element over the ACCGAC element. AtCYS4 transcript and protein levels were elevated in transgenic DREB2C overexpression lines with corresponding decline of endogenous cysteine peptidase activity. We propose that AtCYS4 functions in thermotolerance under the control of the DREB2C cascade.






Similar content being viewed by others
References
Almoguera C, Prieto-Dapena P, Díaz-Martín J, Espinosa JM, Carranco R, Jordano J (2009) The HaDREB2 transcription factor enhances basal thermotolerance and longevity of seeds through functional interaction with HaHSFA9. BMC Plant Biol 9:75–86
Belenghi B, Acconcia F, Trovato M, Perazzolli M, Bocedi A, Polticelli F, Ascenzi P, Delledonne M (2003) AtCYS1, a cystatin from Arabidopsis thaliana, suppresses hypersensitive cell death. Eur J Biochem 270:2593–2604
Benchabane M, Schlüter U, Vorster J, Goulet M-C, Michaud D (2010) Plant cystatins. Biochimie 92:1657–1666
Callis J (1995) Regulation of protein degradation. Plant Cell 7:845–857
Chen H, Hwang JE, Lim CJ, Kim DY, Lee SY, Lim CO (2010) Arabidopsis DREB2C functions as a transcriptional activator of HsfA3 during the heat stress response. Biochem Biophys Res Commun 401:238–244
Chen H, Je J, Song C, Hwang JE, Lim CO (2012) A proximal promoter region of Arabidopsis DREB2C confers tissue-specific expression under heat stress. J Integr Plant Biol 54:640–651
Chiu RS, Nahal H, Provart NJ, Gazzarrini S (2012) The role of the Arabidopsis FUSCA3 transcription factor during inhibition of seed germination at high temperature. BMC Plant Biol 12:15–30
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Diop NN, Kidrič M, Repellin A, Gareil M, d’Arcy-Lameta A, Pham Thi AT, Zuily-Fodil Y (2004) A multicystatin is induced by drought-stress in cowpea (Vigna unguiculata (L.) Walp.) leaves. FEBS Lett 577:545–550
Gaddour K, Vicente-Carbajosa J, Lara P, Isabel-Lamoneda I, Díaz I, Carbonero P (2001) A constitutive cystatin-encoding gene from barley (Icy) responds differentially to abiotic stimuli. Plant Mol Biol 45:599–608
Gong M, Li YJ, Chen SZ (1998) Abscisic acid-induced thermotolerance in maize seedlings is mediated by calcium and associated with antioxidant systems. J Plant Physiol 153:488–496
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database. Nucleic Acids Res 27:297–300
Hong JK, Hwang JE, Lim CJ, Yang KA, ** ZL, Kim CY, Koo JC, Chung WS, Lee KO, Lee SY, Cho MJ, Lim CO (2007) Over-expression of Chinese cabbage phytocystatins 1 retards seed germination in Arabidopsis. Plant Sci 172:556–563
Huang B, Xu C (2008) Identification and characterization of proteins associated with plant tolerance to heat stress. J Integr Plant Biol 50:1230–1237
Huang Y, **ao B, **ong L (2007) Characterization of a stress responsive proteinase inhibitor gene with positive effect in improving drought resistance in rice. Planta 226:73–85
Hwang JE, Hong JK, Je JH, Lee KO, Kim DY, Lee SY, Lim CO (2009) Regulation of seed germination and seedling growth by an Arabidopsis phytocystatin isoform, AtCYS6. Plant Cell Rep 28:1623–1632
Hwang JE, Hong JK, Lim CJ, Chen H, Je J, Yang KA, Kim DY, Choi YJ, Lee SY, Lim CO (2010) Distinct expression patterns of two Arabidopsis phytocystatin genes, AtCYS1 and AtCYS2, during development and abiotic stresses. Plant Cell Rep 29:905–915
Jakoby M, Dröge-Laser W, Vicente-Carbajosa J, Tiedemann J, Kroj T, Parsy F (2002) bZIP transcription factors in Arabidopsis. Trends Plant Sci 7:106–111
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: β-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Kim SY (2006) The role of ABF family bZIP class transcription factors in stress response. Physiol Plant 126:519–527
Larkindale J, Knight MR (2002) Protection against heat stress-induced oxidative damage in Arabidopsis involves calcium, abscisic acid, ethylene, and salicylic acid. Plant Physiol 128:682–695
Larkindale J, Vierling E (2008) Core genome responses involved in acclimation to high temperature. Plant Physiol 146:748–761
Lata C, Prasad M (2011) Role of DREBs in regulation of abiotic stress responses in plants. J Exp Bot 62:4731–4748
Lee SJ, Kang JY, Park HJ, Kim MD, Bae MS, Choi HI, Kim SY (2010) DREB2C interacts with ABF2, a bZIP protein regulating abscisic acid-responsive gene expression and its overexpression affects abscisic acid sensitivity. Plant Physiol 153:716–727
Lescot M, Dehais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouze P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327
Li S, Fu Q, Huang W, Yu D (2009) Functional analysis of an Arabidopsis transcription factor WRKY25 in heat stress. Plant Cell Rep 28:683–693
Li S, Zhou X, Chen L, Huang W, Yu D (2010) Functional characterization of Arabidopsis thaliana WRKY39 in heat stress. Mol Cells 29:475–483
Li Z, Yue H, **ng D (2012) MAP kinase 6-mediated activation of vacuolar processing enzyme modulates heat shock-induced programmed cell death in Arabidopsis. New Phytol 195:85–96
Lim CJ, Yang KA, Hong JK, Choi JS, Yun D-J, Hong JC, Chung WS, Lee SY, Cho MJ, Lim CO (2006) Gene expression profiles during heat acclimation in Arabidopsis thaliana suspension-culture cells. J Plant Res 119:373–383
Lim CJ, Hwang JE, Chen H, Hong JK, Yang KA, Choi MS, Lee KO, Chung WS, Lee SY, Lim CO (2007) Over-expression of the Arabidopsis DRE/CRT-binding transcription factor DREB2C enhances thermotolerance. Biochem Biophys Res Commun 362:431–436
Margis R, Reis EM, Villeret V (1998) Structural and phylogenetic relationships among plant and animal cystatins. Arch Biochem Biophys 359:24–30
Martínez M, Díaz-Mendoza M, Carrillo L, Díaz I (2007) Carboxy terminal extended phytocystatins are bifunctional inhibitors of papain and legumain cysteine proteinases. FEBS Lett 581:2914–2918
Martínez M, Cambra I, González-Melendi P, Santamaría ME, Díaz I (2012) C1A cysteine-proteases and their inhibitors in plants. Physiol Plant 145:85–94
Massonneau A, Condamine P, Wisniewski J-P, Zivy M, Rogowsky PM (2005) Maize cystatins respond to developmental cues, cold stress and drought. Biochim Biophys Acta 1729:186–199
Matsukura S, Mizoi J, Yoshida T, Todaka D, Ito Y, Maruyama K, Shinozaki K, Yamaguchi-Shinozaki K (2010) Comprehensive analysis of rice DREB2-type genes that encode transcription factors involved in the expression of abiotic stress-responsive genes. Mol Genet Genomics 283:185–196
Neuteboom LW, Matsumoto KO, Christopher DA (2009) An extended AE-rich N-terminal trunk in secreted pineapple cystatin enhances inhibition of fruit bromelain and is posttranslationally removed during ripening. Plant Physiol 151:515–527
Otlewski J, Jelen F, Zakrzewska M, Oleksy A (2005) The many faces of protease-protein inhibitor interaction. EMBO J 24:1303–1310
Park HC, Kim ML, Kang YH, Jeon JM, Yoo JH, Kim MC, Park CY, Jeong JC, Moon BC, Lee JH, Yoon HW, Lee S-H, Chung WS, Lim CO, Lee SY, Hong JC, Cho MJ (2004) Pathogen- and NaCl-induced expression of the SCaM-4 promoter is mediated in part by a GT-1 box that interacts with a GT-1-like transcription factor. Plant Physiol 135:2150–2161
Pernas M, Sánchez-Monge R, Salcedo G (2000) Biotic and abiotic stress can induce cystatin expression in chestnut. FEBS Lett 467:206–210
Sakuma Y, Liu Q, Dubouzet J, Abe H, Shinozaki K, Yamaguchi-Shinozakia K (2002) DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression. Biochem Biophys Res Commun 290:998–1009
Sakuma Y, Maruyama K, Osakabe Y, Qin F, Seki M, Shinozaki K, Yamaguchi-Shinozakia K (2006) Functional analysis of an Arabidopsis transcription factor, DREB2A, involved in drought-responsive gene expression. Plant Cell 18:1292–1309
Schöffl F, Prandl R, Reindl A (1998) Regulation of the heat-shock response. Plant Physiol 117:1135–1141
Schramm F, Larkindale J, Kiehlmann E, Ganguli A, Englich G, Vierling E, Koskull-Döring P (2008) A cascade of transcription factor DREB2A and heat stress transcription factor HsfA3 regulates the heat stress response of Arabidopsis. Plant J 53:264–274
Solomon M, Belenghi B, Delledonne M, Menachem E, Levine A (1999) The involvement of cysteine proteases and protease inhibitor genes in the regulation of programmed cell death in plants. Plant Cell 11:431–444
Su P, Li H (2008) Arabidopsis stromal 70-kD heat shock proteins are essential for plant development and important for thermotolerance of germinating seeds. Plant Physiol 146:1231–1241
Turk V, Bode W (1991) The cystatins: protein inhibitors of cysteine proteinases. FEBS Lett 285:213–219
Van der Vyver C, Schneidereit J, Driscoll S, Turner J, Kunert K, Foyer CH (2003) Oryzacystatin I expression in transformed tobacco produces a conditional growth phenotype and enhances chilling tolerance. Plant Biotechnol J 1:101–112
Vierling E (1991) The roles of heat shock proteins in plants. Annu Rev Plant Physiol Plant Mol Biol 42:579–620
Wang Y, Zhan Y, Wu C, Gong S, Zhu N, Chen S (2012) Cloning of a cystatin gene from sugar beet M14 that can enhance plant salt tolerance. Plant Sci 192:93–99
Wu C (1995) Heat shock transcription factors: structure and regulation. Annu Rev Cell Dev Biol 11:441–469
Yoshida T, Sakuma Y, Todaka D, Maruyama K, Qin F, Mizoi J, Kidokoro S, Fujita Y, Shinozaki K, Yamaguchi-Shinozaki K (2008) Functional analysis of an Arabidopsis heat-shock transcription factor HsfA3 in the transcriptional cascade downstream of the DREB2A stress-regulatory system. Biochem Biophys Res Commun 368:515–521
Zhang X, Liu S, Takano T (2008a) Two cysteine proteinase inhibitors from Arabidopsis thaliana, AtCYSa and AtCYSb, increasing the salt, drought, oxidation and cold tolerance. Plant Mol Biol 68:131–143
Zhang X, Wollenweber B, Jiang D, Liu F, Zhao J (2008b) Water deficits and heat shock effects on photosynthesis of a transgenic Arabidopsis thaliana constitutively expressing ABP9, a bZIP transcription factor. J Exp Bot 59:839–848
Acknowledgments
This work was supported by the Next-Generation BioGreen 21 Program (SSAC, grant no. PJ009514032013, Rural Development Administration, Republic of Korea) and Basic Science Research Program (Grant No. 2010-0009175, through the NRF funded by the Ministry of Education, Science and Technology, Republic of Korea). J. Je was supported by BK21 fellowship from the Ministry of Education, Science and Technology, Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
11248_2013_9735_MOESM3_ESM.tif
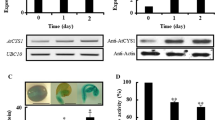
Supplementary Fig. S1 Analysis of AtCYS4 overexpression and knock-down lines. A Schematic diagram of the AtCYS4 gene from the putative transcription start site (TSS; -1) to the end of the AtCYS4 ORF (+492). The position of the T-DNA insertion (+74) in 5’-UTR of the knock-down mutant is shown by the inverted triangle. The position and orientation of the P4 to P5 primers that were used for RT-PCR is shown by the arrows. B Comparison of AtCYS4 transcript levels. RT-PCR analysis was performed on total RNA isolated from 10-day-old plants of WT, two independent Prom 35S :AtCYS4 transgenic lines (OX18, OX21) and the AtCYS4 knock-down mutant (cys4-KD) using primers P2 and P5. Actin2 gene (At3g18780) is shown as loading control. C Comparison of AtCYS4 protein levels. Shown is a western blot of resolved total protein extracts of 10-day-old plants that was performed using anti-AtCYS4 polyclonal antibody and visualized using an ECL system. D Comparison of endogenous CP activities. CP activities were measured in total protein extracts of 10-day-old plants and are expressed as a percentage of the mean CP activity of WT extracts. Bars represent mean ±S.E. (n = 3; *P < 0.05, **P < 0.01 by Student’s t-test). (TIFF 147699 kb)
Rights and permissions
About this article
Cite this article
Je, J., Song, C., Hwang, J.E. et al. DREB2C acts as a transcriptional activator of the thermo tolerance-related phytocystatin 4 (AtCYS4) gene. Transgenic Res 23, 109–123 (2014). https://doi.org/10.1007/s11248-013-9735-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11248-013-9735-2